A team of scientists from Australia has recently identified a frameshift deletion mutation in ORF7a of the delta variant of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The mutation results in the production of a truncated peptide with altered functionality. The study is currently available on the medRxiv* preprint server while awaiting peer review.

 *Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
Background
SARS-CoV-2, the causative pathogen of coronavirus disease 2019 (COVID-19), is an enveloped, positive-sense, single-stranded RNA virus with a genome size of about 30kb. The genome of SARS-CoV-2 encodes four structural proteins, including envelope, membrane, nucleocapsid, and spike proteins, which play important roles in viral budding and entry into host cells.
In addition, the genome encodes multiple accessory proteins, including ORF3b, ORF6, and ORF7a, which are involved in the suppression of host immune responses by antagonizing type 1 interferon (IFN) response. Studies investigating SARS-CoV-2 accessory proteins have shown that ORF7a acts as an antagonist of a host pro-apoptotic protein, namely bone marrow stromal cell antigen 2 (BST2). A reduction in BST2 expression is known to increase the replication of SARS-CoV-2 in the host cell, leading to increased release of infectious virions from the host cell.
In some SARS-CoV-2 variants detected in March 2020, an 81-nucleotide in-frame deletion in ORF7a has been identified. This mutation is associated with the removal of signal peptide and beta strand sequences. Despite occurring frequently, deletion mutations in ORF7a are generally transient, most probably because of host immunity-mediated reduction in viral fitness.
In Australia, a recent surge in new COVID-19 cases has been observed. These cases are primarily associated with different SARS-CoV-2 delta variant infections. For mutational analysis of these variants, the scientists have sequenced SARS-CoV-2 genomes isolated from recent COVID-19 cases detected in Australia.
Important observations
Based on the study findings, the delta variants currently circulating in Australia are associated with a 17-nucleotide frameshift deletion mutation in ORF7a with minimal other alterations in the gene. The deletion mutation spanning the genome positions 27607–27623 resulted in the generation of a truncated peptide that is incapable of suppressing host cell BST2.
The consensus genome of the first virus of current Australian cases showed 100% similarity with the sequence of a complete ORF7a obtained from the USA. This could be the possible progenitor. Importantly, the first Australian case of the current outbreak and close contacts exhibited only 25% sequence similarity with ORF7a with a 17-nucleotide deletion mutation.
In Australia, the first delta genome with consensus-level ORF7a deletion mutation was identified in June, which accounted for 72% of viral genomes in the Australian COVID-19 outbreak. However, as mentioned by the scientists, it could be an underestimated value as the analysis includes travelers returning to Australia and incomplete sequences.
By analyzing 392,734 whole-genome sequences of the delta variants from the Global Initiative on Sharing Avian Influenza Data (GISAID) database, it was observed that almost all delta variants with ORF7a frameshift deletion mutation originate from current Australian cases.
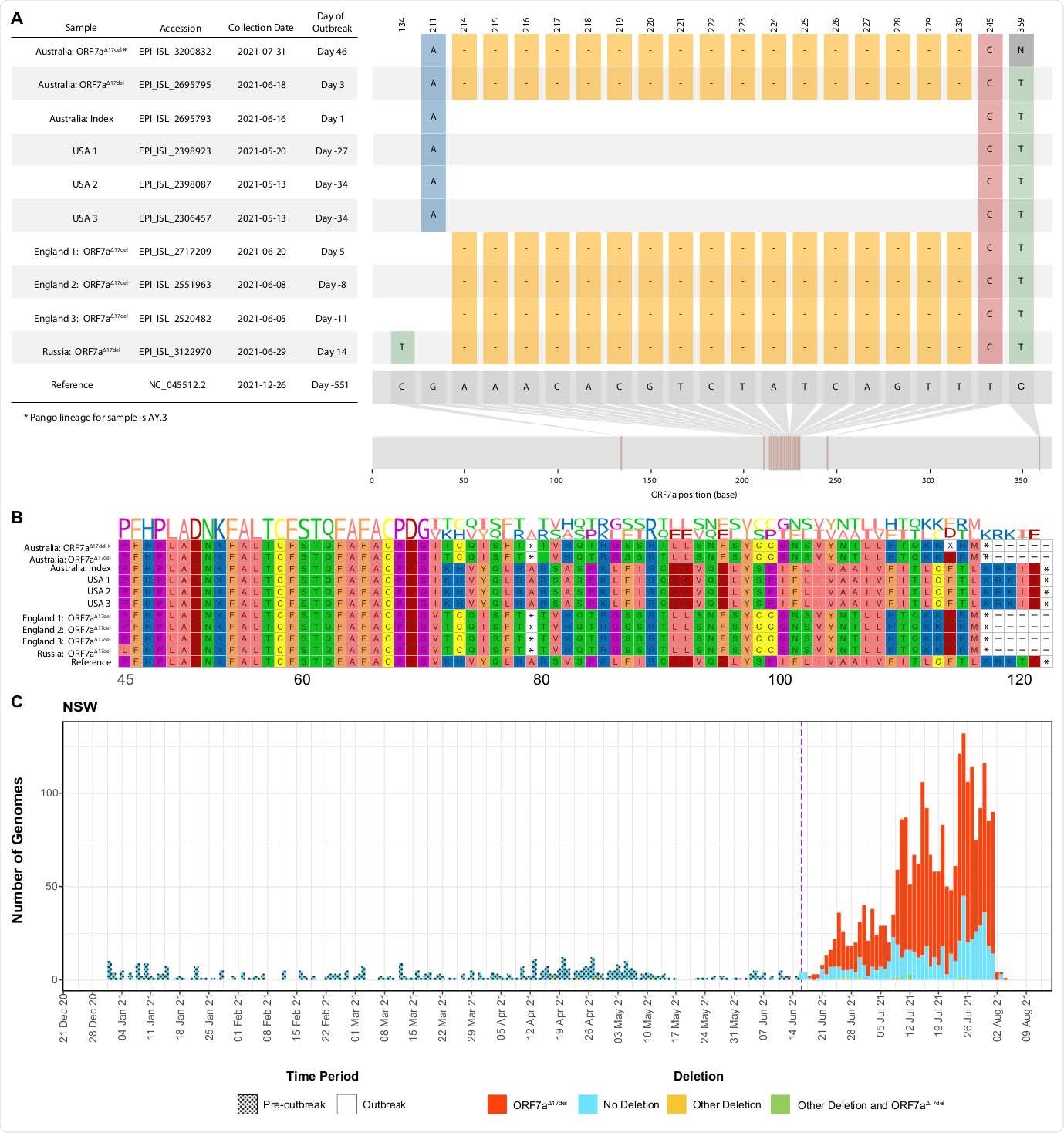
Visualization of a 17-nucleotide deletion in ORF7a of Delta variant SARS-CoV-2 sequences (ORF7a Δ17del) from Australia, 2021, the possible functional consequences at the peptide level, and the increasing frequency in NSW, Australia. Sequences chosen for visualization include the index case of the current outbreaks (lacking ORF7aΔ17del), the first Australian case possessing ORF7aΔ17del, the only sample assigned to lineage AY.3 that possesses ORF7aΔ17del, the three USA samples (all lacking ORF7aΔ17del) that match the Australian index case, and the only four non-Australian samples to possess ORF7aΔ17del. Accession numbers for all samples analyzed for this panel are in the Supplementary Appendix, available with the full text of this letter online. Panel A shows the sample dates of all chosen samples relative to the index case of the current outbreaks (16 June 2021), and all variants in the sample genomes relative to the SARS-CoV- 2 reference genome (created in part using snippet (https://github.com/aineniamh/snipit). Panel B illustrates the consequences of ORF7aΔ17del on the amino acid translation of ORF7a, leading to a premature stop codon after amino acid 78 and a truncated peptide sequence (~64% complete). Amino acids 1–44 are identical to the SARS-CoV-2 reference genome in all samples and are not presented. Panel C demonstrates the number of genomes of all lineages from Australia from 01 January 2021 until 11 August 2021 with ORF7aΔ17del, ORF7aΔ17del plus other deletion(s), other deletions in ORF7a only, or no deletions in ORF7a. The purple dashed line indicates the beginning of the current outbreaks in Australia, with shading also indicating pre-outbreak and outbreak samples.
Study significance
The study reveals a global surge of delta variants with ORF7a 17-nucleotide frameshift deletion mutation that causes the generation of a truncated peptide with different or no activity towards the host restriction factor BST2. Despite the inability to suppress host restrictions, the number of cases with delta ORF7a 17-nucleotide deletion mutation is increasing, indicating that this mutation does not always impair viral fitness.
Collectively, these observations indicate that the ORF7a 17-nucleotide frameshift deletion mutation provides a direct selective advantage by compensating for the loss of BST2 antagonism. However, another reason could be that the mutation has appeared by a chance founder effect, which is defined by a reduction in genetic variation that occurs when a small group of individuals dissociates from a larger population to establish a sub-population.
Given the rapid spread of delta variants worldwide, the scientists suggest that functional studies of variants containing the ORF7a 17-nucleotide deletion mutation are urgently required.

 *Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: medRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.