There is a wide spectrum of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) mutated strains that have infected millions over the past several months. It is crucial to understand the mechanism by which these variants are impacted by antibodies produced by vaccines or previous infections.
 Study: Exposures to different SARS-CoV-2 spike variants elicit neutralizing antibody responses with differential specificity towards established and emerging strains. Image Credit: PHOTOCREO Michal Bednarek / Shutterstock.com
Study: Exposures to different SARS-CoV-2 spike variants elicit neutralizing antibody responses with differential specificity towards established and emerging strains. Image Credit: PHOTOCREO Michal Bednarek / Shutterstock.com

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
In a recent study published on the preprint server medRxiv*, researchers compared the neutralization responses against several SARS-CoV-2 variants of concern (VoCs), including the B.1.617.2 (Delta) strain, in sera from individuals exposed to variant infection, vaccination, or both.
Causes behind mutations and how they affect antibody responses
The mutations in the genes encoding for the spike protein in coronaviruses, which is the main viral determinant of cellular entry, are key areas of interest in research on SARS-CoV-2.
Several spike mutations are likely to have resulted from selective pressure acquired through constant evasion of host immunity and subsequently increased transmissibility. Several recent studies have concluded that sera from vaccinated and naturally infected individuals yield diminished neutralizing activity against certain variants, including the currently dominant circulating SARS-CoV-2 Delta variant.
Now that public vaccination campaigns have been initiated in many countries around the world, it is important to understand the efficacy of serum neutralization titers produced from these vaccines. This is an important correlate of real-world protective immunity.
Based on data acquired from previous mutant strains of SARS-CoV-2, the antibody responses elicited by exposure to the ancestral strain’s spike protein will be less effective at preventing future infection by strains like the Delta variant.
About the study
To address the effects of different mutations on previously inoculated individuals, the researchers of the current study collected serum from subjects who were previously infected with various SARS-CoV-2 variants include the B.1 (D614G mutation only), B.1.429 (Epsilon), P.2 (Zeta), and B.1.1.519 strains. The researchers also collected serum from messenger ribonucleic acid (mRNA) vaccine recipients who were previously infected with the B.1 ancestral spike lineage prior to vaccination, infected with B.1.429 prior to vaccination, or had no prior infection before getting the vaccines.
The researchers derived a quantitative comparison of antibody titers in the sera of people infected by these strains. The neutralization titer of each exposure serum cohort was compared individually against a panel of pseudoviruses, which are artificially engineered viruses, with the newer mutants.
Each pseudovirus represented a different exposure variant, including the B.1.351 (Beta), P.1 (Gamma), B.1.617, and B.1.617.2 (Delta) SARS-CoV-2 variants. The newer SARS-CoV-2 variants had one or more spike mutations in common with one of the exposure variants.
The researchers further demonstrated the effect of serial exposure to different spike mutations to better understand the cross-reactivity of neutralizing antibody responses. They crossed a panel of eight different spike protein mutations against serum pools elicited by seven different exposure variants.
Study findings
The researchers inferred that the neutralizing antibody responses were strongest against variants that had one or more spike mutations in common with that used to trigger the immune response in the first place, otherwise known as the exposure variants.
The most important observation of the current study was that the B.1.617.2 (Delta) was neutralized more effectively by serum elicited by prior exposure to two prior exposure variants of B.1.429 and B.1.1.519. These two variants had separate subsets of spike mutations similar to, overlapping with, mutations in B.1.617.2.
Individuals who were previously infected with the B.1.429 (Epsilon) variant and were subsequently vaccinated were found to elicit greater cross-neutralization of B.1.429 and B.1.617.2 (Delta) as compared to those who were vaccinated alone or had a prior infection with the D614G mutant strain followed by vaccination. This established the fact that exposure to multiple spike variants enhanced cross-neutralization and expanded the specificity of the vaccines.
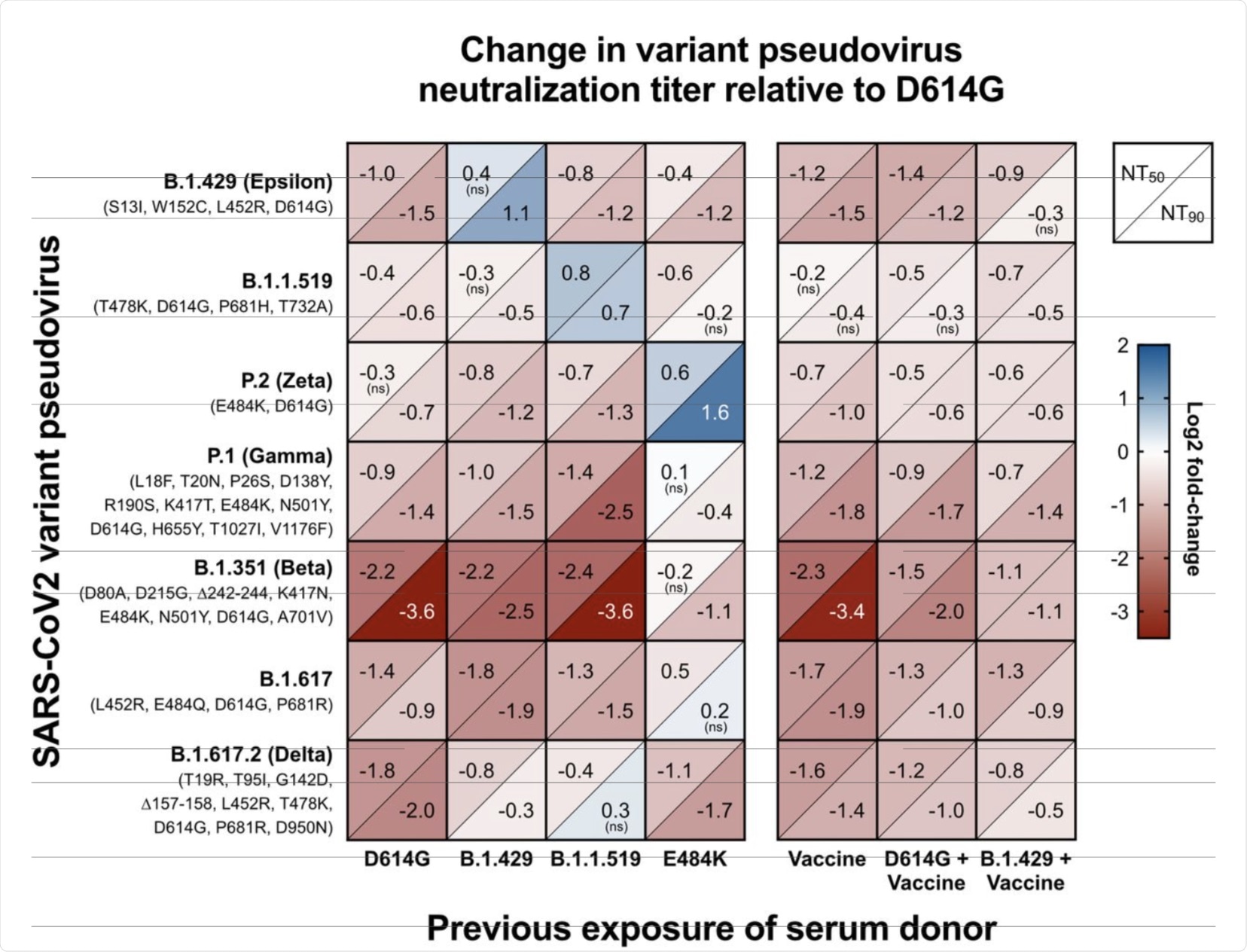 Matrix of normalized neutralization titers for seven different variant pseudoviruses (rows) neutralized by seven different pools of individual sera grouped by exposure (columns). Data is represented as a heat map of the Log2 fold-change in NT50 (top left of each box) and NT90 (bottom right of each box) of each variant relative to D614G pseudovirus. All serum samples were collected at least 14 days after the date of the subject’s positive COVID-19 test or date of second vaccine dose. All titer measurements are the mean of at least three independent experiments, each performed with two technical replicates. Positive Log2 fold-change (blue) indicates an increase in neutralization titer for that variant relative to D614G pseudovirus, while negative Log2 fold-change (red) indicates a decrease relative to D614G. Statistical significance was determined with unpaired t-tests. All values are statistically significant (P-value < 0.05) except where noted with “ns” to indicate the difference in variant neutralization titer is not significantly different from D614G titer in that serum pool.
Matrix of normalized neutralization titers for seven different variant pseudoviruses (rows) neutralized by seven different pools of individual sera grouped by exposure (columns). Data is represented as a heat map of the Log2 fold-change in NT50 (top left of each box) and NT90 (bottom right of each box) of each variant relative to D614G pseudovirus. All serum samples were collected at least 14 days after the date of the subject’s positive COVID-19 test or date of second vaccine dose. All titer measurements are the mean of at least three independent experiments, each performed with two technical replicates. Positive Log2 fold-change (blue) indicates an increase in neutralization titer for that variant relative to D614G pseudovirus, while negative Log2 fold-change (red) indicates a decrease relative to D614G. Statistical significance was determined with unpaired t-tests. All values are statistically significant (P-value < 0.05) except where noted with “ns” to indicate the difference in variant neutralization titer is not significantly different from D614G titer in that serum pool.
Taken together, the findings of the current study aid in understanding the relationship between exposures to different strains and antibody responses. Furthermore, these findings may also help in strategizing booster doses of vaccines.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Laurie, M.T., Liu, J., Sunshine, S., et al. (2021). Exposures to different SARS-CoV-2 spike variants elicit neutralizing antibody responses with differential specificity towards established and emerging strains. medRxiv. doi:10.1101/2021.09.08.21263095. https://www.medrxiv.org/content/10.1101/2021.09.08.21263095v1
- Peer reviewed and published scientific report.
Laurie, Matthew T, Jamin Liu, Sara Sunshine, James Peng, Douglas Black, Anthea M Mitchell, Sabrina A Mann, et al. 2022. “SARS-CoV-2 Variant Exposures Elicit Antibody Responses with Differential Cross-Neutralization of Established and Emerging Strains Including Delta and Omicron.” The Journal of Infectious Diseases 225 (11): 1909–14. https://doi.org/10.1093/infdis/jiab635. https://academic.oup.com/jid/article/225/11/1909/6494558.