A team of Brazilian scientists has recently conducted genomic and phylogenetic analyses of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) genome to understand the viral evolution process better. Their analyses reveal that the E484K mutated variant of SARS-CoV-2 is present in three different lineages in four Brazilian regions and that the mutation may serve as a common solution for viral evolution. The study is currently available on the bioRxiv* preprint server.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Background
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative pathogen of coronavirus disease 2019 (COVID-19), is a member of the Coronaviridae family, which are enveloped positive-sense, single-stranded RNA viruses with a larger genome size of about 30 kb. Since its emergence in December 2019 in China, researchers worldwide have sequenced almost 4,10,000 genomes of SARS-CoV-2 and have shared the information on the Global Initiative on Sharing All Influenza Data (GISAID) database.
Several genomic epidemiological studies of SARS-CoV-2 have been conducted, with the majority focusing on the mutations in viral spike protein, a surface glycoprotein of SARS-CoV-2 responsible for binding with the host angiotensin-converting enzyme 2 (ACE2) and initiating the viral entry process. Studies are currently mainly focusing on three viral lineages, B.1.1.7, B.1.351, and P.1. because of their significant impact on viral transmissibility and pathogenicity. Of these lineages, lineage P.1 is prevalent in Brazil with three mutations (K417T, E484K, and N501Y) in the spike receptor-binding domain (RBD). In Brazil, many viral genomes harbor E484K mutation, which is associated with increased viral transmissibility and immune evasion.
In the current study, the scientists evaluated the mutation characteristics and phylogenetic relationships in currently known lineages in Brazil to understand the adaptive evolution process of SARS-CoV-2.
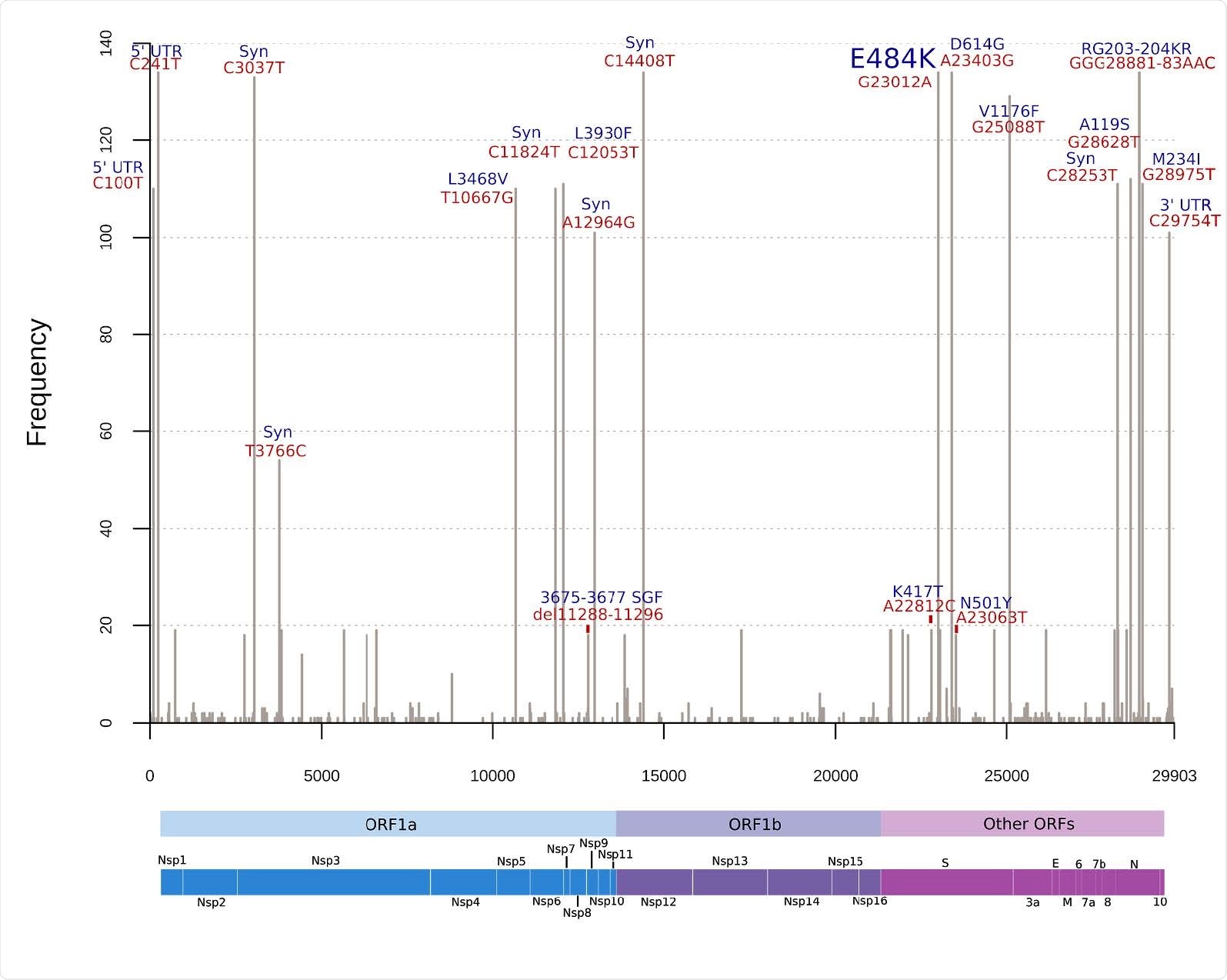
Histogram of frequent mutations observed in the Brazilian SARS-CoV-2 genomes harboring E484K mutation. Red labels above the bars indicate absolute nucleotide position and the blue labels indicate effects of these mutations in the corresponding proteins. As P.1 has only 19 genomes represented and multiple mutations, only main mutations of concern were highlighted. UTR: Untranslated region; Syn: Synonymous substitution; del: deletion; ORF: Open Reading Frame; Nsp: Non-structural protein; S: Spike; E: Envelope; M: Membrane; N: Nucleocapsid.
Important observations
