The SARS-CoV-2 virus infects the human host cell via its interaction with the host cell membrane. The viral envelope is made up of spike protein as well as membrane (M) and envelope (E) proteins. The S protein is a prime target for vaccine development since it mediates viral binding to the host cell receptor, fusion of viral and host cell membranes leading to internalization of the virus, tissue tropism or specific infection of certain tissues, and the range of susceptible hosts.
Why Epitope-Based Vaccines?
The current paper discusses the development of an epitope-based vaccine. The advantages of this approach include the inclusion of a specific antigenic component that elicits an immune reaction by the host. Such vaccines have already been developed against rhinovirus, Dengue virus, and Chikungunya virus, among others.
However, epitopes in SARS-CoV-2 have not yet been identified. This is an RNA virus and thus shows a high mutation frequency. These are found to occur mostly at the spike glycoprotein, and typically increase viral fitness by enabling immune evasion. The S protein is capable of eliciting a more robust and rapid mucosal immunity than other viral proteins. The S proteins of SARS- and MERS-CoV have been used to develop effective vaccines against these viruses. The conserved region of the spike protein in SARS-CoV-2 presents an excellent target for an epitope-based vaccine.
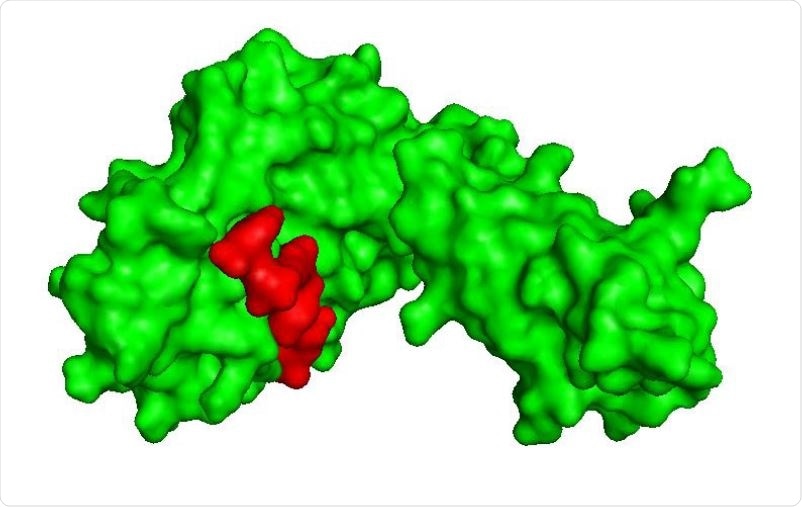
Shows surface interaction of GQTGKIADT T cell epitope (red) with HLA-C*03:03 (green).

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
T Cell Responses for Long-Term Immunity
Durable immunity is mostly due to adaptive T cell responses, especially the effector CD8 T cells, relative to B cell immunity, because once the viral antigens develop antigenic drift during serial passages, specific antibodies to the original antibodies become useless. The current study makes use of the imposing collection of computationally and experimentally derived database of epitopes.
The researchers found 23 T cell epitopes along with the specific MHC I molecules from the immune database, all from the spike protein of SARS-CoV-2. Of these, four epitopes meet the criteria for an effective vaccine candidate epitope; they are stable, strongly antigenic, non-toxic, and non-allergenic.
Identifying the Best Epitope
They confirmed that these epitopes bind with high affinity to the MHC I molecules, resulting in effective viral inhibition at lower doses. They identified the T cell epitope GQTGKIADY that performs the best, being able to bind effectively with HLA-C*03:03. The firmness of the binding is supported by molecular dynamics (MD) simulation results, which show that the GQTGKIADY-HLA-C*03:03 complex becomes stable in 40 ns.
The researchers went on to carry out a population coverage analysis, which showed that HLA-C*03:03 was prevalent in six countries badly hit by the virus: India, China, Italy, United States, United Kingdom, and Russia. This told them that the selected epitope would show effective binding to this prevalent HLA subtype, which is present at a high frequency in many of the most affected countries.
Prior research shows that many B cell epitopes have been identified that have different properties as well as the ability to induce antibody responses at different thresholds. However, the current study focuses on T cell epitopes since these are responsible for durable immunity.
Advantages and Implications
The study is an in silico study, with the actual data sourced from available immune databases. However, this type of study has been validated in earlier papers with wet-lab results. Not only does this save such laboratory resources, but it can help to design the right type of vaccine. The earlier results impart confidence that this T cell epitope could be used to elicit an effective immune response against SARS-CoV-2 infection.
The researchers sum up, “A novel T-cell epitope GQTGKIADY has been proposed as it contains all the features like antigenicity, non-allergenicity, non-toxicity, and stability. Further, GQTGKIADY epitopes binds with HLA-C*03:03 very well.” Its ability to bind to a common HLA antigen prevalent in many countries where the virus has been rampant shows that it could prove to be a very effective vaccine component, producing long-term immunity.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources