Jacopo Annese, President and CEO of the Institute for Brain and Society, a non-profit organization dedicated to democratizing neuroscience and making neuroscience tools and knowledge about the brain more available to the public, discusses his work on the Human Brain Library.
In what ways can the human brain be studied?
There are currently two ways of looking at the human brain. We can look at the brain non-invasively, using techniques such as magnetic resonance imaging (MRI) and computerized tomography.
Methods of Examining the Brain
Methods of Examining the Brain from AZoNetwork on Vimeo.
MRI is a wonderful technique because, although the resolution is low, it still allows quite a detailed view of the brain, while the person is alive.
You can use that to make decisions about treatment or to see what's going on. However, as it's very low resolution, not all the answers can be found.
The other way of looking at the brain is post-mortem, which is necessary when you want to see tissue at a microscopic level.
Histological techniques are used to process brain tissue and look at individual cells and fibers and thereby observe pathological phenomena that create disease.
For example, with Alzheimer's Disease when you look at the brain using MRI, you can see shriveling, or a loss in mass, but you don't really see the actual hallmark lesion of Alzheimer's, which is the plaques, the tangles.
In order to see those, you have to work on post-mortem, donated brains. You slice them and stain the slices (which is actually an old procedure as histology started 200-years ago).
It is still the only way we have to look at individual cells and the processes that caused the disease. It's extremely important and it's still the gold standard.
These days of course, just like anything else, this branch of neuroscience has entered the digital domain and so now we use digital imaging to look at cells.
We can catalogue them and there are beautiful, quantitative tools that allows us to study the individual brain in microscopic resolution.
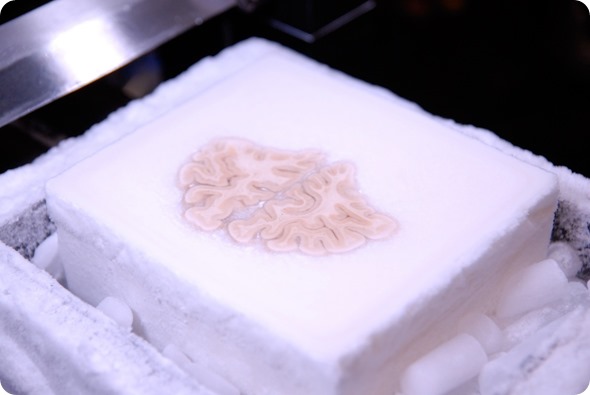
Image of the frozen brain at the level of the frontal lobes during the cutting procedure. Image credit: Huron Digital Pathology.
How much do brains vary from person to person?
What interested me the most, as a brain scientist, was not so much what we have in common in terms of brain anatomy, but what distinguishes each and every individual.
Since the brain and its connectivity, the way it develops and the way it matures is very unique, more so than any other organ of the body, it's very different in different people.
Mapping the Brain Universe
Mapping the Brain Universe from AZoNetwork on Vimeo.
There's not one brain that looks like another. Even in monozygotic twins, the brains are different, because the twins have grown, been raised differently and made different choices. It's really wonderful.
That's why we talk about the brain as a personal universe, because you could never really stop studying the brain. Even if you precisely mapped the brain of one human being, you would almost have to start from scratch when you mapped somebody else's brain.
I love the technology enabling us to digitize a brain to cellular and even sub cellular resolution. When you zoom in, you find layers and layers of detail and then the differences between two brains become overwhelming.
Essentially, we wanted to catalogue brains, which is why I called our project the Brain Library rather than the Brain Bank.
Our function is not so much to receive a brain specimen, process it and distribute it, but to receive that donated brain, virtualize it, catalogue it and make all that information such as images and data that we extract from that particular brain, available to the widest possible research community.
How do we do that? Through the web. We essentially digitize everything and then it's put into a format, images or data, that can eventually be downloaded from the Cloud. We call them ‘Brains in the Cloud’, which is a very strange image.
We're beginning to look at different diseases such as Alzheimer's and Parkinson's, not as diseases that act the same way in every patient, but actually diseases that follow the individual pattern of connectivity of the brain.
Just like if you looked at two different cities from the satellite using Google Maps; you would see a different roadmap for San Diego than the roadmap for LA.
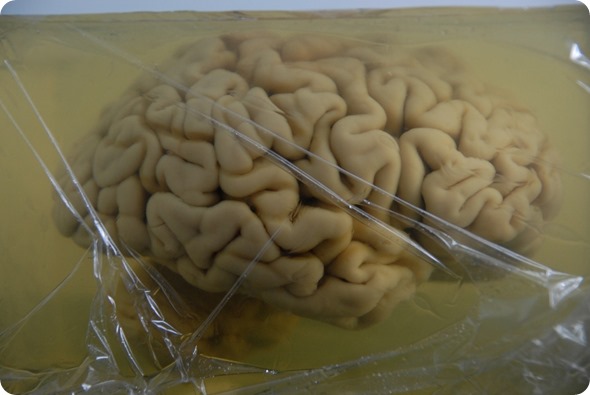
The brain of H.M. is embedded in a gelatin block. Image credit: Huron Digital Pathology.
Who was patient H.M. and why was he famous?
A lot of the motivation to create the technological infrastructure for most of the Human Brain Library stems from one particular, iconic case in neuroscience, which is the case of Patient H.M. We now know his name was Henry Molaison.
When he was a young boy, he suffered from epilepsy. He underwent brain surgery when he was in his late 20s and a part of the temporal lobe was removed from both hemispheres.
This was in 1953 and the surgeons didn't know at the time, that we need those areas to create memories. When this patient woke up, he could not create new memories and he became the most famous amnesiac in the history of neuroscience.
He passed away in 2008. The Brain Observatory was asked to create a map of his brain, because we had already developed new technology.
We were able to histologically process a whole human brain in one piece. The project became more about data sharing.
Since thousands of publications had been written about him over 50 years or more and because it was so iconic, what I wanted to do with this very famous brain, was to create an open source brain.
Instead of a collection that would sit in somebody's lab for years and decades, I wanted to share this brain with the scientific community.
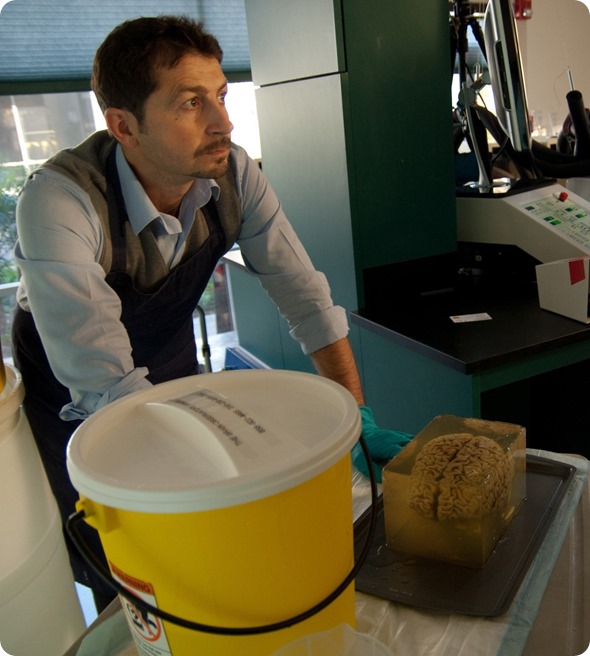

Dr Annese during the preparations for cutting. Image credit: Huron Digital Pathology.
How did you carry out the scanning for patient H.M. and how long did it take?
The project started developing into different phases. A very exciting phase was preserving this brain in the form of slides, so slicing and staining - it was almost like a forensic study.
The goal was to understand what the surgeons did in 1953 that created this syndrome, this amnesia. It essentially started the branch of human memory, but nobody had actually looked inside the brain of this patient, because you cannot really do that in detail until the brain is donated.
We wanted to understand what happened during that fateful surgery. We had to create a histological collection to be able to see the lesion in microscopic detail.
The other part, which is just as exciting, was that, as we do that, we create a digital archive so that this information is not just published in a scientific paper, but is actually shared with the scientific community.
Also, we wanted to share it with the public because the public knew about H.M., especially after he died.
The news came out and his identity and history were revealed. He also became a very fascinating case. If you look at the media and news at the time in 2009, everybody was talking about him.

Dr Annese cutting the brain. Image credit: Huron Digital Pathology.
The first phase was the methodology in the lab to process the brain specimens and form a histological collection. The next step was to virtualize this histological collection.
The third phase was to make it available through the web and essentially create the files that people could view over the web, at home on their computers or at a lab in Germany or Japan, or wherever they were.
When it comes to sharing data images of the brain acquired with MRI, the output is digital images. It's straightforward; you can simply create image servers and hospitals use picture archiving and communication systems (PACS) where these digital images are made available.
Sharing images of the actual tissue that we acquire from physical, histological slides involves a cross-section, an actual physical slice of brain that has been mounted on glass and stained to reveal fibers.
There's an enormous amount of information within this 20 microns of slice, which is mounted on the slide. When you zoom in, you see individual axons that come from different neurons. You may or may not see, it’s a much fainter stain, but there is another dye that we use to essentially stain neurons in the slices.
Whereas we use silver particles to stick to axons for these particular stains, this is a dye that sticks to DNA inside the cells. Therefore, we're able to individualize and count each cell. There are probably hundreds of thousands of cells just in this. For every brain, we create about 2,500 to 3,000 of these.
How have scanners advanced since the post-mortem of patient H.M?
In 2008, when we started with project H.M., we had modified microscopes to do this work.
This is a very large format; the surface that we need to image is very large, which was, of course, a bottleneck, because taking microscopic pictures of such a large format, would take us about a day and half, about 19 hours.
Your pipeline gets delayed when you have so many. Imagine the assembly line; you create a lot of slices and then they all get crammed at the digitizing stage.
Digitizing Pathology
Digitizing Pathology from AZoNetwork on Vimeo.
These days, the technology has improved a lot. There are line scanners and technology that can essentially do what we used to do in 19 hours in just one hour.
H.M. was one brain, but the goal of the Human Brain Library is to digitize 1,000 brains, because we want to have a cross-section of different patients, and likewise, of different people without neurological disorders.
To represent the validity, we need to have very many brains. Each brain, when digitized, is potentially 1 petabyte of data, so the challenges are in the speed and ease of digitization, the pipeline, and then, of course, storage.
Technology came to our rescue in a sense, over the course of the project, because now the image compression has become much better. This means you can still visualize the detail you need, but in file, they are much smaller than they needed to be before.
The speed of digitizing has also improved a lot, which is crucial to us, because we need to make thousands and thousands of slides available to researchers.

Image credit: Huron Digital Pathology.
We want to share our microscopes and create a virtual microscope that people can use, wherever they are. I think this technology could be made available not only to experts, but to anyone who is able to view the images via a screen.
For example, a middle school student doing a project about the human brain, may want to show some pictures of neurons. Rather than going to Google and looking at some random pictures, they could access the Brain Library, choose which particular person they would like to represent and look at areas of their brain.
Motivation to do the preparations is also important since this technology is almost like quality control for a lot of the work that we do because, now, we cannot hide anything.
I would never want to hide anything, but when all you have to present your image in a published journal, is a one-inch square, you've got to choose what's the best way of presenting your data, your results.
This way, however, I'm able to have supplementary data for my paper and show them everything. Even if in the actual journal there's only going to be a sample, the reader can then go online and view the whole brain.
For me, that's accountability, because if that brain is not well done, then people are going to say "oh yeah, that image looked nice, but the rest of the brain is terrible" or "wait a minute, this doesn't match."
It's good to have accountability in research and I think this technology can contribute a lot towards that.
At the time, when we digitized the first batch of slides from H.M., we had engineered microscopes that we built from some basic model we had re-engineered. It was a bit of a Frankenstein-type piece of equipment.
Developing new equipment and new technology is a very long and expensive process, so it was a huge investment at the time for my lab.
It was okay for a one-off scenario; once you got it done it was great, but it was evident that if we had to industrialize that process, it wouldn’t have then been practical because it also required a lot of technician time to set up.
Now, the technology has advanced so that instead of a microscope, in terms of speed, it's almost like a document being scanned on a very high resolution flatbed scanner.
The technology of digitizing these histological, pathological, slides is more similar to a flatbed scanner than a microscope.
The challenge there for the industry and those who developed this, is, of course, gaining or acquiring images at higher speeds and with greater detail.
For us, what was important, was to accelerate the process of digitizing the materials because when you have hundreds and thousands of slides, you don't want to be held back. To us at least, this new technology is very important because we have very large volumes of slides.
The project is ambitious. If you want to archive hundreds of thousands of slides that represent each of these people's brains in detail, we need to create this archive in a faster way.
The other point is, again, the file format and the ability to share it with other researchers. There's really no point in having all this digital information on my hard drive in the lab, but with compression and using cloud technology, this can be shared.
Can you please outline the project you are currently working on with the brain of a painter? What are the main aims of this work?
One of the last cases that we digitized was the donated brain from a person who was an artist, a painter who specialized in painting horses. This was very interesting to me, personally, because we were essentially creating art - brain art.
These images are very beautiful to look at and some stains are just gorgeous. We produce them as art that could be displayed in a home or in an office.
The idea of this artist's brain kind of transcending into more art, makes it seem almost as if he didn't finish creating art, even after he had passed.
We looked at his brain to find out what made him an artist and how his brain enabled him to have that talent.
We knew there were papers published recently, that used MRI to give an outside view of the gross anatomy of the brain and showed that there are some morphologies you can find in the brain of people who are trained in drawing.
It's not really far-fetched anymore to consider what it is about a person’s brain that enables them to be an artist.
That's why we do what we do. Apart from being a tool for diagnosis to help find treatments for diseases, the idea of the Brain Library is to try to really understand what is at the basis of being human. For example, what makes people choose a profession or have passions that are different from those of somebody else.
In this case, this was a person who discovered early on that he had a talent and pursued that as a career. I'm convinced that we're going to find something in his brain. We need to study many brains of artists, because one result is only anecdotal.
If we see differences between his brain and the brain of somebody who was not an artist, who was maybe a businessman, or an engineer, we cannot then draw conclusions.
That's why the Human Brain Library has to collect so many brains. We need to generalize the subjective information from each individual person, into some patterns where we can then say "yeah, this actually is true."
This is what they do now with MRI, because it's easier. You can recruit 30 people, perform scans, with each taking 10 minutes and everybody goes home happy. No donated brains are required and it's much simpler.
There are MRI studies of thousands of people, looking at patterns among people with a disease or comparing people who do certain things.
This is the next level of analysis. It's post-mortem, so it makes it a little more ghoulish as a project, but yet fascinating, because we are really looking into the underlying biology of what these people were when they were alive.
That makes it very human and is why we call it the Human Brain Library. We connect the person to the brain. Different people have different brains and as you zoom in, you find that the differences get increasingly bigger.
One day, we will have maps that will essentially tell us that this is how artists are wired and enable us to look at the roots of creativity, the roots of genius, if you will. Nowadays, it's very simplistic.
There are scientists, there are artists, but the scientist could be an artist, as many scientists are. They live their science as an art, or they're very artistic in the way that they do their science.
I think the brains of these people will tell us what they are really capable of. One day, we may even be able to predict whether a child could be a good artist, based on what their brain looked like.
They may be an engineer, but we should know. There are people who discover painting and that they have talent when they're 80, because they never had the opportunity to discover it before, but their brain would already have been wired to allow them to do that. If they had known, they could have started painting earlier.
How do you think this technology will impact brain research moving forwards?
The field of neuroscience, just like any other field of science, has become very reliant on technology. It's up to the investigator or scientist to maintain an oversight and his own judgment because technology can also take over.
You can delegate science to technology, but what makes the discoveries is human intuition. It’s the balance of these two, when human intuition can use technology for discovery, that has led to so many advances lately.
In terms of digital technologies for anatomy or pathology, the great advance was to be able to share this information. Before, every hospital research lab had their own collections and it was very insular.
Now, there's definitely more collaboration. This enabled cross-validation between different labs and a lot of cooperation. For example, somebody in Japan could be studying the same images that I am studying here in my lab.
We could do it together and at the same time, like a conference call. They would see the same detail that I do and be working on the same case.
That's what's really good about this technology. I think in the future, if I had a company that created this sort of technology, the pull would be to make it as user friendly as possible.
The boundary between science and consumers is blurring.
I've been using more and more off-the-shelf consumer products to do my research. The computer I'm sitting next to has 192 GB of RAM.
When I bought it, it cost about $11,000. Now, it's half the price and it has 250 GB of RAM. We need it to manipulate volumes that we create out of imaging that are very, very large in terms of size.
For 50-60 GB, we need a lot of RAM memory to be able to visualize the data, but now it's become affordable.
I think this process is going to continue, and, in the case of a microscope scanner, for example, eventually anyone should be able to use it.
You wouldn't need to know MATLAB or how to program. You wouldn’t need to know how to use or be trained on very difficult software.
I think the next step is to make this technology, which is scientific software and hardware, just as easy to use as the camera that you buy in a store because people who make consumer products know that if they're too difficult to use, people are going to just leave them alone.
Scientists still have a lot of graduate students who can sit there for days figuring out how software works. I think the steep learning curves that we always see in our field should be eliminated.
I should be able to do what I do picking students from middle school, who are much smarter than I am already, with a lot of technologies that I don't know how to use. I think that's the next step.

Tissue sections mounted on glass slides before staining. Image credit: Huron Digital Pathology.
Can you tell us a little bit about the Brain Observatory concept?
The idea behind the Brain Observatory is a laboratory that's also open to the public. Whereas labs or research centers usually have organized tours and, occasionally, a show and tell, it’s really not the space where research happens and the public is generally not on the premises.
We want to have a location that will be very accessible, so we actually want to have it downtown, next to other retail stores, that attract foot traffic.
Our non-profit initiative is about brain health, we want to make science directly available and the information and knowledge about the brain accessible to the public.
One way of doing it is a retail approach to the lab. We would open these Brain Observatories in different cities, they would essentially be living showrooms of research and science, because researchers would actually be working there, but there would also be areas that are like a brain exhibit, a brain museum.
Imagine the “Body Works” exhibit, but for the brain. You can imagine having, of course, MRI machines so that people can actually also use that location to have MRI scans.
If you wanted to be even more creative, you could have cafeterias. We know that cafeterias or food attract foot traffic and today, museums, for example, all have cafeterias.
In this case, it would be a cafeteria or a restaurant themed around food that is healthy for the brain. That would also serve our educational mission, to make people informed about which foods and lifestyles are brain healthy and which ones are not.
The mission of the Institute for Brain and Society was essentially to make the knowledge and tools that researchers or healthcare professionals have available to the public and basically make them available to everybody, including the layperson. The idea is to democratize the technologies that are actually very useful in terms how our brains are doing.
Just to put it in context, we now routinely check our blood pressure; we check our skins for moles that could be malignant and generally do a lot of routine health checks that we now take for granted.
The only organ that we don't check regularly is the brain. I think everybody would agree that the brain is probably one of the most important parts of our body.
Without my brain, I couldn't speak about this and I could not have obtained my degree. I could also not have loved who I have loved in my life.
In my opinion, it has always been strange to me, that people do not seem interested in checking how their brains are doing, and it’s not because I’m a neuroscientist.
I always make an analogy with the black box of an airplane. If the airplane crashes, then the technicians would look at the black box to try to find out what happened. This is actually how we use diagnostics now for the brain.
If I had a symptom of a stroke, then eventually a doctor would refer me for an MRI, so they could see what happened. Maybe they'd see the stroke or that there was fluid in the brain and, at that point, they would intervene.
However, the stroke, the fluid, the dementia all happen long before we actually show symptoms. If we could see this happening in the brain regularly, we would start doing something about it. I would prevent getting a stroke if I saw some indication that that might happen.
What the non-profit Institute for Brain and Society does, is try to make these technologies more affordable and more available, so that we encourage people to check their brain, with the same ease as they check other things.
Of course, our challenge is to make this technology accessible, both financially and logistically. Where are these MRIs and how can we send somebody to have an MRI?
It's also culture. Education is one of our biggest concerns because we need to make people conscious of the fact that having a routine health check for the brain can actually improve their life or even save their life in the future.
I think once we can help them understand and interpret these images that they gather of their brains, we can then coach them about ways in which they can actually preserve their health.
If we see, for example, in an elderly individual, that there is a pattern that is not very positive, there are ways in which you can influence the outcomes, even in the case of dementia.
There are several publications connecting the Mediterranean diet with lower incidence of dementia.
I'm not saying that's the silver bullet, but if I saw in my brain that there were some areas becoming atrophied at a faster rate than they should, I think I would be very likely to eat the foods that I know have shown some positive effects on the brain or engage in activities that we also know are helpful for oxygenation of the brain.
I think information is very important. I think being able to view your brain is very valuable and that's what we're trying to do for the public, for everybody.
Which foods should people eat to maintain a healthy brain?
There are super foods for the brain. I would say olive oil is the king of brain foods.
We often think about whether wine is good or bad; a lot of people ask me that. It really boils down to what's in the wine. I personally try to drink organic wines. In Italy, there's a trend now that's been happening for a while, referred to as biodynamic agriculture.
Essentially, it's very similar to organic agriculture, so there is no use of pesticides and also the use of sulfites is limited or eliminated. Taken as fermented grape juice, red wine with the skins and the tannin from the skins is very rich in polyphenols, which are very good for the brain. There are other foods that are rich in anti-oxidants, which are very good.
There is no brain diet that I could concoct. I could be very close to something that we know would increase your chances of not becoming cognitively impaired with old age.
The health of the brain is essentially also the health of the heart, so exercise is very important. A healthy heart is a healthy brain.
Even though Alzheimer's is the most commonly used name for dementia, many dementias are not caused by Alzheimer pathology, but by vascular problems such as hypertension. I think the neurologists and the cardiologists should always work together on somebody's health.
The body is always holistic. The brain doesn't sit alone in the body; it's connected with everything. It's about awareness.
If I see a suspicious mole getting bigger, I can seek advice, but I cannot see my brain. The only tool now available for me to take a view of my brain is MRI.
Personally, I have a scan every year, just to check and I did it on my last birthday, when I turned 50 two months ago. It was great to know that I'm doing pretty well.
I'm in the right percentile in terms of size, health and the way it looks and there was nothing to worry about. It made me feel as if I can step into the second half of my life knowing I have a healthy brain.
I can work and embark on a big project such as the Human Brain Library knowing I have a healthy brain to do it.
Where can readers find more information?
About Dr Jacopo Annese
Dr. Jacopo Annese graduated from the University of Rome in Italy with bachelor’s and master’s degrees in biology and zoology.
He earned a master’s degree in neurological sciences at University College London, U.K., and completed the Ph.D. program in cognitive neuroscience at Dartmouth College, Hanover, NH.
He subsequently trained in computational anatomy at the Montreal Neurological Institute, Canada, and at the Laboratory of Neuroimaging at the University of California, Los Angeles.
His research has been featured in a variety of broadcasting and publishing media, including PBS NOVA, National Geographic, BBC, Discovery Channel, The New York Times, Bloomberg News, Financial Times, Esquire Magazine, and Discover Magazine.
Dr. Annese's primary goal in the field of neuroscience is to conduct research that is open to public engagement and promotes the highest standards in data sharing and collaboration within the scientific community.