The study findings revealed that SARS-CoV-2 variants with E484K mutation are widely distributed in many regions in Brazil. The mutation was introduced in Brazil in October 2020. Because the E484K mutation is found in different viral lineages simultaneously with other mutations, the scientists suggest that this particular amino acid substitution may act as a common driving force for viral evolution in different genetic variants of SARS-CoV-2. In the E484K mutation, a negatively charged amino acid (glutamic acid) is substituted with a positively charged amino acid (lysine). Thus, it can be expected that the mutation has a significant impact on viral sustainability and adaptive evolution.
The structural analysis revealed that a new site for ACE2 binding (amino acid 75) is generated because of the E484K mutation. This appears to create a significantly stronger interaction between ACE2 and the native binding site located at the RBD and ACE2 interface (amino acid 501).
A highly diverse range of genetic mutations was observed in all Brazilian lineages with E484K mutation. On average, about 19 and 30 mutations were observed in B.1.1.33 and P.1 lineages, respectively. Further genomic analysis of the most recently emerged lineage P.2 indicated that both P.1 and P.2 lineages are evolving rapidly and have been circulating in Brazil for a more extended period.
The analyses of new specific single substitutions were also carried out in the current study. One such mutation is A27V, present in the N-terminal domain (NTD) of the spike protein. Studies have suggested that mutations in the spike NTD can allosterically alter the spike RBD – ACE2 interaction dynamics and facilitate immune evasion. Moreover, evidence shows that viral lineages carrying E484K mutation are associated with increased escape from antibody-mediated neutralization. Although the actual impact of E484K and other substitutions on immune evasion is still not clear, the scientists suggest that E484K mutation is associated with increased viral fitness under natural selection pressure.
Study significance
Overall, the study indicates that the E484K mutation acts as a critical evolutionary event for different viral lineages in terms of increasing viral fitness. The E484K mutation could potentially increase the infectivity and immune evasion potency of SARS-CoV-2. According to the scientists, more studies investigating the effectiveness of anti-SARS-CoV-2 human sera against E484K mutated variants of SARS-CoV-2 are urgently needed.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources