A research group from Nigeria found evidence of long-distance transmission and dispersal between bat coronaviruses isolated in Africa and the ones Europe and Asia, as described in a new paper available on the preprint server bioRxiv*.
The coronavirus disease 2019 (COVID-19) pandemic, caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), became the defining global public health crisis of our time. This virus belongs to the broad group of coronaviruses, which are known to have a zoonotic potential of spreading from animals to humans.
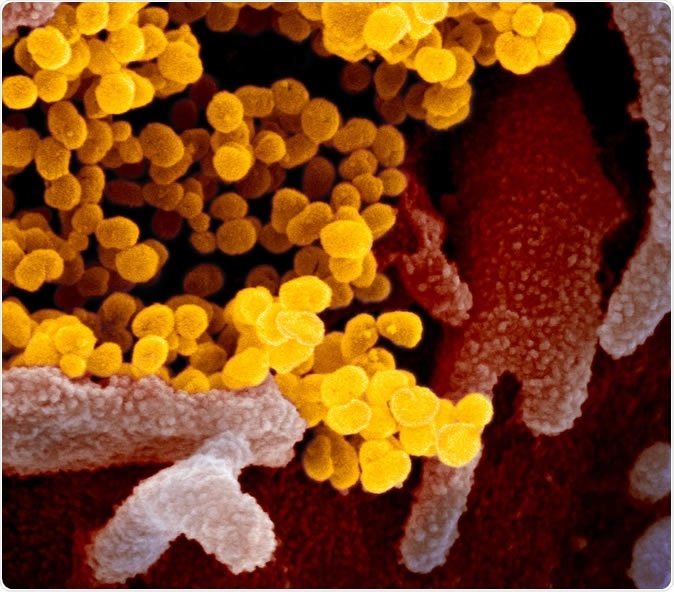
Novel Coronavirus SARS-CoV-2 This scanning electron microscope image shows SARS-CoV-2 (yellow)—also known as 2019-nCoV, the virus that causes COVID-19—isolated from a patient in the U.S., emerging from the surface of cells (pink) cultured in the lab. Image captured and colorized at NIAID's Rocky Mountain Laboratories (RML) in Hamilton, Montana. Credit: NIAID

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
The rich fauna and biodiversity in Africa created a hotspot for emerging viral diseases. Additional opportunity for viruses is an assortment of bats that may serve as reservoirs, and that may efficiently spread the pathogen (for example, African fruit bats are known for covering thousands of miles during their migration cycles).
The evidence points toward African bats as a potential reservoir for several betacoronaviruses that may cause serious outbreak events. Some studies have also suggested that the SARS-CoV-2 likely spilled over into the human population by a zoonotic event involving SARS-related betacoronavirus.
Consequently, a research group from the Federal Medical Center in Abeokuta, Covenant University in Otta, and the University of Ibadan in Nigeria decided to appraise the phylogenetic diversity and evolutionary dynamics of African betacoronaviruses among bats, as well as their possible dispersal across the continent.
Analyzing the evolutionary tree
Three data sets were generated from the Virus Pathogen resource databases for the purposes of this study. More specifically, the authors obtained sequences of the partial RNA-dependent RNA polymerase (RdRP) gene of bat coronaviruses from seven African, four European and three Asian countries.
Information such as host species, country of origin, and collection dates was combined with sequence data in order to conduct phylogenetic determination accurately. Cluster analysis was carried out with the specific software to identify sequence similarity and reduce data duplication.
Phylogenetic trees were inferred with the use of MEGA software for manual and automatic sequence alignment, distribution analysis (both geographically and evolutionary) was conducted by using Markov Chain Monte Carlo implemented in BEAST (a platform for Bayesian phylogenetic molecular sequence analysis) and visualized with SpreaD3 interactive platform.
A virus that jumps continents
The most prevalent bat species sampled in this study was Peters's dwarf epauletted fruit bat (Micropteropus pusillus). At the same time, Cameroon was the country with the highest distribution of bat species sampled in this study. However, recognized data gaps should be taken into account when appraising study findings.
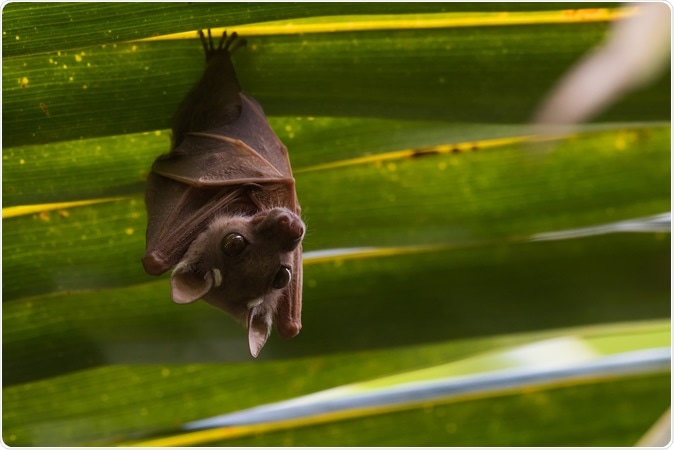

Peter's Dwarf Epauletted Fruit Bat (Micropteropus pusillus). Image Credit: Dave Montreuil / Shutterstock
"This result does not necessarily represent the true picture of bat species diversity in Africa, as some countries lack sequence information for bats due lack of surveillance," caution study authors.
Actually, thus far, only a handful of studies have been conducted in Africa on coronaviruses among bats, which leaves a considerable rift in epidemiologic information regarding beta bat coronaviruses in Africa.
This study adds additional knowledge on the phylogeny of beta coronavirus sequences, which revealed that a majority of the African strains fall within the lineage D – consisting of strains from Cameroon, the Democratic Republic of the Congo, Kenya, Madagascar, and Nigeria. It corroborates previous reports that identified a single clade circulating throughout the African continent.
Moreover, the researchers hinted towards inter-species transmission among the lineage D betacoronaviruses, allowing for potential recombination and rapid evolution of this lineage. Also, the circulation of two distinct lineages B and C among African bat species was demonstrated.
Finally, phylogeographic dispersal of bat coronaviruses revealed a plethora of inter-continental and intra-continental spread events. The former shows viral dispersal from China and Hong Kong to Central and Southern Africa, while the latter demonstrates the spread between Cameroun, the Democratic Republic of the Congo, and South Africa, as well as directly from Cameroon into Madagascar.
Study shortcomings and conclusions
Although these insights are pivotal for further tracking of coronaviruses, one major study limitation was the inability to analyze spike protein sequence data of these viruses. Such an approach would have provided more pertinent data on their evolution concerning transmission and infectivity.
In addition, the population demography reported in this study might not realistically depict the viral population since the size of the utilized dataset is limited and might not represent the true demographic population of bat coronaviruses in Africa.
"We have, however, shown the importance of molecular surveillance of viruses with zoonotic potential such as coronaviruses," say study authors. "We advocate for broader trans-continental studies involving full genome sequences of betacoronaviruses to further understand the drivers for their emergence and zoonotic spillovers into the human population", they conclude.
In any case, we have to be aware that bat SARS-related coronaviruses will continue to spill over to the human population. The chances of another deadly pandemic are indeed rare; however, those chances increase with the frequency of such spillovers. Thus the scientific community has to stay vigilant.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources