Scientists stated the importance of understanding various aspects of the severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), the causal agent of the ongoing coronavirus disease 2019 (COVID-19) pandemic. Some factors, such as vaccine hesitancy, shortage of vaccines in many parts of the world, and the continual emergence of new SARS-CoV-2 variants, have prolonged the pandemic.
 Study: Numb-associated kinases are required for SARS-CoV-2 infection and are cellular targets for therapy. Image Credit: NIAID
Study: Numb-associated kinases are required for SARS-CoV-2 infection and are cellular targets for therapy. Image Credit: NIAID

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Background
Scientists have developed some direct-acting antivirals (DAAs), e.g., nirmatrelvir (protease inhibitor) and molnupiravir (polymerase inhibitor), for the treatment of severely infected COVID-19 patients. However, extensive use of DAAs could lead to the emergence of resistant viral variants. Hence, there is a need for new host-targeted antiviral drugs that could suppress viral replication with a high barrier to resistance.
Previous studies have revealed that SARS-CoV-2 infection is mediated by the binding of the spike protein of the virus with angiotensin-converting enzyme 2 (ACE2) of the host. Some of the factors associated with viral infection are neuropilin-1 (NRP-1) and cellular transmembrane protease serine 2 (TMPRSS2). After the virus gains entry to the targeted host cell, the virions assemble in the endoplasmic reticulum-Golgi intermediate compartment (ERGIC) and egress through the secretory pathway.
The Numb-associated kinases (NAK) family of Ser/Thr kinases contains adaptor-associated kinase 1 (AAK1), BMP-2 inducible kinase (BIKE/BMP2K), cyclin G-associated kinase (GAK), and serine/threonine kinase 16 (STK16). Interestingly, these kinases do not share many similarities in the kinase domain. One of the important functions of these kinases is the regulation of intracellular membrane trafficking. AAK1 and GAK are associated with phosphorylation of endocytic adaptor protein complex 2 (AP2M1) and secretory AP complex 1 (AP1M1), which enhances binding of the virus to the host. Previous studies have also reported that BIKE is an accessory protein that binds the endocytic adaptor NUMB and phosphorylates AP2M1. The primary function of STK16 is that it regulates various physiological activities, such as Golgi assembly, trans-Golgi network (TGN) protein secretion, and TGF-beta signaling.
A New Study
Previous studies have indicated the role of AAK1 and GAK in enabling intracellular trafficking of viruses, such as hepatitis C virus (HCV), dengue virus (DENV), and Ebola virus (EBOV), during the entry and assembly phases. Based on in silico studies, AAK1 was proposed as a cellular target of the SARS-CoV-2 virus. However, there is a gap in research regarding NAKs as potential targets for anti-SARS-CoV-2 therapy.
A new study posted to the bioRxiv* preprint server has focussed on assessing four members of the NAK family that are associated with SARS-CoV-2 infection. These kininases could act as a target for the development of effective COVID-19 therapeutics.
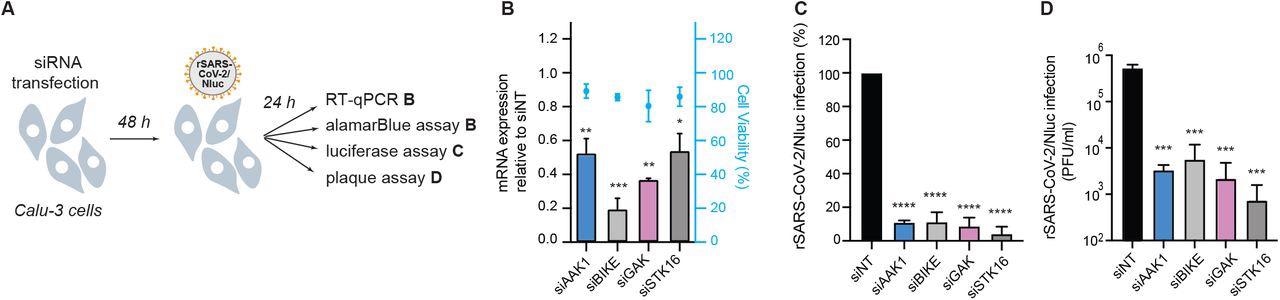
NAKs are required for SARS-CoV-2 infection. (A) Schematic of the experiments shown in panels B-D. (B) Confirmation of siRNA-mediated gene expression knockdown and cell viability in Calu-3 cells transfected with the indicated siRNAs. Shown is gene expression normalized to GAPDH and expressed relative to the respective gene level in the non-target (siNT) control at 48 hours post-transfection measured by RT-qPCR and cell viability measured by alamarBlue assay. (C) WT SARS-CoV-2 infection measured at 24 hours post-infection (hpi) of the indicated NAK-depleted Calu-3 cells with rSARS-CoV-2/Nluc (USA-WA1/2020 strain; MOI=0.05) by Nluc assays. (D) Viral titers measured at 24 hpi of the indicated NAK-depleted Calu-3 cells with rSARS-CoV-2/Nluc (USA-WA1/2020 strain; MOI=0.05) by plaque assays. Data in all panels are representative of 2 or more independent experiments. Individual experiments had 3 biological replicates, means ± standard deviation (SD) are shown. Data are relative to siNT (C, D). *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001 by 1-way ANOVA followed by Dunnett’s multiple comparisons test. PFU, plaque-forming units.
Key Findings
In this study, researchers integrated virological, genetic and pharmacological approaches to investigate the role of four NAKs in SARS-CoV-2 infection. Additionally, the authors provided a proof of concept to show that NAK inhibitors could be used to treat SARS-CoV-2 infection. Previous studies have used artificial intelligence to identify AAK1 as a target candidate for SARS-CoV-2 treatment. The researchers have used the RNAi approach to show that AAK1 is essential for SARS-CoV-2 infection. Additionally, scientists provided genetic evidence to prove that BIKE, GAK and STK16 are proviral factors of SARS-CoV-2 infection.
This study provided evidence that NAKs regulate SARS-CoV-2 entry. A decreased level of NAKs was seen to reduce the entry of pseudo- and wild-type SARS-CoV-2 through Nluc assays and other specific assays related to measuring viral RNA at 2 hpi. Interestingly, time-of-addition experiments showed that pharmacological inhibition of AAK1/BIKE potently inhibited viral infection. A previous study showed that the conserved tyrosine motif in the cytoplasmic tail of ACE2, mediated interaction with AP2M1, which is essential for SARS-CoV-2 infection when the level of TMPRSS2 is low. The current study revealed that in high TMPRSS2 expressing Calu-3 cells, NAKs are essential for the entry of SARS-CoV-2. Additionally, researchers also observed that NAKs are involved with regulating later stages of the SARS-CoV-2 life cycle (e.g., assembly and/or egress) but not RNA replication.
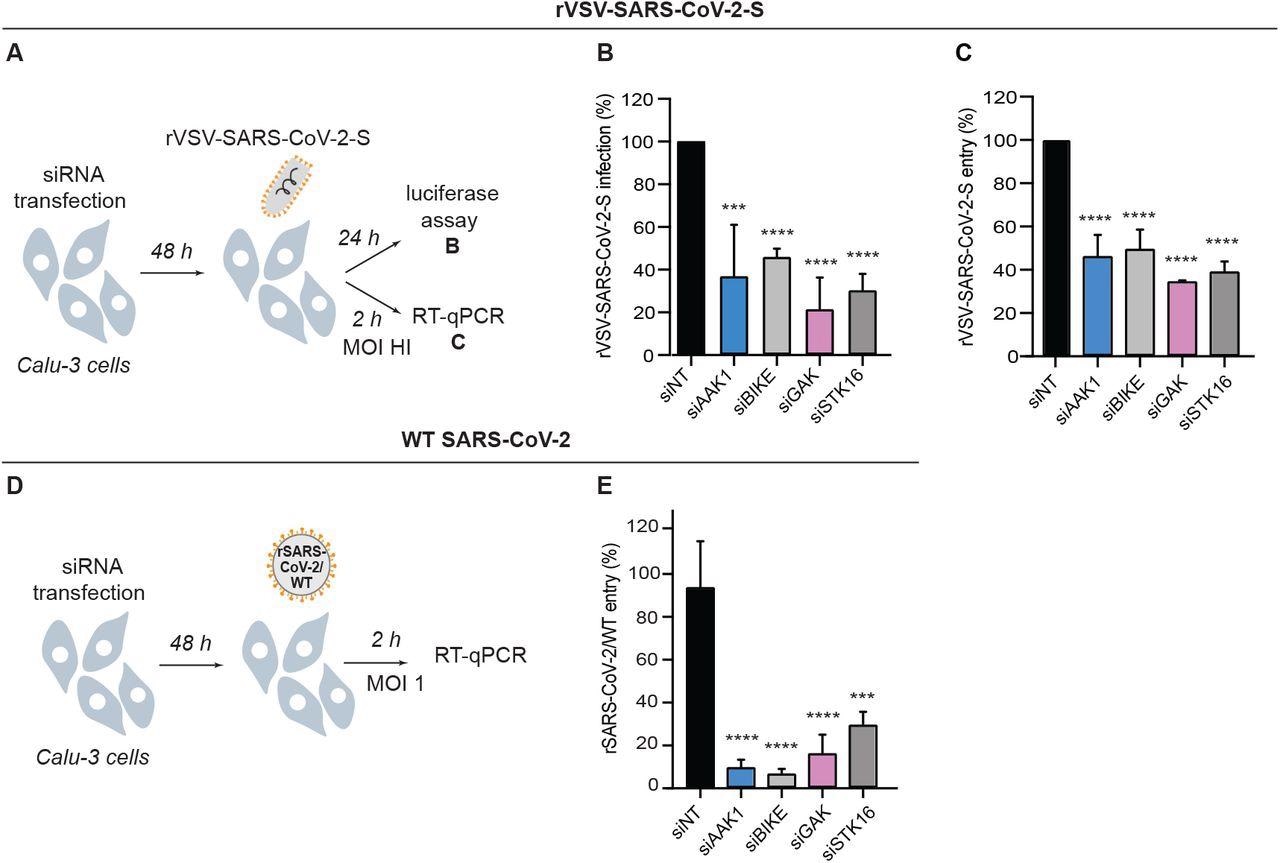
NAKs are required for the entry of pseudovirus and WT SARS-CoV-2. (A) Schematic of the experiments shown in panels B and C. (B) rVSV-SARS-CoV-2-S infection at 24 hpi of Calu-3 cells depleted of the indicated NAKs-by siRNAs (Figure 1B) measured via luciferase assays. (C) rVSV-SARS-CoV-2-S entry at 2 hpi of Calu-3 cells (high MOI) depleted of the indicated NAKs by siRNAs measured via RT-qPCR. (D) Schematic of the experiment shown in panel E. (E) rSARS-CoV-2/WT entry at 2 hpi of Calu-3 cells (MOI=1) depleted of the indicated NAKs by siRNAs measured via RT-qPCR. Data in all panels are representative of 2 or more independent experiments. Individual experiments had 3 biological replicates, means ± SD are shown. Data are relative to siNT (B, C, E). ***P < 0.001; ****P < 0.0001 by 1-way ANOVA followed by Dunnett’s multiple comparisons test.
In this study, researchers provided robust evidence to show that NAKs can serve as a potential target for the development of anti-SARS-CoV-2 therapy. The finding of this study is in line with previous studies that revealed AAK1 inhibitors could effectively suppress SARS-CoV-2 infection. Gefitinib, RMC-242, and SGC-GAK-1 are NAK inhibitors that exhibited moderate antiviral activity against SARS4-CoV-2. Interestingly, the STK16 inhibitor also revealed the anti-SARS-CoV-2 effect. Scientists used artificial intelligence to identify baricitinib, a NAK inhibitor, which displayed an anti-inflammatory effect. The authors also stated that combining compounds with anti-AAK1/BIKE and anti-GAK activities could provide a synergistic antiviral effect against SARS-CoV-2.
Conclusion
The current study provided robust evidence that showed NAKs are a potential target for the development of antiviral therapy for the treatment of SARS-CoV-2 infection. They can also be used for developing treatments of other coronavirus infections.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Karim, M. et al. (2022) Numb-associated kinases are required for SARS-CoV-2 infection and are cellular targets for therapy, bioRxiv, 2022.03.18.484178; doi: https://doi.org/10.1101/2022.03.18.484178, https://www.biorxiv.org/content/10.1101/2022.03.18.484178v1
- Peer reviewed and published scientific report.
Karim, Marwah, Sirle Saul, Luca Ghita, Malaya Kumar Sahoo, Chengjin Ye, Nishank Bhalla, Chieh-Wen Lo, et al. 2022. “Numb-Associated Kinases Are Required for SARS-CoV-2 Infection and Are Cellular Targets for Antiviral Strategies.” Antiviral Research 204 (August): 105367. https://doi.org/10.1016/j.antiviral.2022.105367. https://www.sciencedirect.com/science/article/pii/S016635422200136X.