Since the emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in Wuhan, China, in late December 2019, more than 240 million recorded infections and over 4.89 million deaths have been attributed to this novel betacoronavirus. Globally, the economy has been severely disrupted, with health systems in many countries nearing collapse. Fortunately, before the end of 2020, there were multiple vaccines that display high efficacy approved for use, which includes the Moderna and Pfizer/BioNTech messenger RNA (mRNA) -based vaccines, which provide 90% protection against the disease.
The D614G variant of SARS-CoV-2 was the first variant that had a survival advantage, i.e., it was more transmissible than the original strain. This variant did not evade immunity and appeared to be more sensitive to neutralizing antibodies. However, more worrisome was the emergence of rapidly spreading variants towards the end of 2020, including B.1.351, P.1, B.1.1.7, and B.1.427/B.1.429.
To address the gaps in the knowledge, a team of researchers from the University of California and the Gladstone Institute of Virology conducted 39-parameter phenotyping by cytometry by time of flight (CyTOF) on 33 longitudinal specimens from 11 mRNA-vaccinated individuals, of which six had previously been infected and recovered from COVID-19.
This study is available as an accepted manuscript on the elifesciences.org website.
.jpg)
Study:
mRNA vaccine-induced
T cells respond identically to SARS-CoV-2 variants of concern but differ in longevity and homing properties depending on prior infection status. Image Credit: Fusebulb / Shutterstock
Can SARS-CoV-2-specific T cells elicited by vaccination recognize B.1.351 and B.1.1.7 variants?
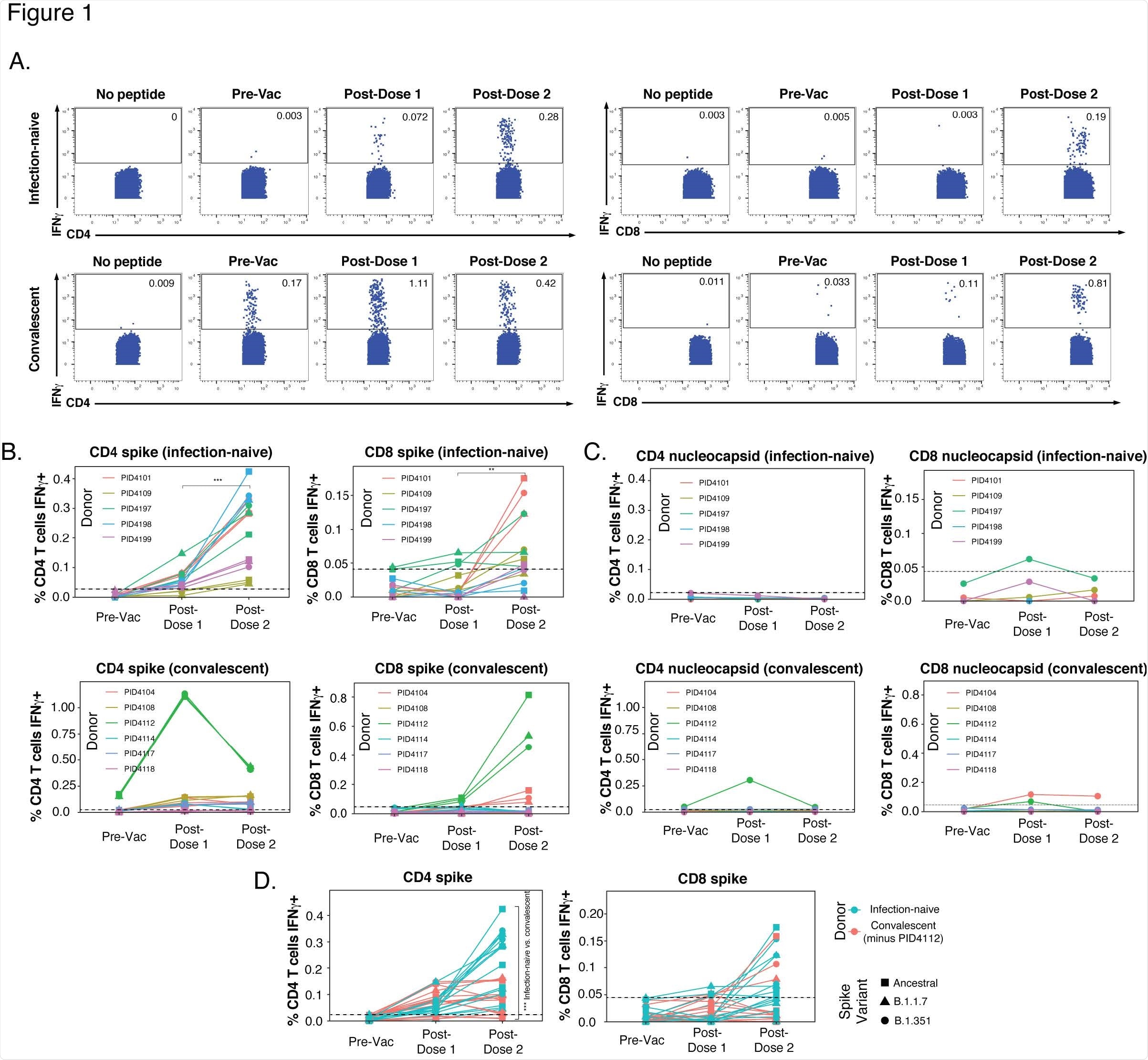
Within the participants who were infection-naïve, a spike-specific CD4+ T cell response was primed following the first vaccination, which was boosted with the second dose. Exposure to ancestral and variant spikes stimulated similar numbers of cells for each participant, suggesting ancestral and variant spikes are recognized equally by vaccine-elicited spike-specific CD4+ T cells.
A weaker response was observed in vaccine-elicited CD8+ T cells, but slight improvements were seen following the second dose.
In comparison to the infection-naïve participants who displayed spike-specific CD4+ T cells of which were boosted following the second dose, previously infected individuals did not show a consistent upward trend. One previously infected participant had a significant frequency of pre-vaccination SARS-CoV-2-specific CD4+ T cells that only increased >1% of the total CD4+ T cell frequency following the first dose and then dampened after the second dose.
The results of the levels of spike-specific T cell response together suggest that the second dose of vaccine boosts the CD4+ T cell response in infection naïve individuals, individuals with previous infections show a different response which is mainly weaker than in infection naïve individuals, and CD4+ T cells exhibit a more robust response when compared to CD8+ T cells.
Six-hour stimulation with spike peptides does not induce significant expression of IL4, IL17, or activation markers in SARS-CoV-2-specific T cells. (A, B) CD4+ T cells were assessed for expression of the Th2 cytokine IL4 (A) or the Th17 cytokine IL17 (B) following 6 hours of stimulation with ancestral spike peptides using PBMC specimens from a representative infection-naïve individual (PID4197) before vaccination (Pre- Vac), or two weeks after dose 1 or dose 2 of vaccination. (C) CD4+ T cells were assessed for co-expression of the activation-induced markers (AIM) Ox40 and 4-1BB following 6 hours of stimulation, using the same specimens as panel A. (D) CD8+ T cells were assessed for co831 expression of the AIM CD69 and 4-1BB following 6 hours of stimulation, using the same specimens as panel A. Baseline specimens not treated with peptide are shown as a comparison control. Numbers correspond to percentages of cells within the gates. Note that the activated (AIM+) cells that appear in stimulated specimens probably do not reflect peptide-specific stimulation as AIM+ cells are also detected in the baseline specimens.
Vaccination-induced spike-specific CD4+ T cells from previously infected individuals exhibit 257 unique phenotypic features of increased longevity and tissue homing
When the results from a previously infected individual who was an outlier were removed, the magnitude of the spike-specific CD4+ T cell response appeared to be lower in previously infected individuals when compared to infection-naïve participants following complete vaccination. Although, there were no significant differences detected when all the participants were included. However, phenotypic differences were observed in the spike-specific CD4+ T cells from the previously infected and infection-naïve.
Differences in CD4+ T cells were much more pronounced following the second dose of vaccination. Previously infected fully vaccinated individuals appeared to have more spike-specific Tcm and Tn cells and less spike-specific Ttm cells. In contrast, there was no difference observed in infection-naïve and previously infected individuals regarding the frequency of Tfh and Treg cells.
The authors then assessed the potential longevity of the spike-specific CD4+ T cells by determining the percentage of CD127+ cells that express low levels of the terminal differentiation marker CD57. In comparison to infection-naïve individuals, previously infected individuals had more long-lived CD127+CD57-spike-specific CD4+ T cells.
The marker CXCR4 was the second most expressed among the spike-specific CD4+ T cells of previously infected individuals and was recently reported to direct bystander T cells to the lungs during an active SARS-CoV-2 infection and has been shown to be co-expressed with the T resident memory/activation marker CD69. In addition, the spike-specific CD4+ T cells from previously infected individuals displayed highly elevated CXCR4+CD69+ cells, suggesting an increased ability to enter pulmonary tissue.
Implications
T cells are extremely essential orchestrators and effectors vital for an efficient immune response. Due to T cell longevity, these cells may hold the key to long-term immunity to viruses such as SARS-CoV-2. The authors in this study designed a longitudinal study that assessed the frequency and phenotypic characteristics of SARS-CoV-2-specific T cells.
If current strategies to vaccination do not result in robust cellular and humoral responses, then strategies that elicit mucosal-homing SARS-CoV-2-specific B and T cells may function greater at achieving desired immunity. Therefore, due to cellular and humoral responses likely being the key to controlling viral replication, future studies should consider the extent to which the breadth, isotypes, and functional features of spike-specific antibodies elicited by vaccination are associated with the phenotypic characteristics of vaccine-elicited SARS-CoV-2-specific T cells.
Journal reference:
- mRNA vaccine-induced T cells respond identically to SARS-CoV-2 variants of concern 2 but differ in longevity and homing properties depending on prior infection status, Jason Neidleman, Xiaoyu Luo1, Matthew McGregor, Guorui Xie, Victoria Murray, Warner C. Greene, Sulggi A. Lee, and Nadia R. Roan, eLife, 2021.10.12, https://elifesciences.org/articles/72619