The coronavirus disease 2019 (COVID-19), caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), leads to asymptomatic or mild symptoms in most cases. However, in a significant minority, it causes moderate or severe disease, with lung damage and sometimes multi-organ dysfunction, the origin of which is currently a matter of intense debate.
Moreover, some patients continue to have long-term symptoms, termed Post-Acute Sequelae of SARS-CoV-2 (PASC) or long-COVID. These may manifest in the brain, lungs, or cardiovascular system. However, there is not much clarity about the presence of the virus in extrapulmonary tissues, or the time required to clear the virus from the body, especially the brain.
Autopsies of non-survivors of COVID-19 have shown that the virus can infect multiple organs. Still, there is little evidence of direct viral injury or inflammatory injury secondary to viral infection.
A new study aimed to answer some of these questions from detailed autopsy samples from multiple organs and tissues in fatal cases of COVID-19. The research is released prior to peer review on the Research Square* preprint server.
Study: SARS-CoV-2 infection and persistence throughout the human body and brain

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
How Was the Study Done?
The scientists carried out autopsies on 44 patients to determine how the virus was distributed across body tissues and cell types; whether the virus was replicated in the tissues where it was found to be present; which types of cells were infected, and how long the virus persisted.
The methods they used included droplet digital polymerase chain reaction (ddPCR), a sensitive test for the quantitative detection of the gene products of the virus in tissue samples; in situ hybridization (ISH) to confirm the above results; immunohistochemistry (IHC) in brain samples to confirm the presence of the virus; detection of subgenomic ribonucleic acid (sgRNA) to detect recent viral replication in tissues that showed evidence of viral RNA, and virus isolation in cell culture to confirm that replication-competent virions were present in tissues outside the lung.
Finally, they used high-throughput, single-genome amplification, and sequencing (HT-SGS), a method of sequencing the viral genomes to help detect the presence and distribution of quasispecies within a single infected individual. This was carried out on six individuals.
What Did the Study Show?
The investigators detected the presence of viral RNA in all cases and in 79 of 85 organs or tissues or body fluids sampled. The respiratory tract, as expected, showed the highest viral load, with more than 100,000 Nucleocapsid (N) gene copies/ng RNA input, in individuals with early COVID-19. However, except for the reproductive organs, every sampled tissue group from several individuals had 100 or more N gene copies.
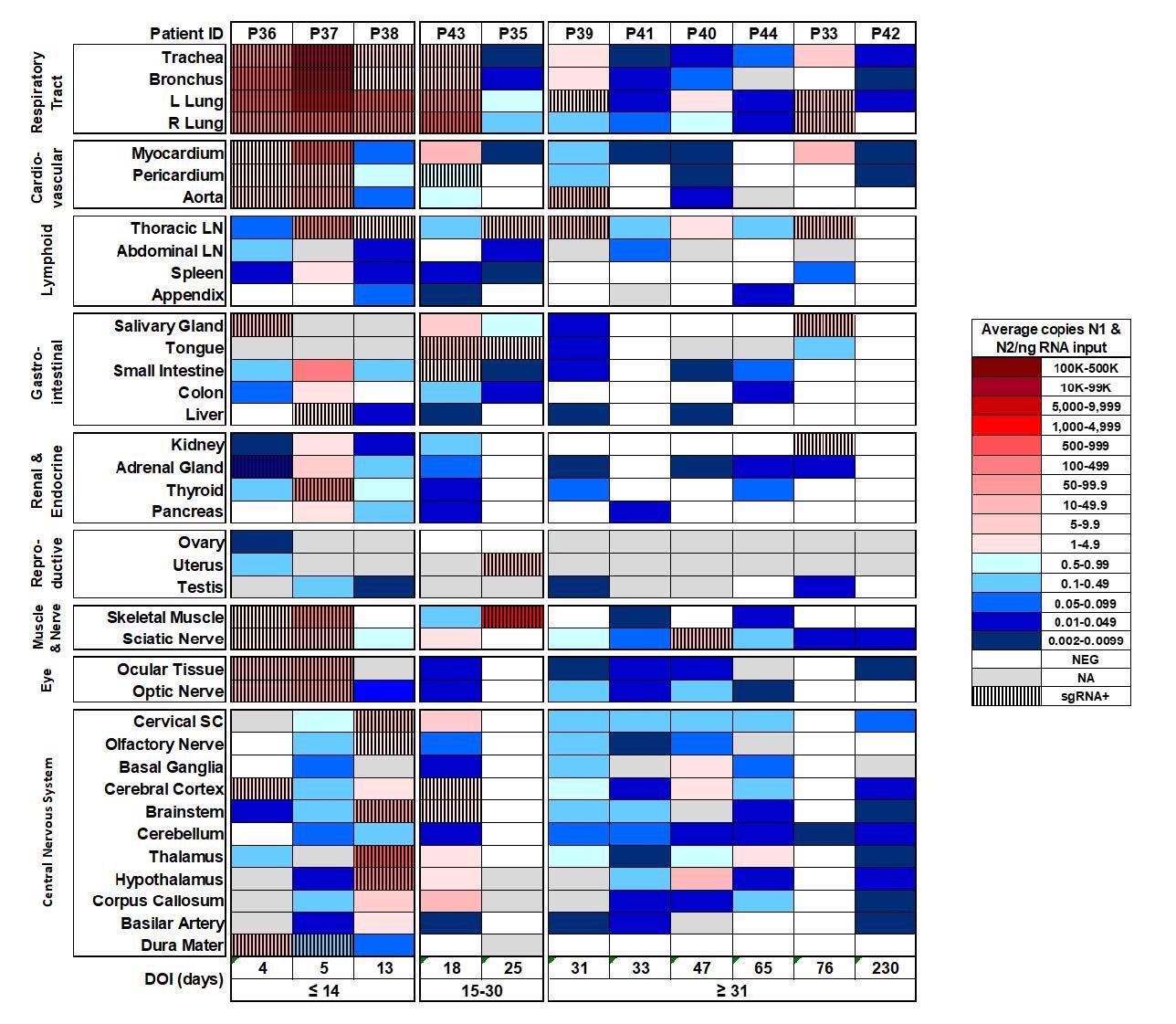
Distribution, quantification, and replication of SARS-Cov-2 across the human body and brain. The heat map depicts the highest mean quantification of SARS-CoV-2 RNA (N) via ddPCR present within the tissues of eleven COVID-19 autopsy patients who underwent whole body and brain sampling. Patients are aligned from shortest to longest duration of illness (DOI) prior to death, listed at the bottom of the figure, and grouped into early (≤14 days), mid (15-30 days), and late (≥31 days) DOI. Tissues are grouped by tissue category beginning with the respiratory tract at the top and central nervous system at the bottom. Viral RNA levels range from 0.002 to 500,000 N gene copies per ng of RNA input, depicted as a gradient from dark blue at the lowest level to dark red at the highest level. Tissues that were also positive for sgRNA via real-time RT-PCR are shaded with black vertical bars. L/left, LN/lymph node, NA/not acquired, R/right, SC/spinal cord.
The viral load fell by a log or more among those with mid-range COVID-19 and still more with late disease. Yet there were several cases with 5 or more N gene copies/ng RNA input in multiple tissues. Interestingly, in late cases, some individuals with undetectable viral RNA in plasma still showed low-level viral RNA in multiple tissues.
All the six individuals who had late COVID-19 were found to have viral RNA in brain tissue, and in 5/6 it was present at almost every sampled location – including one patient who died and was therefore sampled at 230 days from infection.
Almost 98% showed viral RNA in the respiratory tissue, and high proportions (>85%) also had it in lymphoid tissue and brain tissue. In over 70%, cardiovascular and gastrointestinal tissue showed the presence of viral RNA, and ~60% or more of renal, endocrine, muscle, skin, fatty tissue and peripheral nervous tissue. In ~58%, the eyes contained viral RNA.
The search for viral sgRNA, indicative of recent viral replication, was found in multiple tissues in over 80% of early cases. This was also found in the blood, pleural fluid and the vitreous humor of the eyes. Among mid- and late-persistence cases, it was found in one or more tissues in two-thirds and 42%, respectively.
The detection of sgRNA in half or more of 270 cases beyond the 14th day from symptom onset indicates the occurrence of prolonged viral replication up to day 99. This could not be due to either contamination by blood, or by lung tissue, since only 12 cases had detectable viral RNA in the blood. Of these 12, only two, both with early disease, also had sgRNA in plasma, with higher cycle threshold levels than in the tissues.
The presence of sgRNA was confirmed by ISH, which showed the extensive presence of the virus across the different sampled tissues. In addition, IHC demonstrated viral proteins in the brain samples.
Viral isolation was performed successfully from samples from multiple tissues from six individuals. Sequencing showed that in five early cases, the spike sequences dominant in each tissue were identical with each other, but with differences in the minor isolates. The scientists concluded that the virus could infect a large variety of cells without requiring different receptors to spread and replicate outside the lung.
Yet, variants were found in the spike protein of the virus in the brain samples, with minor brain variants showing the G1219V residue, while no brain sample out of nearly 500 showed the presence of D80F, found in all 31 lung samples. This suggests that different genetic variants of the virus reveal compartmentalization in their propensity to infect different tissues while also confirming the actual widespread presence of the virus in multiple tissues.
The results, therefore, confirm the ability of the virus to establish replicative infection in all these tissues. The virus was also found within white cells at multiple lymphoid sites, and in the colonic mucosa – either due to phagocytosis or actual infection of tissue-resident macrophages.
What Are the Implications?
The results showed that the virus is present in most body tissues, with replication at these sites rather than spillover effects, as demonstrated using very sensitive and accurate methods, including single-copy sequencing. The very few intra-individual genetic variants of the infection are attributable to a low intrinsic mutation rate and the fact that there is little immune selection pressure in early infection.
This may also explain why genetic compartmentalization was not seen in all individuals. The presence of a differential selection pressure operating in the brain due to antiviral antibodies may be suggested by the clear difference in the spike sequence in several minor brain variants but not other tissues, but more study will be required to validate this.
While the highest viral load is in the respiratory tract, early dissemination and widespread infection appear to be possible. This seems to be the effect of viremia, occurring early in infection, and causing viral seeding throughout the body. Another study has produced findings that support this conclusion as well.
Extensive seeding and replication of the virus can co-exist with asymptomatic or mild COVID-19, as seen in this cohort. Moreover, the virus is seen to be able to persist for months, mainly in the respiratory tract, but in more than half the late cases, also in the heart muscle, chest lymph nodes, peripheral nerves, the eye, and the brain.
The viral load was much lower in late cases, but both pulmonary and extrapulmonary tissues showed similar viral burdens, indicating that the latter was poor at clearing the virus. This could be due to a lower level of immune response to the virus in these tissues compared to the respiratory tract.
The late persistence of viral RNA and sgRNA could indicate that the infection was caused by a defective virus, as in persistent measles leading to subacute sclerosing panencephalitis. PASC has been postulated to be caused by local and systemic inflammation in response to the virus. However, the findings of this study do not support this with no signs of significant inflammation outside the respiratory tract even months after the primary infection.
Further research into the way the virus can sustain replicative infection and the host cell's response to viral persistence will be necessary in understanding how PASC occurs and can be managed.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Chertow, D. et al. (2021). SARS-CoV-2 Infection and Persistence Throughout the Human Body and Brain. DOI: https://doi.org/10.21203/rs.3.rs-1139035/v1. https://assets.researchsquare.com/files/rs-1139035/v1_covered.pdf?c=1640020576
- Peer reviewed and published scientific report.
Stein, Sydney R., Sabrina C. Ramelli, Alison Grazioli, Joon-Yong Chung, Manmeet Singh, Claude Kwe Yinda, Clayton W. Winkler, et al. 2022. “SARS-CoV-2 Infection and Persistence in the Human Body and Brain at Autopsy.” Nature, December, 1–6. https://doi.org/10.1038/s41586-022-05542-y. https://www.nature.com/articles/s41586-022-05542-y.