Parallel line analysis (PLA) is often used to analyze biological assays in laboratories that follow GMP (good manufacturing practice) and GLP (good laboratory practice) regulations. This method is typically employed for the comparison of dose-response curves where a direct measurement of a product is not available, but rather, an effect is measured (Figure 1).
Using parallelism methods, users are able to discern whether there is a similarity in the biological response of two substances or if two biological environments influence the dose-response curves to the same substances.
Testing for parallelism is necessary when calculating the relative potency of a compound and takes up a central role in several pharmaceutical drug development applications, such as analyte confirmation, concentration estimation, cross-reactivity, drug comparison, inhibitory studies, interfering substances, and matrix compensation.
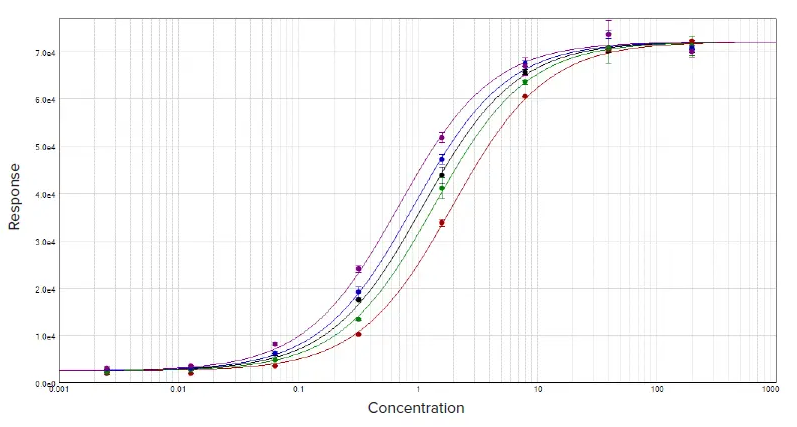
Figure 1. Parallel line analysis of dose response data sets with a constrained global 4-parameter curve fit. Image Credit: Molecular Devices UK Ltd
Two curves are determined to be parallel when one function can be obtained from the other using a scaling factor either to the right or to the left on the x-axis. ƒ(x) = ƒ(rx), where x is the dose and r is the scaling factor, or relative potency.1
The setting of the relative potency typically begins at one for the reference curve (known agent), and the scaling factor employed to convert the reference curve into the test curve (unknown agent) becomes the relative potency of the unknown agent.
This approach functions well when applied to linear regression curve fits where the slope remains intact across the concentration range (Figure 2). However, when faced with non-linear regression curve fits, such as the 4-parameter and 5-parameter logistics, the slope of sigmoidal dose-response curve can fluctuate across the entire concentration range (Figure 1).
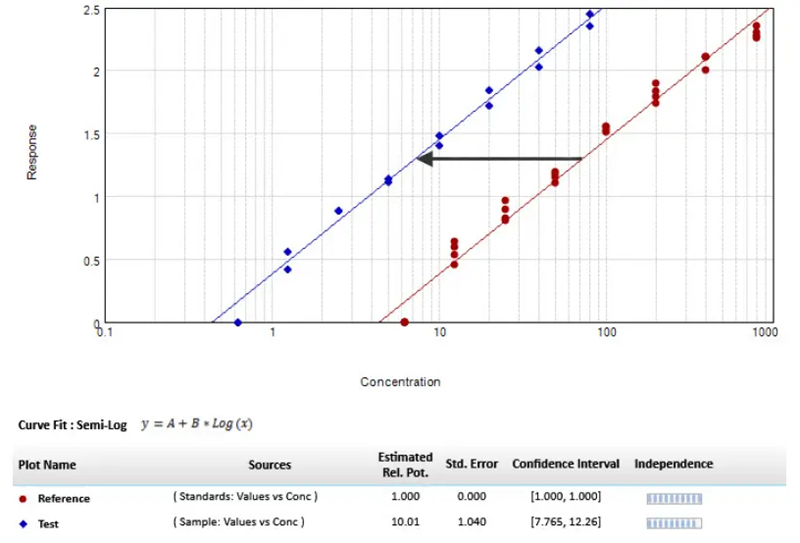
Figure 2. Parallel line model for linear regression. The relative potency is set to one for the reference curve (red circle), and the scaling factor used to transform the reference curve into the test curve (blue diamond) is the relative potency of the unknown agent. Image Credit: Molecular Devices UK Ltd
Testing parallelism methods can be split into two categories, which are determined by how the parallelism hypothesis is tested: response comparison tests and parameter comparison tests.1
This article provides a discussion of both methods and explains how to apply them in both SoftMax® Pro GxP and Standard Software to test for parallelism. A protocol has been performed using the F-test probability with the F-test, alongside an application of the chi-squared probability with the chi-squared test.1,2,3
Moreover, a parameter comparison method was developed using Fieller’s theorem. This protocol, named the Parallelism Test, can be found in the SoftMax Pro Protocol Library in the Data Analysis subfolder. Each method can be applied for both linear and non-linear regression curves.
