COVID-19 cases continue to increase all over the world. Several vaccines have now been approved and are being administered. However, with new variants of the virus emerging, there have been questions around how effective the vaccines will be.
The genome in coronaviruses is encoded by ribonucleic acid (RNA), making them vulnerable to RNA interference, especially when they are delivered to the lungs. However, there are few therapies that target viral RNA.
Small interfering RNA (siRNA) are short double-stranded RNA molecules. These interfere with the expression of specific genes and prevent translation. siRNA molecules can be delivered to the lungs via the nose or intravenously.
In a new study, researchers screened several siRNAs that target the highly conserved regions of SARS-CoV-2 and can inhibit virus replication. They reported their results in a paper published on the bioRxiv* preprint server.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Selecting siRNA
The team designed siRNAs targeted to the virus RNA polymerase, helicase, and the 5’ untranslated leader region (5’UTR). The team characterized the SARS-CoV-2 genome for structural features and sequence conservation, and looked for sequences that were absent in the human transcriptome.
Using this information, they generated about 9,500 candidate siRNA sequences using computational siRNA design tools. From these, they selected 18 siRNAs for further testing. In vitro testing of these candidates showed varying responses, with about five showing the best decrease in expression of virus genes, also called repression. Two of these can target both SARS-CoV and SARS-CoV-2.
siRNA can be stabilized by further chemical changes, which allows it to be more potent in vivo and show stronger virus repression. The authors chose siUTR3 for further chemical modification because its target site is in a highly conserved region of 5’UTR, important for viral processing and expression of viral RNA.
A 2’ O-methyl modification in siUTR3 increased serum stability and maintaining the level of virus repression, but the potency of SARS-CoV-2 repression was reduced. Further optimization of the modifications can lead to a more stable and potent siRNA for in vivo delivery.
Previous studies on RNA viruses have shown that a single siRNA can lead to the virus mutating, while combinations can reduce the emergence of viral variants. So the team tested three potent siRNAs alone and in combination. The combination showed the same level of virus repression as each of them alone, although their individual concentrations in the combination were lower.
Delivery vehicle for siRNA
The team had previously designed “stealth” lipid nanoparticles (sLNPs) for intravenous delivery of siRNAs to the lungs. The thick mucus associated with COVID-19 may likely hinder delivery by aerosols, and an intravenous method may prove advantageous. They modified this formulation to deliver SARS-CoV-2 siRNA to a mouse model expressing the human angiotensin-converting enzyme 2 (ACE2).
Mice infected with SARS-CoV-2 and injected with the sLNP-siRNA showed an improved chance of survival, and these mice had lower weight loss compared to untreated mice. Although the virus was fully repressed three days after siRNA administration, this effect was lost after six days, suggesting the repression is only transient and appears a day or two after injection. Daily injections may thus be required during peak viral loads.
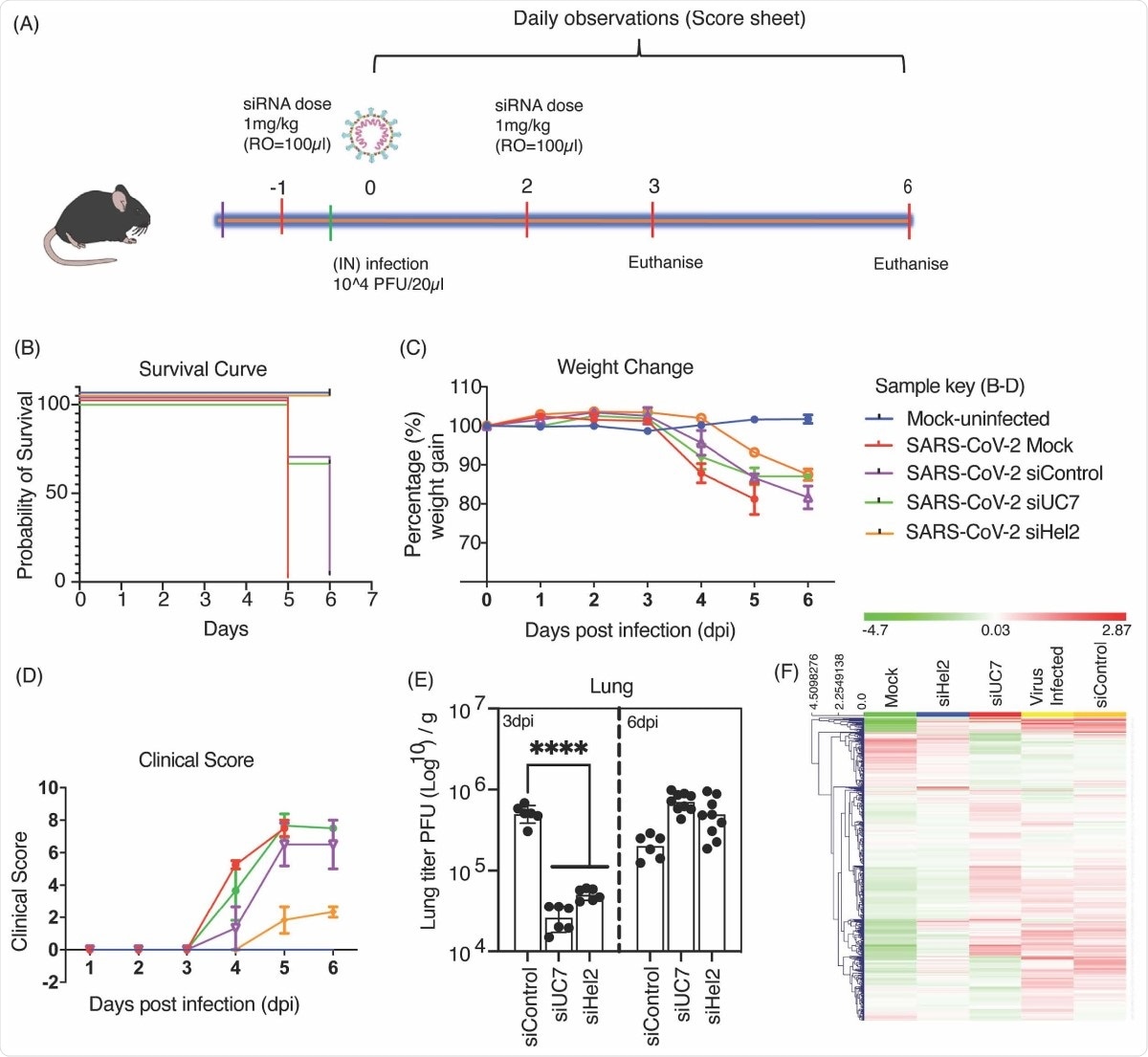
Intravenous administered sLNP-siRNA repression of COVID-19 in vivo. (A) 7–13-week-old K18-hACE.2 female and male mice were intranasally infected with either PBS or 104 PFU/ 20 μL of SARS-CoV-2 (Australian VIC1 strain, passage 4). (A) Mice were intravenously treated with 1 mg/kg in 100 μL of siRNA packaged into HFDM lipid nanoparticles (LNP) by retroorbital administration at -1 and 2 days post infection (dpi). At 3dpi and 6dpi lung and brain tissues were harvested and homogenized for immunoplaque assays. (B-D) Mouse survivorship, probability of survival, body weight (weight change), and clinical score were evaluated at the indicated days post-infection (dpi). (B) Weight loss >15% was taken as an endpoint and mice were euthanized. (C) Mice were weighed and scored daily until the experimental end point (6 dpi), for disease progression. (D) The clinical score was evaluated based on locomotion, behaviour and appearance. Each data point represents the average ± SEM of 3 to 4 mice. (E) The amount of infectious virus particles in lung tissues at 3 and 6 dpi were titrated by immunoplaque assays on Vero E6 cells, using a SARS-CoV-2 N protein specific antibody and expressed as PFU per gram of tissue. Each data point represents a technical replicate, where one mouse is equivalent to 3 technical replicates and bars represent the average ± SEM. (F) An unsupervised hierarchical cluster heatmap of immune gene expression in the lungs at 6dpi. Each row is a gene and each column is a treatment group. Rows are Z-score normalised (Green: low expression and red: high expression). A p value of <0.0001 (****) is considered statistically significant when assessed by 2-way ANOVA (Dunnett’s post- test) when compared against siControl.
The researchers did not see any immune dysregulation because of the siRNAs. They also further modified the formulation to ensure the reduction of any immune stimulation because of the lipid.
They did this by reducing the amount of a cationic lipid, which has been seen to activate the immune system. Reducing the amount of this lipid in the formulation reduced the positive charge by almost half. Similar to the previous formulation, this formulation also showed effective repression of the virus and better survival of the mice.
Both the Moderna and Pfizer vaccines use LNP as a delivery vehicle. Their success in LNP clinical translation helps the idea that LNPs are safe to use as delivery vehicles for mRNA and RNAi. Building upon this, this study shows LNPs can also deliver siRNA and have the potential for use as a COVID-19 treatment.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources