The coronavirus disease 2019 (COVID-19) pandemic, caused by SARS-CoV-2, has presented unprecedented problems to international societies. To date, the SARS-CoV-2 pandemic has caused over 513 million cases and more than 6.24 million mortalities worldwide. COVID-19 has a wide range of clinical manifestations and illness courses, ranging from asymptomatic (AS) SARS-CoV-2 infection to a critical or severe condition. Notably, the underpinnings for short-term nucleic acid test positive (STNP) and long-term nucleic acid test positive (LTNP) in SARS-CoV-2 infection are not known.
Additionally, miRNAs are short single-stranded RNA compounds harboring a genome of 18 to 28 nucleotides that have been linked to several pathological and physiological mechanisms in animals and humans, like immunological responses, development, apoptosis, and inflammation. Interestingly, although miRNAs have been demonstrated to play significant roles in viral infections, their relationships with COVID-19 are unknown.
About the study
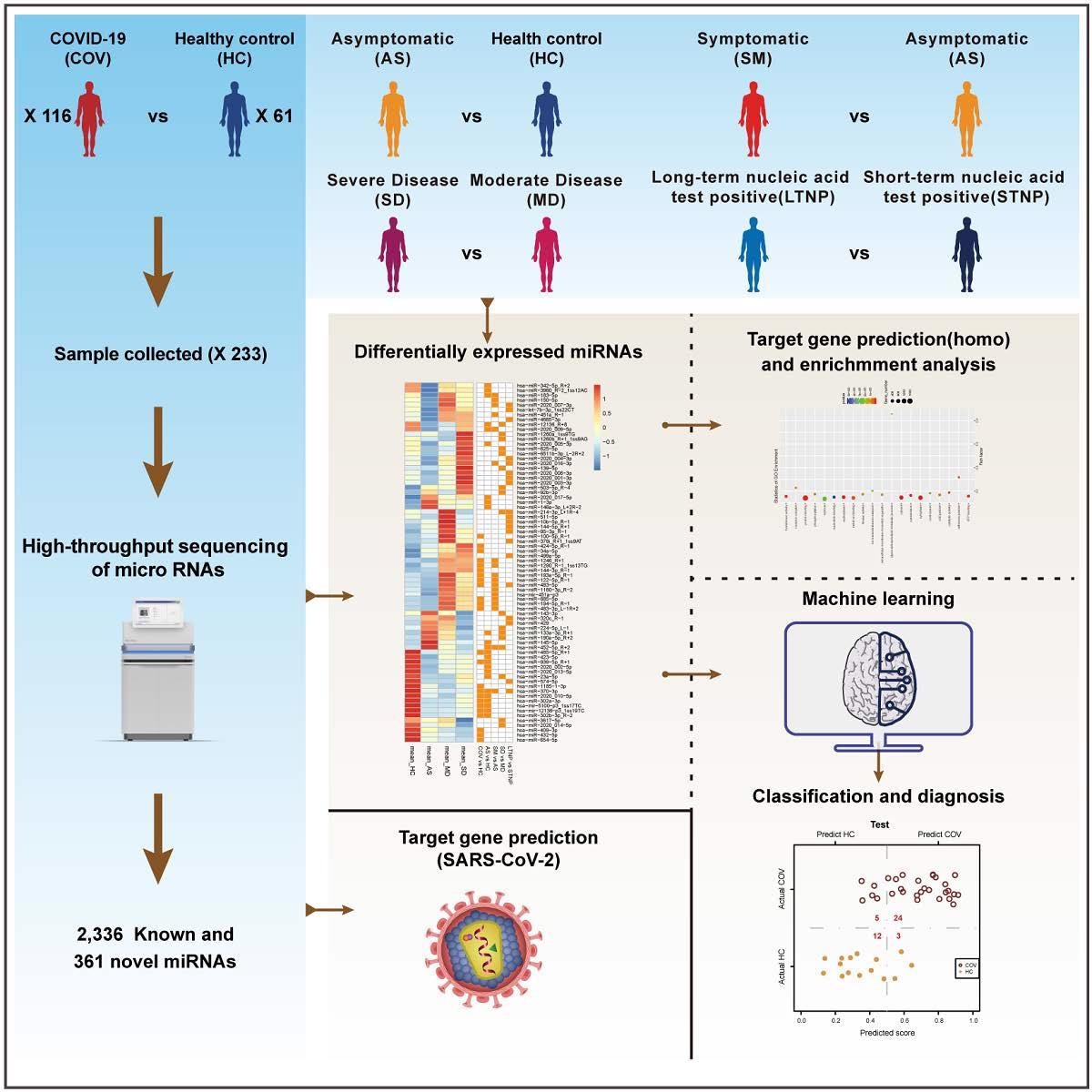
In the present work, the researchers investigated the possible roles of miRNAs in the pathogenesis of SARS-CoV-2 infection by performing a high throughput analysis of the plasma miRNA library. The team developed and evaluated a machine learning model based on differentially expressed miRNAs (DE-miRNAs) between several compared cohorts to categorize COVID-19 patients with distinct clinical manifestations.
The authors presented the heatmap of 85 DE-miRNAs derived by comparing 11 groups, including SARS-CoV-2-infected, healthy controls, AS, severe disease (SD), and moderate disease (MD) COVID-19. Further, DE-miRNAs associated with various severity of COVID-19 and SARS-CoV-2 persistence were determined. The researchers also discovered miRNAs addressing cellular genes linked with the SARS-CoV-2 lifecycle and viral genome. Moreover, they elucidated the possible functions of the representative DE-miRNAs in COVID-19 pathogenesis.
Results
The study results discovered 361 new and 2,336 established miRNAs in 233 plasma samples from 116 SARS-CoV-2 patients and 61 healthy controls employing high throughput sequencing test. The authors found 85 DE-miRNAs among the currently identified miRNAs. In the COVID-19 patients, these DE-miRNAs like homo sapiens (hsa)-main immunogenic region (miR)-370-3p, hsa-miR-885-5, hsa-miR-146a-3p, hsa-miR-10b-5p, and hsa-miR-214-3p were linked to SARS-CoV-2 infection, viral persistence, and illness severity.
Additionally, a panel of miRNAs was identified that could directly attack SARS-CoV-2 viral genes. Moreover, the scientists discovered several miRNAs that could address the cellular genes implicated in the life cycle of SARS-CoV-2, such as angiotensin-converting enzyme 2 (ACE2), translocase of outer mitochondrial membrane 70 (TOMM70), AXL, and transmembrane protein 106B (TMEM106B). The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway and gene ontology analysis demonstrated that DE-miRNAs were correlated with viral infections, lung diseases, and immunological responses.
The authors tested their machine learning technique for the categorization of COVID-19 patients with diverse clinical presentations based on DE-miRNAs among different groups. Further, the results depicted six DE-miRNAs, including hsa-miR-302a-3p and hsa-miR-302b-3p_R-2, that might be utilized as biomarkers to differentiate the COVID-19 patients from the healthy controls. Additionally, previous studies have correlated four out of the six biomarkers with inflammation, virus infection, lung disease, and immune response.
According to the present data, miR-302 might protect COVID-19 patients by addressing the C-X-C motif chemokine ligand 8 (CXCL8)-linked pathways. Further, hsa-miR-146a-3p might have a protective function in AS COVID-19 patients and could be a biological marker for therapeutic and diagnostic approaches aiming at this category. The authors also discovered numerous De-miRNAs, such as hsa-miR-122-5p and hsa-miR-1246, that could distinguish AS and symptomatic (SM) SARS-CoV-2 patients.
Additionally, in the LTNP versus STNP groups, the researchers found 20 DE-miRNAs, nine of which, including hsa-miR-429 and hsa-miR-483-5p, were substantially linked with the length of viral persistence in SARS-CoV-2 patients.
Conclusions
According to the authors, this was the initial large-scale analysis of miRNA characteristics using high throughput investigation on the plasma samples procured from SARS-CoV-2 AS volunteers in tandem with the SM patients harboring different clinical presentations and the healthy control subjects.
Overall, the present study discovered a multitude of DE-miRNAs linked to SARS-CoV-2 infection, disease severity, viral persistence, and clinical symptoms in COVID-19 patients. It also recognized a screen of miRNAs possibly addressing the viral genomic regions or host genes implicated in a range of pathways of SARS-CoV-2. The authors mentioned that the current findings should help researchers better understand how miRNAs play a role in the pathophysiology of SARS-CoV-2 infection and uncover possible molecular targets and biomarkers for COVID-19 therapy and diagnostics.
Journal reference:
- Zeng, Q., Qi, X., Ma, J., Hu, F., Wang, X., Qin, H., Li, M., Huang, S., Yang, Y., Li, Y., Bai, H., Jiang, M., Ren, D., Kang, Y., Zhao, Y., Chen, X., Ding, X., Ye, D., Wang, Y., Jiang, J., Li, D., Chen, X., Hu, K., Zhang, B., Shi, B., Zhang, C., Distinct miRNAs associated with various clinical presentations of SARS-CoV-2 infection, ISCIENCE (2022). DOI: https://doi.org/10.1016/j.isci.2022.104309, https://linkinghub.elsevier.com/retrieve/pii/S2589004222005806