Testing for parallelism
Response comparison test
Biological systems often behave in unexpected ways and tend to add noise and variation to the data. Therefore, it is crucial to select the correct curve fit model and apply a weighting factor.
Applying a model that has the appropriate accommodation capacity for these variations can be a necessary and important step to consider first before parallelism analysis. If the wrong curve fit model is selected, bias can be inserted into the parallelism metrics and may produce an incorrect conclusion.
It can be difficult to calculate the relative potency of non-parallel curves, due to the rare occurrence of curve fits that are perfectly parallel for assay data, particularly when observing non-linear regressions.
Using the response comparison method, the tested curves can be inserted into a constrained model where the curves are forced to be parallel, or an independent model where the curves are fitted independently.
Statistical metrics are then applied simultaneously to compare differences in how well points fit between the constrained model and the independent model, which might relate to non-parallelism.
When fit to the constrained model, curve parameters are identical across the board, with the exception of the parameter that describes the x-value. For a linear curve, the x-value is the intercept and for a non-linear curve, the x-value is the midpoint between the upper and the lower asymptotes, expressed as EC50.
SoftMax Pro Software is equipped with tools that can identify the relative potency of linear fits together with an evaluation of non-linear curves by applying the constrained or global fit model and the independent model for relative potency estimation.
Testing the null hypothesis
The parallelism metrics are often a function of the residual sum of squared-errors (RSSE) when applying the response comparison method. These metrics determine to what extent the constrained model matches the data.
One particular method applies the Extra-Sum-of-Square analysis.1,2 This is a form of analysis of variance (ANOVA) using a statistical regression technique where the null hypothesis determines that the constrained model is correct, or that the curves are parallel.
To test the null hypothesis, a variety of statistical techniques can be applied, including the F-test probability with the F-test,1,2 or the chi-squared probability with the chi-squared test.3
Across both methods, the probability is noted as a number between zero and one. As the probability moves closer towards one, it is increasingly likely that the curves are parallel.
It should be noted, however, that the F-test presents some limitations. These include false positives for independent curves that exhibit a good fit, or false negatives for independent curves exhibiting a poor fit.
These two statistical methods can be accessed using SoftMax Pro Software. Values of the test and the probabilities can be easily obtained by applying the following formulas:
- ChiSquaredPLA (PlotName@GraphName): Returns the value of the chi-squared statistic for a reduced curve fit
- ChiProbabilityPLA (PlotName@GraphSection): Returns the chi-squared probability distribution value for a reduced curve fit
- FStatPLA(PlotName@GraphSection): Returns the value of the F-test statistic for a reduced curve fit
- FProbPLA(PlotName@GraphSection): Returns the value of the F-test probability for a reduced curve fit
Note: These formulas can only be used when the Global Fit (PLA) option is activated in the Curve Fit Settings dialog.
PlotName@GraphName is used to designate the name of the plot, including the name of the graph. For instance, Plot#1@Graph#1 or Std@Standardcurve.
Since the chi-squared value is calculated using all plots in the designated graph, the designation of a particular plot is considered arbitrary. Thus, a protocol titled ‘Parallel Line Analysis Using F-test and Chi-squared Test’ was developed for testing parallelism in accordance with these two statistical testing methods.
Once data acquisition has been completed or imported into the protocol, the calculations will run automatically to evaluate whether or not the null hypothesis (that the curves are considered parallel) is true.
When using this protocol, the probability results for the F-test and the chi-squared test must exceed 0.05 to determine that the curves are parallel. Generally speaking, when using this setting, users can have 95 % confidence in the fact that the null hypothesis is true. The confidence level can be adjusted as necessary to an appropriate level of non-parallelism for the assay performed.
Noise and weighting
Noise is characterized as the unpredictable variations present in the measured response. It is a vital factor to consider when parallelism is assessed, as it impacts the ability to identify non-parallelism. When encountering high noise levels, parallelism metrics are not suitable for measuring non-parallelism as they are lost in the noise.
The F-test and chi-squared metrics manage the expected noise levels differently when calculating their relative probability. While noise has no bearing on the F-test probability, the chi-squared probability is greatly dependent on noise levels, which means that data variances must be carefully estimated. When applying the chi-squared method, it is necessary to have inverse variance weighting.
As covered in the application note, ‘Selecting the best weighting factor in SoftMax Pro GxP and Standard Software’, bioassays typically exhibit increased variability in the upper echelons of the curve. With unweighted regression, the data expressed in the points from the top of the curve can dominate the parallelism results. This means information at the lower end of the response curve has little to contribute.
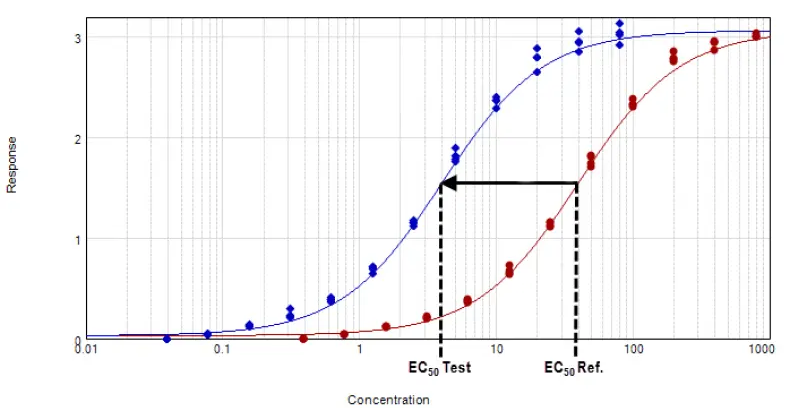
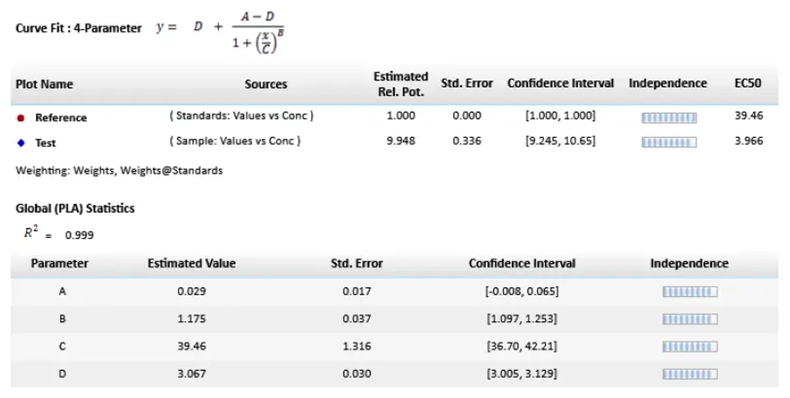
Figure 3. Parallel line model for non-linear regression. The relative potency is determined in the linear region of the curve where the response changes relative to the concentration at 50 % effective dose or EC50. The curves tested are fitted to the constrained model. The parameters describing the curves are identical for all curves except for the X-value in the 4-parameter curve fit equation. Image Credit: Molecular Devices UK Ltd
In the software protocol, the weighting factor is the inverse of the variance, but adjustments can be made to make this a more suitable weighting factor, if necessary. Additionally, there is the flexibility to specify that the weights are to be considered inverse variances (Figure 4G).
The chi-squared profile method for parameter confidence intervals (Figure 4H) can only be activated when the ‘Weights are Inverse Variances’ box is checked. An optimal weighting factor ensures that the most variable data points do not dominate any results.
How to apply PLA in SoftMax Pro GxP Software
In the software, PLA is available across all global curve fits, with the exception of point-to-point, log-logit, and cubic spline curves. In a graph section, the same curve fit functions will be applied across all of the plots. PLA can be implemented as displayed in Figure 4.
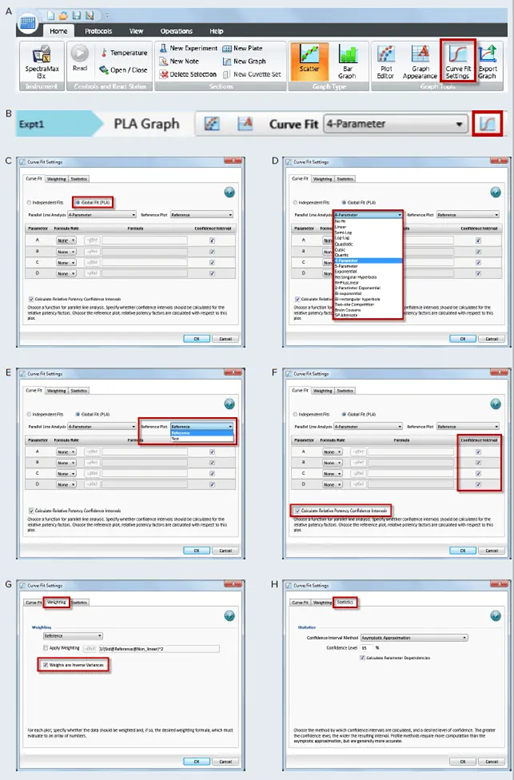
Figure 4. How to apply PLA in SoftMax Pro GxP and Standard Software and estimate relative potency. Select a graph section with multiple plots. Click Curve Fit in the Graph Tools section on the Home tab in the ribbon (A) or in the toolbar at the top of the graph section (B). (C) In the Curve Fit Settings dialog, select Global Fit (PLA). (D) Select any curve fit option except point-to-point, log-logit, or cubic spline from the dropdown list. (E) Select a plot for the Reference Plot list. (F) If applicable, select the curve fit parameters and the Relative Potency Confidence Intervals. (G) If necessary, click the Weighting tab. See application note “Selecting the best weighting factor in SoftMax Pro GxP and Standard Software”. You may also directly select the inverse of the variance weighting factor. (H) If applicable, click the Statistics tab. (I) When all curve fit options have been selected, click OK. The curves tested are fitted to the constrained model. The parameters describing the curves are identical for all curves except for the parameter describing the X-value as shown in Figure 3. For non-linear functions, the minimum and maximum responses (lower and upper asymptote, respectively) are also constrained to be the same for all curves. Image Credit: Molecular Devices UK Ltd
Parameter comparison method
In contrast to the response comparison methods that directly analyze the differences in the dose-response curve, the parameter comparison methods compare the parameters of unconstrained curves against an approximate confidence region one by one.
The parameter pairs must correspond within the determined confidence interval in accordance with a given level of confidence. This type of assessment is known as equivalence testing and evaluates whether there is a degree of parallelism less than the threshold specified. The slope ratio method employed by European Pharmacopoeia is one such example of equivalence testing.
Fieller’s theorem
Fieller’s theorem uses statistics to calculate the confidence interval for the ratio of two parameters.4 The tInv function can be applied to calculate the estimated ratio and consequently follows the t distribution with degrees of freedom, for which the probability is p.
Statistical formulas in SoftMax Pro Software grant users the ability to determine the confidence interval for the ratio of a curve fit parameter between two curves, for instance, reference and test curves.
A protocol (Figure 5) has been developed, incorporating these calculations with a probability set to 0.1 (90 % confidence), which can be adjusted as necessary. To establish whether the reference and the test curves can be considered parallel, the calculated confidence interval is contrasted against a fixed confidence interval that is characterized by a certain confidence level.
In the protocol, a 90 % confidence level is applied, meaning that the calculated confidence interval must be within 0.9 and 1.1 to determine whether the curve is parallel.
When assessing linear regression curves, this test is used for the slope values of the reference and the test curves, which are explained by applying the B parameters in SoftMax Pro Software. However, for non-linear regression curves, parameters depicting the upper asymptote and the slope are tested. Testing of the lower asymptote is not required, as this is a mathematical limitation of the Fieller’s theorem.
At a lower concentration, the parameter exhibits wild variances and produces an intermediate calculation containing imaginary numbers, which result in an incorrect final calculated confidence, or at best, one that cannot be calculated.
This United States Department of Agriculture (USDA) Center for Veterinary Biologics has addressed this issue by recommending that the lower asymptote should be set to zero while applying the slope and the upper asymptote for the relevant tests.5
Likewise, the Parallelism Test protocol built into the SoftMax Pro Software sets the lower asymptote to zero automatically and instinctively tests the slope (Parameter B). Either Parameter A or Parameter D serves as the upper asymptote to establish whether the reference and the test curves can be considered parallel (Figure 5).

Figure 5. Response comparison method in SoftMax Pro Software to assess parallelism. The confidence level and the probability are set to 90 % and 0.1, respectively, but can be adjusted as needed. Once the lower (rBCILower and rDCILower) and upper (rBCIUpper and rDCIUpper) values of the confidence interval for the parameter ratio have been calculated, they are compared to a defined confidence interval (lval and uval). If the calculated confidence interval values are within that defined confidence interval, then the reference and the test curves can be considered parallel for that parameter. Image Credit: Molecular Devices UK Ltd
Conclusion
A large number of biological assays require parallelism to be determined between pairs of dose-response curves. This article has discussed certain response comparison methods, including the F-test and chi-squared test, and parameter comparison methods, alongside an illustration of Fieller’s theorem.
SoftMax Pro Software allows users to choose between a constrained or an unconstrained curve fit model and offers state-of-the-art statistical formulas for comprehensive analysis.
The software also comes equipped with pre-written protocols that are developed alongside the various sections of each test, so the appropriate method can be easily selected to evaluate parallelism for some or all of the parameters describing two curves.
In conjunction with SoftMax Pro GxP Software’s data integrity features, evaluating parallelism and determining relative potency can be conducted in FDA 21 CFR Part 11 and EudraLex Annex 11 compliant laboratories.
References
- Gottschalk, P.G. and Dunn, J.R. (2005). Measuring Parallelism, Linearity, and Relative Potency in Bioassay and Immunoassay Data. Journal of Biopharmaceutical Statistics, 15(3), pp.437–463. DOI: 10.1081/bip-200056532. https://www.tandfonline.com/doi/full/10.1081/BIP-200056532.
- Vandeginste, B. (1989). Nonlinear regression analysis: Its applications, D. M. Bates and D. G. Watts, Wiley, New York, 1988. ISBN 0471-816434. Price: £34.50. Journal of Chemometrics, 3(3), pp.544–545. DOI: 10.1002/cem.1180030313. https://analyticalsciencejournals.onlinelibrary.wiley.com/doi/10.1002/cem.1180030313.
- Draper, N.R. and Smith, H. (1998). Applied Regression Analysis. Wiley Series in Probability and Statistics. DOI: 10.1002/9781118625590. https://onlinelibrary.wiley.com/doi/book/10.1002/9781118625590.
- Buonaccorsi, J.P. (2006). F ieller’s Theorem. Encyclopedia of Environmetrics. DOI: 10.1002/9780470057339.vaf004. https://onlinelibrary.wiley.com/doi/10.1002/9780470057339.vaf004.
- United States Department of Agriculture Center for Veterinary Biologics Standard Operating Policy/Procedure. 2015. Using Software to Estimate Relative Potency. USDA Publication No. CVBSOP0102.03. Ames, IA.
About Molecular Devices UK Ltd
Molecular Devices is one of the world’s leading providers of high-performance bioanalytical measurement systems, software and consumables for life science research, pharmaceutical and biotherapeutic development. Included within a broad product portfolio are platforms for high-throughput screening, genomic and cellular analysis, colony selection and microplate detection. These leading-edge products enable scientists to improve productivity and effectiveness, ultimately accelerating research and the discovery of new therapeutics. Molecular Devices is committed to the continual development of innovative solutions for life science applications. The company is headquartered in Silicon Valley, California, with offices around the globe. For more information, please visit www.moleculardevices.com.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.