The rollout of vaccines against the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), agent behind the notorious and catastrophic coronavirus disease 2019 (COVID-19) pandemic, was expected to help usher in the end of the extreme social and economic devastation associated with containment measures.
Natural selection and VOCs
The emergence of recent mutant strains, called variants of concern (VOCs), in the UK, South Africa and Brazil, has shattered expectations of a speedy end to the pandemic. Prominent vaccine makers are now stressing the need for repeated boosters after a year of the original course, or less, along with updated vaccines to overcome the loss of efficacy against the newer variants.
The SARS-CoV-2 genome is a large one, composed of ribonucleic acid (RNA), with a high frequency of mutations in many key genes. The viral spike is a prominent site of mutation, altering the epitopes or recognition sites for many of the commonly encountered neutralizing antibodies elicited by natural infection or vaccines.
Many of the newer variants are selected by the use of convalescent serum or therapeutic antibodies, as well as vaccine-induced antibodies, allowing them to rapidly supersede earlier variants in the local region in which they emerge. Moreover, they are often associated with more serious diseases.
Natural selection acts not only at the population level, however, but also in individuals with chronic infection. Repeated sequencing of the virus from samples obtained from the same patient over time, in such cases, shows that antibody-escape mutations are selected when the patient is treated with high-titer convalescent plasma containing polyclonal antibodies.
Such patients also tend to shed virus for weeks at high levels, thus favoring transmission of such immune-evading variants produced by prolonged selection pressures to those around them.
Important VOCs
The UK variant, B.1.1.7, is highly infectious and also more deadly than its preceding D614G mutant strain. It has a high rate of viral replication, which has led to its rapid rise to dominance in the region.
The SA variant B.1.351 is adept at escaping neutralization altogether and causes reinfections among those who already had the infection once. Both these VOCs, along with P.2, are resistant to monoclonal antibodies.
B.1.351 and B.1.429 resist neutralization by convalescent plasma and vaccine-induced antibodies. The latter property is shared by multiple other VOCs, including B.1.1.7, B1.298, P.2, and P.1.
Study aims
The current study aims to explore the emergence of such mutants so as to design treatment protocols that prevent the occurrence of such scenarios.
The role of chance in the emergence of VOCs
The researchers sought to understand the inherent stochasticity in viral evolution. This will assist the development of efforts to target it. In other words, their aim was to increase the role of such unpredictability, or chance, and thus reduce the role of natural selection and other predictable factors that improve viral fitness.
The favorable factors in this kind of situation include the stochastic generation of genetic diversity in the viral genome. This occurs through random errors in the replication of the genome within the host cells. In fact, the viral population in any infected host exists as a quasispecies, comprising a number of related sequences that result from de novo mutations occurring during viral population.
As the virus propagates within the different organs of the same host, different sequences come to make up the quasispecies. Genetic drift and obstacles to transmission within specific tissues play their roles in opposing the expansion of some viral mutations beyond the original small population, however advantageous they are.
Transmission of viral variants that arise in such a situation is necessary to allow them to survive and to expand their numbers. A limiting factor is the chance that they will die out because their numbers do not fulfill the low threshold required to establish a new infection.
Along with this, all variants arising in a single host do not have the same infectivity or ability to establish themselves widely. Those mutations that increase the rate of replication have the advantage in frequency among the quasispecies, and thus are more likely to be transmitted.
At least five infections must occur with the same strain for it to be established in the population without extinction. This is especially so since most of the spread originates from a limited number of individuals. This means that mutations associated with rapid replication can emerge at the population level much faster.
A higher viral load also favors the emergence of multiple mutations if the infection is prolonged.
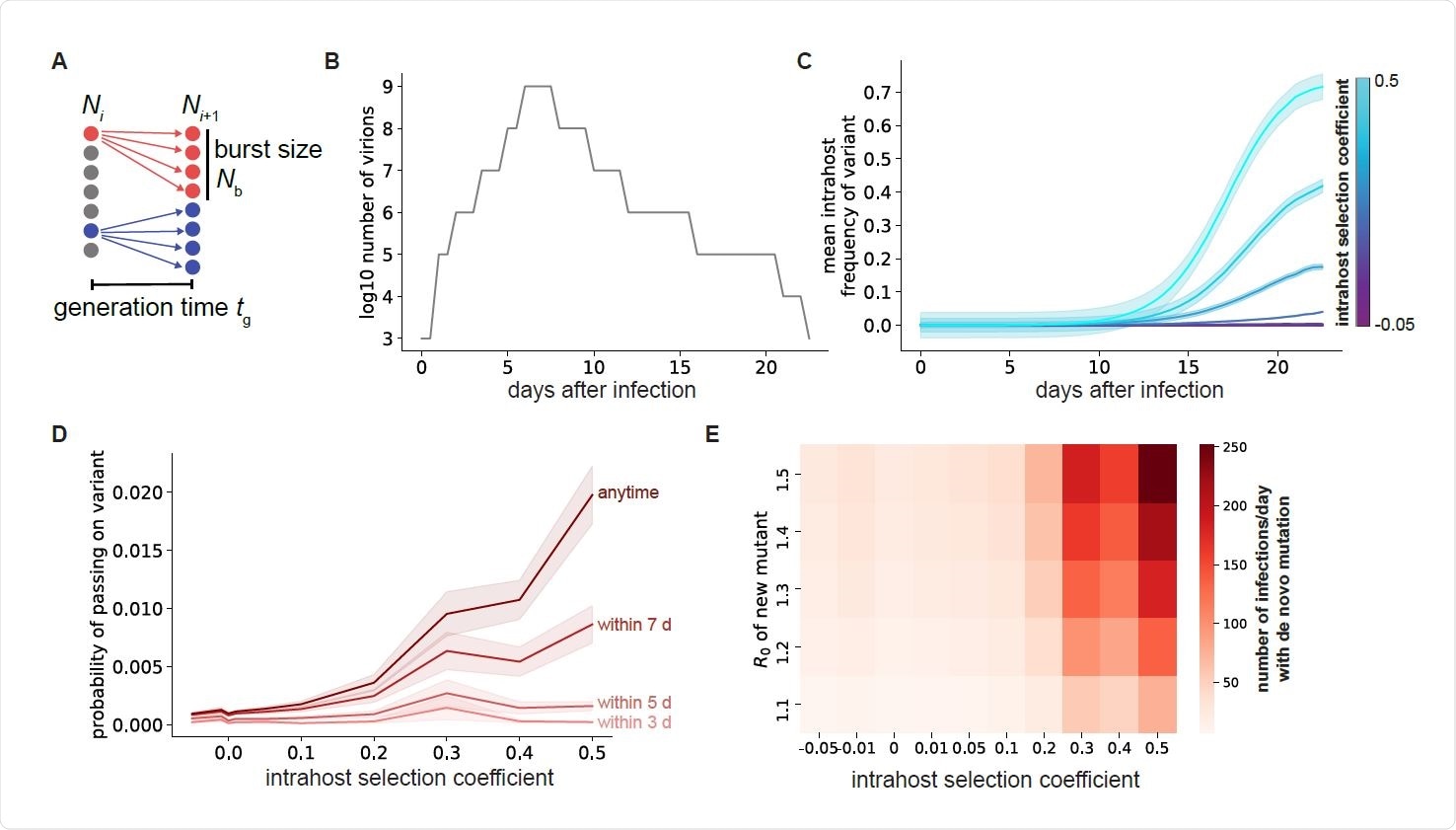
Selection within individuals with COVID-19 leads to selection of more fit viral variants. A. Schematic of viral replication model used to simulate SARS-CoV-2 evolutionary dynamics. B. Sputum viral load curve for a typical COVID-19 infection. The x-axis represents the time starting from the initial transmission event that started the infection. C. Mean frequency of variants with point mutations within individuals with COVID-19 for different mutation fitness effects (colors). D. Probability of a specific single mutation to be present in at least one virion transmitted if transmission occurs within the first 3-7 days of infection (lighter red curves) or anytime during infection (darker red). For C and D, shaded areas represent ± SEM, n=1000 simulations per condition. E. Number of total new single mutant infections generated per day that establish a surviving variant lineage, assuming all infections are of standard length with viral load profile given in B and that transmission occurs within the first 7 days of infection.
Transmission of single mutations depends on fitness
The study shows that fitter mutants show greater expansion, increasing the chances that at least one such particle will reach a new host and establish an infection. The later in infection this occurs, the higher the chances of successful transmission of an advantageous variant.
This is because, over time, such variants become more frequent as a result of natural selection. If they have a moderate transmission advantage enabling them to spread through the population, they can give rise at the population level to a rapidly emerging new lineage with beneficial point mutations.
Chronic infection and high viral load
Patients who harbor the virus for longer periods of time and have higher viral loads are more likely to spread the virus efficiently. The converse happens with reducing viral loads. This is due to the increased number of actively replicating viral particles present in both these situations.
Paired-mutation combinations
A recent study showed that a patient with immunocompromise developed COVID-19, which persisted for many weeks. This was associated with the rise of a mutation that allowed the virus to escape neutralization by antibodies, even though it was less infective.
However, the latter deficit was overcome by the co-occurrence of another mutation that increases the ease of transmission. Such highly advantageous combinations obtained via an in-between period when the virus is actually less fit may require prolonged infections, as in this case.
This is because this type of combination is present in very few viral particles over the short period of a typical SARS-CoV-2 infection. Chronic infections (30 days or more from the onset of symptoms) are thus necessary to arrive at a frequency that is high enough to lead to a realistic probability of successful transmission to another host.
Thus, just as such individuals increase the rate at which new strains with multiple or clustered mutations arise, a population with a higher number of such chronic infections will be more likely to produce variants with two mutations and with greater fitness.
What are the implications?
Several opportunities exist to hinder the emergence of new fitter variants by this process. First, the viral load within patients should be rapidly reduced over the full spectrum of symptoms, even in asymptomatic cases. This should make use of three or more antiviral agents that prevent viral replication by affecting proteins other than the spike, thus minimizing the chances of rapid resistance.
The result will be a significantly lower risk that immune-evading variants will occur following widespread exposure to convalescent plasma.
Secondly, long-term infections must be prevented irrespective of whether the patient is symptomatic or not, in view of the threat they pose in terms of the accelerated emergence of viral variants.
This provides a rationale for the requirement that a patient is tested negative before ending isolation, whatever the severity of the illness.
Transmission must also be blocked so that such variants do not get a chance to establish themselves. This includes the use of NPIs and vaccination. Contact tracing and genomic sequencing of secondary cases must also be persevered with to detect dangerous variants early.
The transmissibility of such variants must be specifically reduced to limit their spread by using multiple preventives that bind to different sites on the virus.
These measures are largely impractical at the present time, requiring further investments into research to differentiate chronic infections from false-positive PCRs.
A grimmer possibility is also presented, namely, that chronic infections predispose to the evolution of the virus into more lethal rather than less virulent forms, as was earlier supposed. This is seen with the current pandemic but has also been observed with other global outbreaks of HIV, the Spanish flu, and the rabbit disease called myxomatosis or the “white blindness.”
And finally, these findings should inform the development of new vaccines and other preventives that interrupt the routes of viral evolution and transmission, helping to contain the pandemic.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Van Egeren, D. et al. (2021). Controlling long-term SARS-CoV-2 infections is important for slowing viral evolution. medRxiv preprint. doi: https://doi.org/10.1101/2021.04.10.21255251, https://www.medrxiv.org/content/10.1101/2021.04.10.21255251v1
- Peer reviewed and published scientific report.
Van Egeren, Debra, Alexander Novokhodko, Madison Stoddard, Uyen Tran, Bruce Zetter, Michael S. Rogers, Diane Joseph-McCarthy, and Arijit Chakravarty. 2021. “Controlling Long-Term SARS-CoV-2 Infections Can Slow Viral Evolution and Reduce the Risk of Treatment Failure.” Scientific Reports 11 (1): 22630. https://doi.org/10.1038/s41598-021-02148-8. https://www.nature.com/articles/s41598-021-02148-8.