A group of researchers from Australia and China recently demonstrated how SARS-CoV-2 spike glycoprotein subunit 1 can induce a pro-inflammatory signaling pathway (leading to cytokines deployment and epithelial damage in human bronchial epithelial cells), but also highlighted the potential of peptide-based antivirals in tackling this novel coronavirus. Their paper is currently available on the bioRxiv* preprint server.
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) – a causative agent of the ongoing coronavirus disease (COVID-19) pandemic – is especially prone to infecting human bronchial epithelial cells that support its replication. Naturally, this prompts the innate immune response, which is critical in the early containment of the infection and subsequent viral spread.
Spike glycoprotein of SARS-CoV-2 interacts with host angiotensin-converting enzyme 2 (ACE2) as a key initial step to viral replication. From the pathophysiological perspective, the receptor-binding domain (RBD) of subunit 1 (S1) represents the pivotal region for binding to the aforementioned receptor.
This interaction kicks off heighten inflammatory response, which is primarily driven by early endoplasmic reticulum (ER) stress and its adaptive unfolded protein response (UPR), but also by activation of mitogen-activated protein (MAP) kinase signaling pathways.
More specifically, such early induction of ER-UPR results in activation of MAP kinase, and then both pathways synergistically lead to the activation of nuclear factor-κB (NF-κB) and production of pro-inflammatory cytokines such as interleukin-6, interleukin-1β, tumor necrosis factor-α (TNF-α) and C-C motif chemokine ligand 2 (CCL2) – the latter three contributing to acute lung injury.
At the moment, there are no specific antiviral or anti-inflammatory drugs available that can impact clinical outcomes in individuals with COVID-19. A broad-spectrum antiviral drug remdesivir is an option for compassionate use against COVID-19, as demonstrated by several different clinical. Nonetheless, we need more candidates that will hamper these specific inflammatory processes.
This research question was of significant interest to the research group from Australia and China, led by Dr. Alan C-Y Hsu from the University of Newcastle and Hunter Medical Research Institute in New South Wales, Australia.
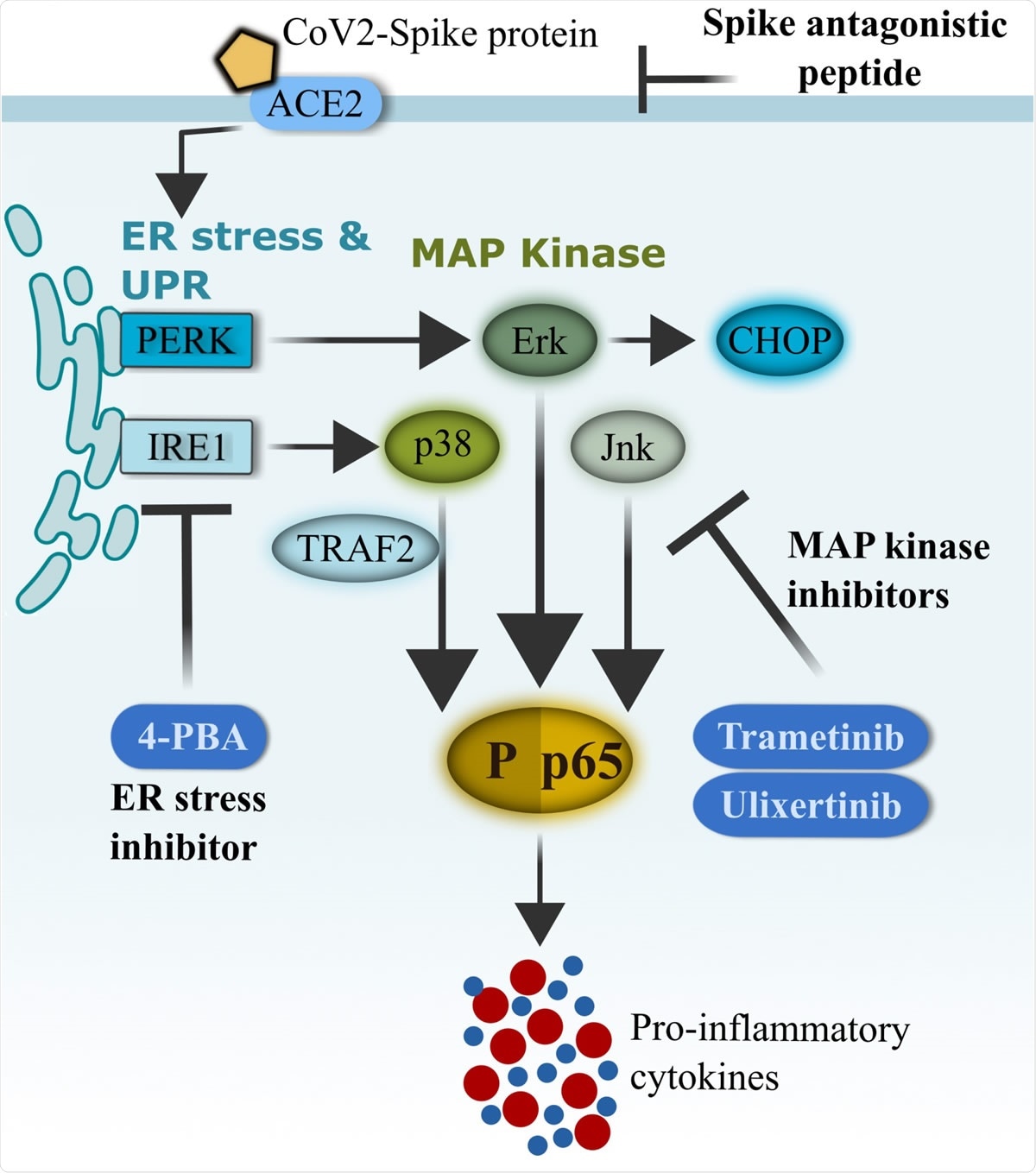
Schematics of CoV2-S1-mediated inflammation via ER-UPR and MAP kinase pathways. SARS-CoV-2 (CoV2) spike protein binds with ACE2 on the surface of human bronchial epithelial cells and rapidly facilitates the induction of ER stress and unfolded protein response (UPR). Activation of UPR (PERK and IRE1�) promotes the activation of MAP kinases, and the two pathways synergistically drive the activation of NF-κB and production of pro-inflammatory cytokines. FDA-approved ER-UPR inhibitor 4-phenylburic acid (4- PBA) and MAP kinase inhibitors (trametinib and ulixertinib) suppressed CoV2-S1-induced ER stress and MAP kinase activities, resulting in reduced NF-κB-mediated expression of pro inflammatory cytokines.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
Highly potent antagonistic peptides
Since SARS-CoV-2 S1 has been shown to induce NF-κB activation via its interaction with ACE2 and early activations of ER-UPR and MAP kinase signaling, these scientists designed a series of SARS-CoV-2-S1-antagonistic peptides – and identified a peptide (designated AP-6) that successfully inhibits this interaction.
In order to investigate whether SARS-CoV-2 spike glycoprotein induces the production of pro-inflammatory cytokines, they have utilized a minimally immortalized human bronchial epithelial cell line BCi-NS1.1, derived from human primary bronchial epithelial cells.
One of the main aims of this study was to assess if SARS-CoV-2 S1-mediated inflammation can be reduced by their SARS-CoV2-S1 inhibitory peptide AP-6, or with existing FDA-approved pharmacological inhibitors that target ER stress (4-phenylburic acid) or inhibit MAP kinase (trametinib and ulixertinib).
Tackling inflammatory storm and downstream consequences
In this study, the researchers confirm the notion that the SARS-CoV-2 S1 subunit and RBD instigate early ER-UPR and MAP kinase activations, subsequently leading to highly characteristic hyper-inflammatory response.
"Our data strong indicates that inflammation could be triggered by SARS-CoV-2 S1 even before viral replication occurs or in the absence of viral replication", explain study authors. "As SARS-CoV-2 S1 is present throughout viral replication cycles and infection, our data demonstrate that spike proteins are likely to be a major contributor to inflammation", they add.
Furthermore, their results imply that triggered inflammatory storm and downstream consequences can be successfully inhibited by S1-inhibitory peptides, as well as existing FDA-approved ER stress inhibitor 4-phenylbuic acid and MAP kinase inhibitors trametinib and ulixertinib.
Viable treatment implications
"Collectively, SARS-CoV-2 S1 induced heightened production of inflammatory cytokines that are primarily driven by MAP kinase and ER stress cross-talks", summarize study authors in their bioRxiv paper.
And AP-6 truly demonstrates the feasibility of this proof-of-concept antiviral strategy specific for certain coronaviruses. This is rather important, because inflammatory injury resulting from the activation of the described inflammation cascade may be the first step to the acute respiratory distress syndrome (ARDS).
Therefore, as antiviral drugs and vaccines are being developed and evaluated, existing FDA-approved ER stress and MAP kinase inhibitors could be deployed immediately in clinical trials as a potential treatment option for those with severe COVID-19.
Article source:

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
Journal reference:
- Preliminary scientific report.
Hsu, A.C.Y. et al. (2020). SARS-CoV-2 Spike protein promotes hyper-inflammatory response that can be ameliorated by Spike-antagonistic peptide and FDA-approved ER stress and MAP kinase inhibitors in vitro. bioRxiv. https://doi.org/10.1101/2020.09.30.317818.