The findings from recent studies suggested widespread SARS-CoV-2 infection in wild WTD populations in the United States. Thus, the high susceptibility of WTD to SARS-CoV-2 and their ability to transmit the virus to other deer emphasize the need for a better understanding of the infection mechanism and transmission dynamics of SARS-CoV-2 in this potential reservoir species.

Study: From Deer-to-Deer: SARS-CoV-2 is efficiently transmitted and presents broad tissue tropism and replication sites in white-tailed deer. Image Credit: Tom Reichner / Shutterstock.com
About the study
In the present study, the researchers assessed replication, shedding patterns, and transmission dynamics of SARS-CoV-2 in WTD. The study included 15 eight-month-old fawns that were randomly allocated to six experimental groups.
The three fawns categorized as a control group were un-inoculated, while six fawns were intranasally inoculated with SARS-CoV-2 and maintained in a separate room during the 22-day experimental period. The remaining six contact animals were allocated to three experimental groups that were maintained in pairs in separate rooms.
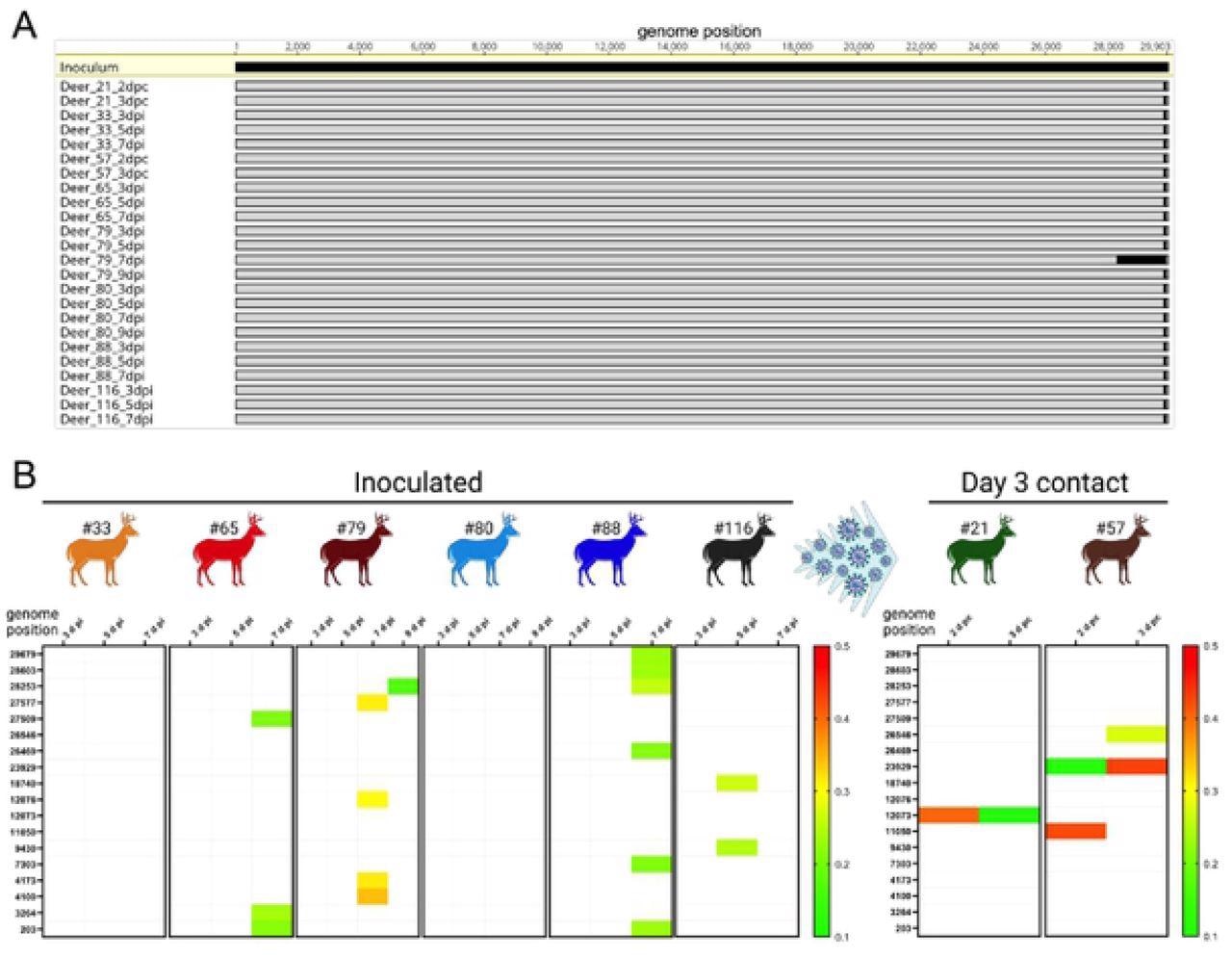 Low genetic diversity was observed following SARS-CoV-2 replication and transmission in white-tailed deer. Genome sequences were obtained directly from nasal secretions from all the 6 inoculated fawns collected on days 3, 5, 7, and/or 9 post-inoculation (pi) and from oronasal secretions from the day 3 contact animals (n = 2) collected on days 2 and 3 post-contact (pc). A) Whole-genome sequence analyses of SARS-CoV-2 sequences recovered from all inoculated and contact fawns revealed no amino acid differences in the consensus SARS-CoV-2 genome in comparison to the genome sequence of the inoculum virus isolate NYI67-20. B) Minor variant viral populations distributed throughout the virus genome from inoculated fawns on days 3, 5, 7, and/or 9 pi and from oronasal secretions from the day 3 contact animals (n = 2) collected on days 2 and 3 pc.
Low genetic diversity was observed following SARS-CoV-2 replication and transmission in white-tailed deer. Genome sequences were obtained directly from nasal secretions from all the 6 inoculated fawns collected on days 3, 5, 7, and/or 9 post-inoculation (pi) and from oronasal secretions from the day 3 contact animals (n = 2) collected on days 2 and 3 post-contact (pc). A) Whole-genome sequence analyses of SARS-CoV-2 sequences recovered from all inoculated and contact fawns revealed no amino acid differences in the consensus SARS-CoV-2 genome in comparison to the genome sequence of the inoculum virus isolate NYI67-20. B) Minor variant viral populations distributed throughout the virus genome from inoculated fawns on days 3, 5, 7, and/or 9 pi and from oronasal secretions from the day 3 contact animals (n = 2) collected on days 2 and 3 pc.
During the experimental period, all fawns were monitored daily for clinical signs and body temperature. The infection and replication dynamics of SARS-CoV-2 in WTD were monitored by assessing virus shedding post-inoculation.
Samples were collected in the form of nasal and oral secretions, as well as feces from inoculated and control fawns. The presence of SARS-CoV-2 in the samples was tested by real-time reverse transcriptase-polymerase chain reaction PCR (rRT-PCR) and by virus isolation and titrations.
The transmission dynamics of SARS-CoV-2 between WTD were analyzed by adding contact animals on days three, six, and nine post-inoculation. The researchers further investigated the potential changes in the genome of SARS-CoV-2 following replication in WTD by genome sequence analysis after infection and transmission to contact animals.
A second study in adult WTD was also conducted to gain insights into the sites of SARS-CoV-2 replication during acute infection. For this study, SARS-CoV-2 tissue tropism and replication sites were assessed by collecting 24 tissues and processed for rRT-PCR and infectious virus quantification. Tissue samples were processed for standard histological examination.
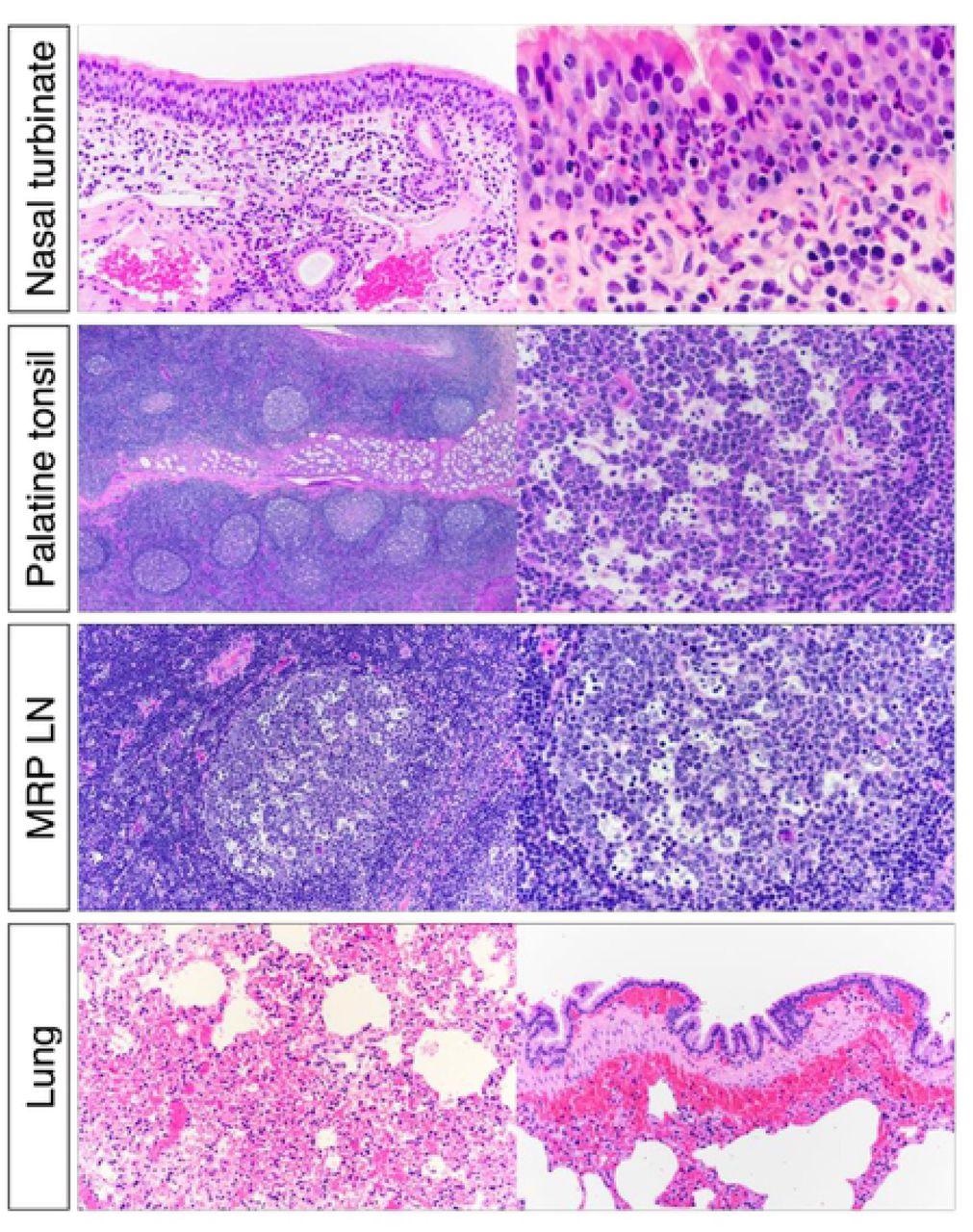 Histological examination of nasal turbinate from white-tailed deer inoculated with SARS-CoV-2. Upper respiratory tract (URT) after inoculation, there was submucosal lymphoplasmacytic infiltrate and frequent mucosal exudation of neutrophils in nasal turbinate. Numerous germinal centers of the palatine tonsils were characterized by lymphoid depletion and lymphocytolysis. Within the lung there was diffuse congestion with multifocal submucosal hemorrhages associated with larger bronchi.
Histological examination of nasal turbinate from white-tailed deer inoculated with SARS-CoV-2. Upper respiratory tract (URT) after inoculation, there was submucosal lymphoplasmacytic infiltrate and frequent mucosal exudation of neutrophils in nasal turbinate. Numerous germinal centers of the palatine tonsils were characterized by lymphoid depletion and lymphocytolysis. Within the lung there was diffuse congestion with multifocal submucosal hemorrhages associated with larger bronchi.
Study findings
The study findings suggested that the transmission of SARS-CoV-2 in WTD is likely to occur through respiratory or oral secretions. Interestingly, no infectious virus was recovered from feces, which further suggests that efficient transmission of the virus would require close contact between infected and susceptible animals. The absence of virus shedding in feces also indicated that the gastrointestinal tract is not a major target of SARS-CoV-2 in WTD.
No amino acid changes were noted in the SARS-CoV-2 consensus genome throughout the experimental period, thus suggesting low host-selective pressure(s) on the virus in WTD. The results demonstrated a broad tissue tropism of the virus, with active replication taking place in respiratory (nasal turbinate, trachea, bronchus, and lung), lymphoid (palatine and pharyngeal tonsil, retropharyngeal, mandibular and tracheobronchial lymph nodes and thymus), and central nervous tissues (olfactory lobes, cerebrum, caudate nucleus of brain and cerebellum).
High viral loads recovered from nasal turbinate and tonsils indicate that nasal turbinates and tonsils are likely the primary sites of viral replication during infection in WTD, which is similar to observations in other animal species and humans.
“This indicates that nasal turbinate and tonsil are likely the primary sites of viral replication during infection in WTD, which is similar to observations in other animal species and in humans.”
Conclusions
The current study discusses SARS-CoV-2 infection dynamics in WTD, an animal species with the potential to serve as a reservoir for the virus in the field. This study explained the short window of infectivity and transmissibility in WTD and demonstrated that the virus presents broad tissue tropism and multiple replication sites in this species.
In humans, infection with SARS-CoV-2 involves binding of the viral spike (S) protein to its host angiotensin-converting enzyme 2 (ACE2) receptor. The human ACE2 protein shares a high degree of homology with that of WTD.
The understanding of the infection and transmission dynamics of SARS-CoV-2 in WTD is critical to prevent future zoonotic transmission to humans and for the implementation of effective disease control measures.
*Important notice
bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information