When I (James) started graduate school, I had never worked in bacteria before. I originally worked with Drosophila, but when I joined Zemer's lab, we basically came to the table with the question, "How can we address the global need for new antibiotics in light of the escalating rates of antibiotic resistance?"
This is a problem because as the rates of antibiotic resistance go up, we should be making new antibiotics that can treat those antibiotic-resistant infections.
However, only six new classes of antibiotics have been approved in the past 20 years, and none of them can treat Gram-negative bacteria. There is this huge gap in the field that we wanted to address by, one, creating a new pipeline for discovering novel classes of antibiotics, and two, creating a way to quickly and efficiently characterize their mechanism of action.
The aim was to find those mechanisms of action that are unique, such that bacteria would have never seen them before, and the hypothesis was that if a mechanism of action is new, bacteria will have a much harder time becoming resistant to it.
Traditionally, antibiotic research and new leads have been done with a certain style that has sort of been dominated by these sorts of chemically-focused perspectives, and the big-picture idea was can we use some fresh approaches to get something new here.
.jpg)
Image Credit: Rost9/Shutterstock.com
What is the difference between Gram-positive and Gram-negative antibiotics, and why is this important in your work?
So first it is important to make it clear that the bacteria are the ones that come in two flavors, the Gram-positives and the Gram-negatives, and the antibiotics are the compounds that then kill those bacteria. When we say Gram-positive antibiotics we mean an antibiotic that kills a Gram-positive, or a Gram-negative antibiotic that kills a Gram-negative.
The distinction is that Gram-positive bacteria only have one membrane layer and then a cell wall outside of that membrane, whereas Gram-negative bacteria have a second membrane outside of the cell wall; they have an inner membrane, then the cell wall, and then the outer membrane.
Gram is simply the name of the person who invented this Gram stain to test for the second membrane layer. Because of that second membrane, the Gram-negative antibiotics are not stained and in the same way that they exclude that stain, that second membrane is also a mechanism to exclude or prevent entry for many different small molecules.
The reason that Gram-negative pathogens are so insidious and hard to kill is that they have this second barrier, this outer membrane, and so they are harder to actually penetrate.
Please explain your new antibiotic and how it works?
This small molecule, SCH-79797 (SCH) was originally used in mammalian experiments to treat thrombosis, as it is a thrombin R inhibitor. We found that it has potent activity in bacteria, and in short, it works by two different mechanisms that complement one another.
One mechanism is that it is able to penetrate and disrupt the membranes of Gram-positive and Gram-negative bacteria, but then it is also able to, once it is inside of the cell, disrupt intracellular processes.
In particular, it disrupts the ability of the cell to make folate, which is a necessary nutrient that the bacterial cells need to make essential parts of life, DNA, amino acids, proteins. Without these things, the bacteria cell will never grow.
Image Credit: Victor Josan/Shutterstock.com
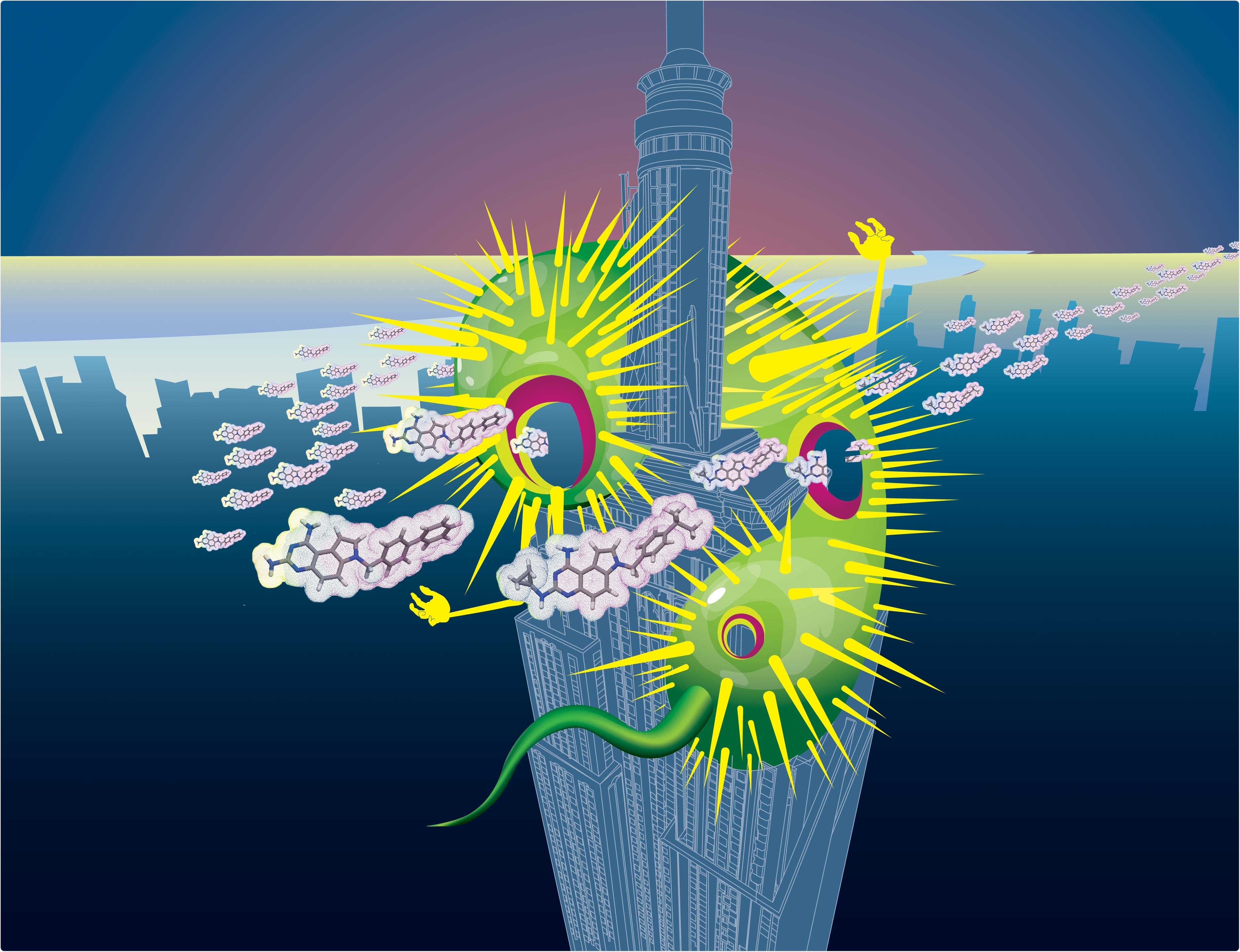
Why has your technique been likened to a poisoned arrow?
The idea is that for an antibiotic to work, it needs to get into the cell, and then it also has to kill. What we have found is that our antibiotic does two things simultaneously.
First, it pokes holes in the bacterial membranes, and so that we liken that to an arrow. That alone is bad for the bacteria - obviously having a hole in your membrane is bad as stuff leaks out. However, it also then inhibits essential folate metabolism processes once it gets inside.
We think of the membrane permeabilization, or the hole poking, as the arrow, and then the inhibition of the ability to then make the building blocks that are needed for the bacteria to grow and divide, the folate inhibition, as the poison.
How did you develop your new antibiotic?
Science is a collaborative effort, and we could not have done this work on our own. We had the help of our co-authors Joe Sheehan, Ben Bratton, and our other collaborators.
Our compound is unique and has a mechanism of action that has not been characterized before. Something else that is interesting about this antibiotic is that we were not able to acquire resistance to it in our experiments. Resistance is one of the primary ways that scientists are able to ascertain the mechanism of action of an antibiotic.
Without this, we had to employ various methods to come at the problem from different angles. We used metabolism; we used flow cytometry, where we looked at the membrane; we used quantitative high throughput imaging under the microscope; we used genetics; we used proteomics.
Basically, to develop this new antibiotic, we employed all these different methods to get different pieces of the puzzle, and then we were able to piece them all together. At the end of the paper, we have this figure where we try to sum up the project.
We did something known as bacterial cytological profiling, or BCP, where we can look at how bacteria are affected by a certain antibiotic. From this, we showed that no antibiotic that we tested looks similar to SCH.
If you combine two antibiotics that have the two separate mechanisms of action of SCH, and co-treat bacteria with them at the same time, those bacteria now look similar to SCH when they die.
We then made a derivative of this molecule which we call Irresistin-16. This was able to have even greater potency and have a greater therapeutic effect in our gonorrhea mouse model than SCH.
How did you test the resistance of your antibiotic?
In many ways, that was the hardest part, as scientifically it is impossible to prove a negative. We cannot prove that something cannot happen, and so we try to be careful and say that we were unable to detect resistance, or the resistance frequency was below the limit of our detection.
This was a really important point that James worked on in a few different ways. The traditional way is to put bacteria on plates that have the antibiotic and the thought is that if one of the bacteria has a mutation that makes it resistant to the antibiotic, then it will grow even in its presence, and you will then be able to isolate the mutant.
James tried that with no success, but then he tried a much more quantitative and careful approach, which convinced us that there was no detectable resistance. He took the drug in half the minimal concentration needed to kill the bacteria. In other words, he took a concentration of the drug where the bacteria can still grow, but not well.
It was a multiple-month long experiment that he did, where every day or two he would come into the lab and he would grow up bacteria in this concentration that was right on the edge where the antibiotic is effective.
The thought was that if there was even minimal mutation that conferred any advantage, those bacteria would grow faster, and because they would grow faster, they would then out-compete their neighbors and would become enriched in the population, and over time you would get this very gradual increase in the resistance.
He tested this on many different antibiotics, including antibiotics that are notoriously hard to get resistance to, and he showed that this concept works. Gradually, he was able to evolve resistance to every other antibiotic that he tried, but to SCH there was zero increase in the resistance. It was a completely flat line.
Why were you unable to use traditional methods to figure out the mechanisms of antibiotics in this research?
The traditional way of figuring out how an antibiotic works is to find resistance mutants. Those usually map to the target of the drug, and then that tells you what the drug is. In this case, there is sort of a double-edged sword.
Clinically, the fact that there is no resistance to these compounds is exactly what we want. That is the Holy Grail of antibiotics - an antibiotic that you cannot get resistance to, but at the same time, the reasons that we had to use all of these different approaches was the same reason that we could not eliminate resistance using the traditional way of figuring out the mechanism of action.
.jpg)
Image Credit: Kateryna Kon/Shutterstock.com
What was the problem with the original SCH 79797 and how did you solve it?
The problem with the original SCH is that it was great at killing bacteria, but when we tested it, it was almost as good at killing many mammalian cells. That is a problem in the sense that for an antibiotic we want something that will kill the bacteria but not kill us.
Bleach, for example, is good at killing bacteria but is not a good antibiotic because it kills our cells just as well. In its original configuration, SCH was not quite like that, but it was in that direction, and so what we then did was make derivatives of SCH.
We were very excited that we found this derivative of SCH, which we named Irresistin-16. The great thing about Irresistin-16 is that it maintains the same level of toxicity towards mammalian cells as SCH, but it kills bacteria at 100 to 1000 times lower levels, so there is more potency towards killing the bacteria.
Since we did not change the level at which we kill the mammalian cells, but we dramatically changed the level at which we kill bacteria. This means we have this 100 to 1000-fold window of concentrations that can be below the concentration of killing the mammalian cells, but above the concentration of killing the bacterial cells.
What examples of bacteria have you shown your antibiotic to be effective against?
One is MRSA, methicillin-resistant Staphylococcus aureus which is a huge problem, especially in hospital settings. Another is Acinetobacter baumannii, which is a Gram-negative bacterium that is notorious for its antibiotic resistance.
This bacteria was actually a big problem here in America during, I believe, the war in Iraq, because this bacteria is very heat tolerant, and so in those conditions, the bacteria would live and subsist on medical equipment or beds. Soldiers who needed amputations or something of that nature, would get this infection and bring it back home.
Neisseria gonorrhoeae was a big one, and that was very important because it is Gram-negative and we are running out of antibiotics to treat drug-resistant gonorrhea. I believe we recently found resistance to the last-resort antibiotic, and so it is imperative. Even the World Health Organization says that this is a problem that needs to be addressed.
One very exciting thing is that the World Health Organization has a strain of Neisseria gonorrhoeae that is resistant to almost all known antibiotics, and yet our drugs were able to kill those strains. That was very exciting for us, and it suggests that this concept that we started with, a genuinely new class of antibiotics, could really work in combatting antibiotic resistance.
Image Credit: Kateryna Kon/Shutterstock.com
What stages are needed before your drug can hopefully be used to treat these infections in humans?
First, we have to make it clear that there are a lot of steps between this and applying it to people. At the moment we have helped treat gonorrhea in mice.
We are actively working on various things to improve the drug. There are three main things. First, we want to kill bacteria even more effectively, which we call potency. Second, we want to make it even less toxic to mammalian cells. Third, various drug-like properties need to be improved.
For example, Irresistin-16 is very hydrophobic and so it does not dissolve very well in water. It would be impractical as a pill, so at the moment we have been injecting it. There are various things like that that need to be optimized. However, we are encouraged that in this very first example we were already able to get it to a place where it worked in an animal.
The other thing that I really want to highlight is that as excited as we are about Irresistin-16 in particular, we also think that this is a general concept that can be used. We think that this idea of a poisoned arrow approach could be applied to many other drugs, and so we hope that either we or others in the field will take this idea that has not really been implemented before, and use it to try to develop more types of new drugs.
Even if our shot-on goal with Irresistin-16 itself misses, our hope is that this will inspire new classes of drugs or research along similar lines to give us more shots at the goal.
How could this antibiotic revolutionize bacterial infection treatment and instigate the development of new drugs?
This is an interesting question for me (James) personally because I think that it is important to recognize that what goes into the paper is not everything that happened. You do not describe all the heartache or all the weird things that you did not quite understand.
Antibiotics have been around for the past 100 years or so, and there are some records that it has been around even longer - people just used it as a natural home-remedy, for example taking moldy bread, making a poultice out of it, and putting it in their wound.
People have been treating bacterial infections for a very long time, and so for me, when I think of this project I do not think about revolutionizing the way we treat the bacterial infections as much as revolutionizing the way we identify new antibiotics to treat those same infections.
It is the second half of the question that I think is the most important, and I think that what I, Zemer, Joe, Ben, and our other coauthors have demonstrated is two main things.
The first thing is that the membrane, which originally was a no-go target in antibiotic development, can be targeted in such a way that it predominantly hurts bacterial membranes more than it hurts human, and I think that opens up the possibility of a whole new class of molecules just by itself.
But I think the other thing that we have come upon is the idea that two-in-one is better than just two. There is a figure in the paper where we demonstrate that combining two antibiotics that have the same separate mechanisms of killing bacteria as SCH is not as powerful or as potent as SCH, and I do not think this is something that has been explored before.
The concept of combination treatment has been explored and all these complications have arisen, and so some people have pursued it and some people have not, but I think what we have demonstrated here is that maybe it is not as hard as we make it, maybe there is an easier way to get this molecule into the bacteria if we combine two separate mechanisms.
I am really excited to see some of the work that comes out of our lab, and others potentially if they pick it up, on doing this dual-mechanism approach. It does not have to be two mechanisms – there is no reason why it cannot be any number of targets. I think it could be really interesting to look at molecules with multiple modalities.
.jpg)
Image Credit: Orawan Pattarawimonchai/Shutterstock.com
What are the next steps for your research?
There are two arms. The first arm is improving the current drugs that we have to make them more and more drug-like and suitable, inching them towards usability as an antibiotic. That means making a lot of derivatives and modifications of them, and so on.
The second arm is: can we generalize these principles? Can we find new antibiotics that combine two mechanisms of action together? Can we make more poisoned arrows? Can we understand how exactly each of the two mechanisms is working?
A big surprise here on the scientific side was that we were able to target bacterial membranes without targeting mammalian membranes, and this specificity was a huge surprise to us.
We used to think a membrane was a membrane, who cares if it is bacterial or human? That is why historically we have avoided trying to target the membrane because we thought that we could not get specificity that way. However, this suggests that this is not the case. Perhaps we could even find ways of targeting viral membranes if that is true, so there are a lot of those sorts of questions that we are now trying to think about.
Have you got anything else you would like to add?
Actually, I do. I feel like in light of the last few weeks, we cannot just sit here and ignore the fact that historically, and in the US in particular, black scientists have been marginalized and underrepresented in every possible way.
Being a black man, a black male in America, is difficult for many different reasons, and trying to be a black man in STEM is even harder.
There is just not a lot of support. Often, there is this assumption that you are not as 'smart', as your peers, and so to be able to actually accomplish a piece of science like this and to be able to say "I discovered something," is really powerful.
These past few weeks in America, we look and there are many different young men who look like me, and I am actually very familiar with many of them in Chicago. Even though I may have all of these scientific accomplishments, in the end, it may mean nothing because in the eyes of many people, I'm black first and that sums up my entire being, complete with negative stereotypes and perceptions. For me, with this paper being published and thankfully getting media attention, I want it to be known that I do stand with those who look like me and with those who are oppressed or marginalized.
I do not want to send a message that I am different, or that I somehow have made it because of something I have accomplished, but instead that I am here, I hurt with you, and I want to use this platform that I have to help change people's perspective on what a black man should or could be.
Where can readers find more information?
Original Paper: https://www.cell.com/cell/pdf/S0092-8674(20)30567-5.pdf?_returnURL=https%3A%2F%2Flinkinghub.elsevier.com%2Fretrieve%2Fpii%2FS0092867420305675%3Fshowall%3Dtrue
A free, open-access pre-print of the paper: https://www.biorxiv.org/content/10.1101/2020.03.12.984229v1
CDC resource guide for antibiotic resistance: https://www.cdc.gov/drugresistance/index.html
About Dr. James Martin
My name is Dr. James Martin. I am a scientist and an educator and have a fierce passion for raising up the next generation of black scientists.
With all the unrest in America right now due to racial injustice, I am proud to say that I am black, and I hope that I can be an inspiration to others to continue pushing forward and be resilient.
About Professor Zemer Gitai
Zemer Gitai is a Professor of Molecular Biology at Princeton University. He graduated with a bachelor's degree from MIT in 1996. After completing his graduate studies at UCSF in 2002 and postdoc at Stanford in 2005, Dr. Gitai joined the faculty of Princeton University as an Assistant Professor.
He was promoted to Associate Professor with tenure in 2012 and a Full Professor in 2016. He is currently the Edwin Grant Conklin Professor of Biology in the Department of Molecular Biology at Princeton University.
Dr. Gitai's research focuses on the cell biology of bacteria and how they interact with their hosts. His lab studies how cells self-organize across spatial scales, using quantitative, molecular, and engineering approaches.
Dr. Gitai has published over 70 original articles in leading journals. His work discovered new components of the bacterial cytoskeleton and established the importance of protein assembly for unexpected processes like metabolism and pathogenesis.
Most recently, Dr. Gitai has focused on understanding bacterial mechanosensing and combating the rise of antibiotic resistance by developing new antibiotics and anti-infectives. Dr. Gitai's achievements have been recognized by many prestigious awards, including the the NIH Director’s Pioneer Award, the NIH New Innovator Award, the Beckman Young Investigator Award, and the HFSP Young Investigator Award.