The immune reaction to viral infection is influenced significantly by NK cells, which are innate lymphocytes. These cytotoxic effector cells react quickly to viral infection via addressing and lysing infected cells.
Studies evaluating the immune reaction in CoV disease 2019 (COVID-19) during the SARS-CoV-2 pandemic have found that NK cells are less prevalent in the peripheral blood of severe SARS-CoV-2 patients versus healthy donors. In addition, immune profiling has revealed substantial, severity-related transcriptional and phenotypic alterations in the peripheral NK cells that persist in SARS-CoV-2 patients' blood.
Though multiple studies have shown NK cells can inhibit SARS-CoV-2 replication in vitro, the mechanism via which NK cells react immediately to SARS-CoV-2-infected cells remains uncertain. This is especially crucial because several viruses adopt techniques to avoid being detected and eliminated by NK cells.
 Study: SARS-CoV-2 escapes direct NK cell killing through Nsp1-mediated downregulation of ligands for NKG2D
Study: SARS-CoV-2 escapes direct NK cell killing through Nsp1-mediated downregulation of ligands for NKG2D

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
About the study
The present research aimed to assess the capacity of SARS-CoV-2 to adjust NK cell identification and lysis of virus-infected cells. The team developed an in vitro model approach that analyzes the NK cell reaction to SARS-CoV-2-infected cells using primary NK cells derived from healthy donors and replication-competent SARS-CoV-2. To further elucidate how the balance between SARS-CoV-2 detection and escape leads to COVID-19, they concentrated on examining the direct killing of SARS-CoV-2-infected target cells by the NK cells.
Adenocarcinomic human alveolar basal epithelial (A549)-angiotensin-converting enzyme 2 (ACE2) cells, identified and lysed by NK cells and are SARS-CoV-2 infectible, were used as a model by the authors to study the NK cell reaction to COVID-19. They introduced NK cells obtained from healthy donors preactivated overnight with interleukin 2 (IL-2) to address cells infected with SARS-CoV-2 for 48 hours to comprehend how exposure to SARS-CoV-2-infected target cells affects NK cell activity and phenotype.
The researcher evaluated whether SARS-CoV-2 modifies NK cells' capacity to annihilate infected target cells. Further, they explored how SARS-CoV-2-infected cells managed to escape being lysed and recognized by NK cells.
The authors examined NK group 2D-ligand (NKG2D-L) expression on the cells that persisted after being co-cultured with NK cells to investigate the relationship between NKG2D-L expression and the killing of SARS-CoV-2-infected cells. The team then tried to figure out how SARS-CoV-2 affects NKG2D-L protein expression across SARS-CoV-2-infected cells.
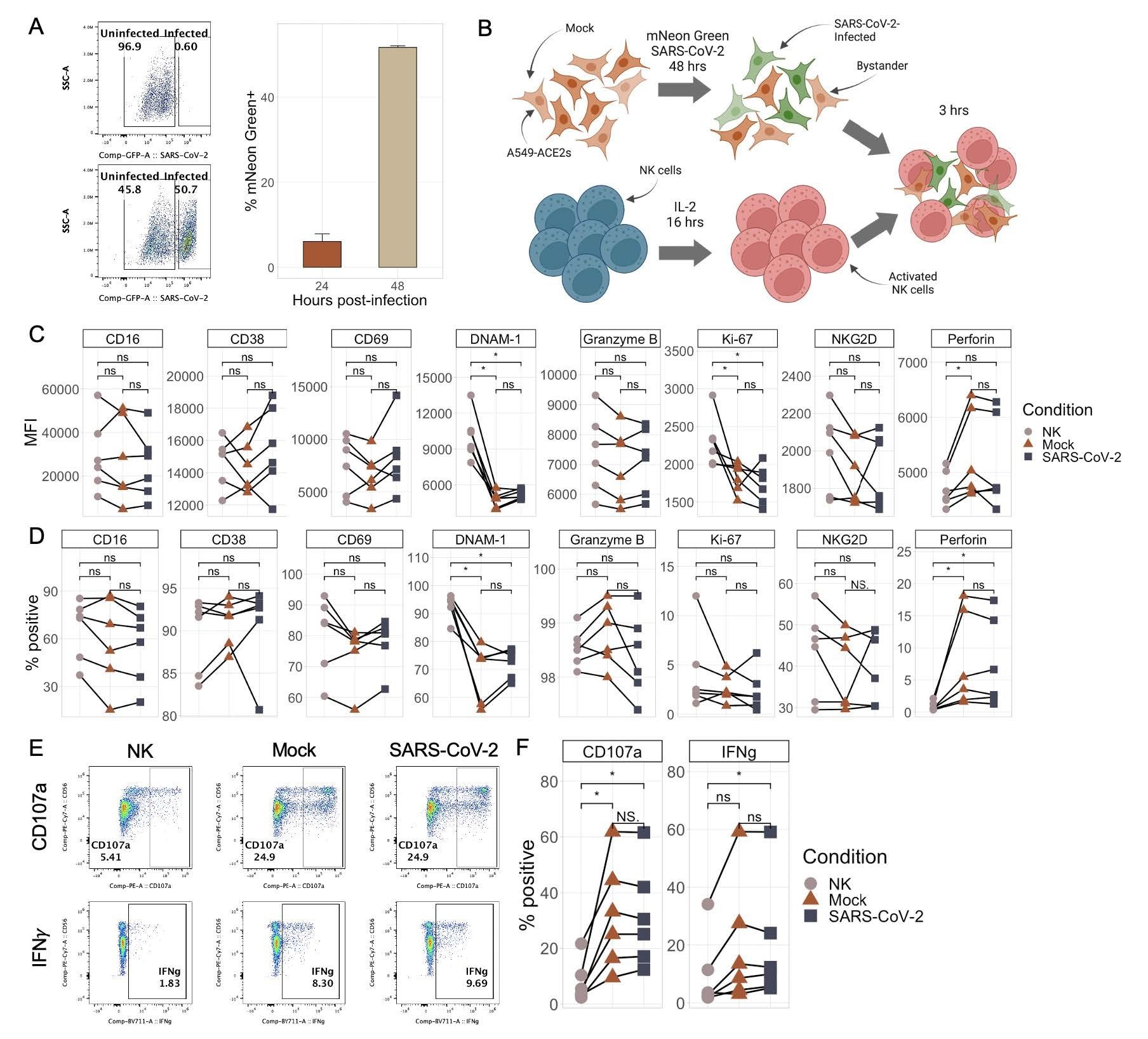 NK cells respond similarly to SARS-CoV-2-infected and mock-infected target cells. A) Representative flow plots (left) and boxplot (right) showing the percentage of mNeonGreen-positive A549-ACE2 cells following infection with either mNeon Green SARS-CoV-2 (MOI 0.5) or media (“mock”) at an MOI of 0.5 for either 24 or 48 hours. Bar plots represent n=4 technical replicates ∓ SD values. B) Schematic illustrating the experimental design of NK cell killing assays. C-D) Plots showing the median fluorescence intensity (C) and % of NK cells positive (D) for eight different NK cell markers by flow cytometry upon culture with no targets, mock-infected targets, or SARS-CoV-2-infected targets. E-F) Representative flow plots (E) and quantitations (F) of percentage of NK cells expressing CD107a and IFNγ upon culture with no targets, mock-infected targets, or SARS-CoV-2-infected targets. Significance values were determined using a paired Wilcoxon ranked-sum test with the Bonferroni correction for multiple hypothesis testing.
NK cells respond similarly to SARS-CoV-2-infected and mock-infected target cells. A) Representative flow plots (left) and boxplot (right) showing the percentage of mNeonGreen-positive A549-ACE2 cells following infection with either mNeon Green SARS-CoV-2 (MOI 0.5) or media (“mock”) at an MOI of 0.5 for either 24 or 48 hours. Bar plots represent n=4 technical replicates ∓ SD values. B) Schematic illustrating the experimental design of NK cell killing assays. C-D) Plots showing the median fluorescence intensity (C) and % of NK cells positive (D) for eight different NK cell markers by flow cytometry upon culture with no targets, mock-infected targets, or SARS-CoV-2-infected targets. E-F) Representative flow plots (E) and quantitations (F) of percentage of NK cells expressing CD107a and IFNγ upon culture with no targets, mock-infected targets, or SARS-CoV-2-infected targets. Significance values were determined using a paired Wilcoxon ranked-sum test with the Bonferroni correction for multiple hypothesis testing.
Results
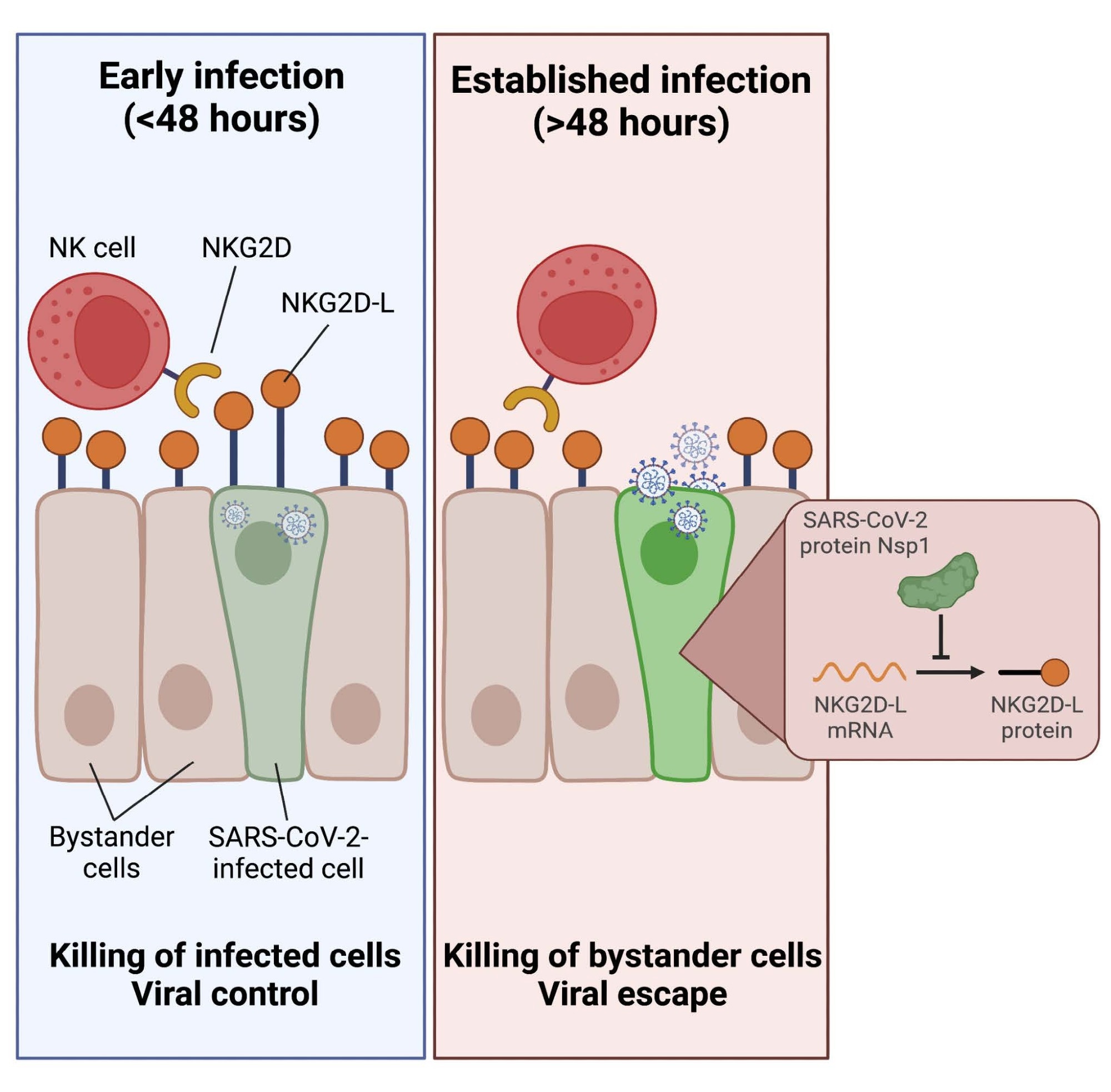
Overall, the study results depicted that NK cells have weak cytotoxic reactions towards SARS-CoV-2-infected targets, selectively killing non-infected bystander cells. Furthermore, the team showed that the SARS-CoV-2-infected cells' significant downregulation of ligands for the stimulating NKG2D receptor was what caused this escape from NK cell-mediated killing.
Indeed, NK cells may efficiently identify and kill infected cells in the early phases of SARS-CoV-2 infection before NKG2D-L downregulation. Nevertheless, the NK cells lose this ability following the expression of viral proteins inside infected cells. The current data show that, when introduced to the culture at later timestamps following SARS-CoV-2 infection, post-expression of viral proteins that dampen the innate immune response, NK cells cannot kill infected cells effectively. Since a small window exists for the elimination of the infected cells before bystander cell killing might occur, the investigators noted that the timing of NK cell migration to the infection site could be crucial in deciding whether NK cells were pathogenic or protective in COVID-19.
Finally, the authors discovered that SARS-CoV-2 non-structural protein 1 (Nsp1) causes the downregulation of NKG2D-L. They also illustrated that transfection with just Nsp1 was enough to impart tolerance to NK cell killing. This observation has crucial implications for NK cell-facilitated regulation of SARS-CoV-2 since the preferred evasion of infected cells with the killing of bystander cells might lead to SARS-CoV-2 pathogenesis.
Conclusions
Collectively, the current investigation deeply examines the NK cell reaction to SARS-CoV-2 and offers new information about the function of NK cells in COVID-19. The researchers discovered that SARS-CoV-2-infected cells evade being destroyed by healthy NK cells in a cell-intrinsic way, leading to the preferential destruction of non-infected bystander cells. The capability of infected cells to escape NK cell identification needs infection to last long enough to facilitate an infected cell to exhibit SARS-CoV-2 encoded proteins. Besides, the present results highlight the significance of studying the temporal dynamics of the NK cell reaction to SARS-CoV-2-infected cells.
Additionally, the team showed that the downregulation of NKG2D-L drives the process of NK cell recognition evasion of SARS-CoV-2. The primary characteristic in the NK cell response to SARS-CoV-2, according to the findings, was the loss of NKG2D-L. The study data further demonstrated that the SARS-CoV-2 Nsp1 protein was responsible for this ligand downregulation and that Nsp1 itself was adequate to induce direct NK cell escape. The current work depicted that NK cell responses to SARS-CoV-2-infected cells might be partially or fully rescued by reducing the activity of Nsp1, proving that this protein was an even more desirable target than previously believed.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
SARS-CoV-2 escapes direct NK cell killing through Nsp1-mediated downregulation of ligands for NKG2D; Madeline J Lee, Michelle W Leong, Arjun Rustagi, Aimee Beck, Leiping Zeng, Susan Holmes, Lei S Qi, Catherine A Blish. bioRxiv preprint 2022, DOI: https://doi.org/10.1101/2022.06.20.496341, https://www.biorxiv.org/content/10.1101/2022.06.20.496341v1
- Peer reviewed and published scientific report.
Lee, Madeline J., Michelle W. Leong, Arjun Rustagi, Aimee Beck, Leiping Zeng, Susan Holmes, Lei S. Qi, and Catherine A. Blish. 2022. “SARS-CoV-2 Escapes Direct NK Cell Killing through Nsp1-Mediated Downregulation of Ligands for NKG2D.” Cell Reports, December, 111892. https://doi.org/10.1016/j.celrep.2022.111892. https://www.cell.com/cell-reports/fulltext/S2211-1247(22)01791-0.