The World Health Organization (WHO) has reported over 361 million confirmed coronavirus disease 2019 (COVID-19) cases, of which include over 5.62 million deaths as of January 2022.
Several vaccines that have been developed against the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), have been successful in reducing the severity and hospitalizations associated with COVID-19. However, infection rates continue to rise due to the emergence of several SARS-CoV-2 variants of concern, as well as in unvaccinated regions.
Study: Longitudinal Assessment of SARS-CoV-2 Specific T Cell Cytokine-Producing Responses for 1 Year Reveals Persistence of Multi-Cytokine Proliferative Responses, with Greater Immunity Associated with Disease Severity. Image Credit: Merlin74 / Shutterstock.com

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Background
Neutralizing antibodies against the SARS-CoV-2 spike (S) protein have been reported to protect against infection and disease. However, some levels of protection have been observed in vaccinated individuals before the development of antibodies.
Moreover, individuals with other betacoronavirus infections who did not possess cross-reactive antibodies were found to have limited disease with SARS-CoV-2. This suggests the presence of other mediators of protection aside from antibodies.
T-cells are known to be important for both disease protection and viral clearance. CD4+ T-cells can enhance antibody maturation, while CD8+ T-cells can mediate the killing of infected cells.
For SARS-CoV-2 infections, T-cell responses have been found to be induced during convalescence and persist for up to eight months after infection. However, a comparison between the role of CD4+ and CD8+ T-cell memory in disease and protection has yet to be fully elucidated. Furthermore, the importance of various T-cell effector cytokines, along with the duration for which T-cell memory persists, remains unclear.
A new study published on the bioRxiv* preprint server assesses SARS-CoV-2 antigen-specific cytokine responses for about one year of follow-up post-symptom onset in a cohort of SARS-CoV-2 infected individuals with varying levels of disease outcomes. The study also mapped epitopes to SARS-CoV-2 structural proteins to determine effector function and killing capabilities of both CD4+ and CD8+ T-cell clones.
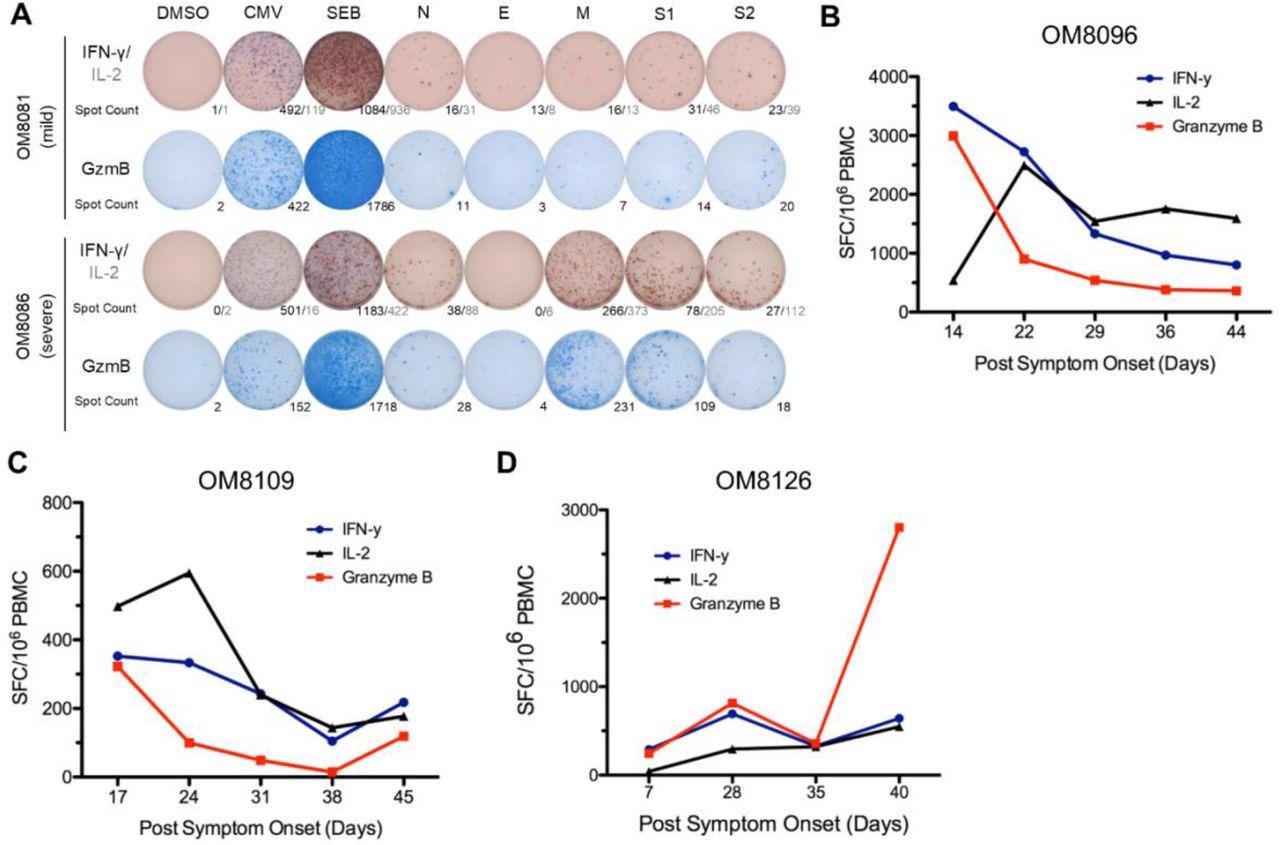
Stronger overall T cell cytokine ELISpot responses detected in severe patients and overall ELISpot responses of acute patients in the first six weeks PSO. (A) Representative cytokine ELISpot responses from individuals with mild or severe illness against SARS-CoV-2 structural proteins (N, E, M, S1 & S2), with DMSO as negative control and CMV and SEB as positive controls. Numbers indicate the number of spot-forming cells for IFN-γ/IL-2 or GzmB. (B)-(D) Three subjects with acute moderate COVID-19 infection were followed weekly shortly after symptom onset for up to 6 weeks PSO. Total additive response to the four SARS-CoV-2 structural proteins was measured by ELISpot for each cytokine.
About the study
The current study included both healthy and SARS-CoV-2-infected individuals. Blood samples were collected from all participants followed by isolation of peripheral blood mononuclear cells (PBMCs).
SARS-CoV-2 15-mer peptides overlapping by 11 amino acids from the four main structural proteins of the SARS-CoV-2 for T-cell epitope mapping were synthesized, along with the generation of matrix peptide pools containing 19-23 15-mers.
Ex vivo ELISpot assays were carried out using human interferon γ (IFN-γ), interleukin-2 (IL-2), and granzyme B (GzmB) antibodies. This was followed by the generation of immortalized B-cell lines and SARS-CoV-2-specific CD4+/CD8+ T-cell clones and lines.
Thereafter, the human leukocyte antigen (HLA) restriction of T-cell clones and lines was determined. Finally, carboxyfluorescein diacetate succinimidyl ester (CFSE) T-cell proliferation assay and cytotoxic T-lymphocyte (CTL) killing assay were performed.
Study findings
Out of the 23 SARS-CoV-2 infected individuals, 14, 6, and 3 experienced mild, moderate, and severe COVID-19 disease, respectively. Two subjects received one dose of Pfizer BNT162b2 COVID-19 messenger ribonucleic acid (mRNA) vaccine, while one received two doses of the Pfizer vaccine in between the study.
Three hospitalized individuals were sampled at seven to ten-day intervals during and after their hospital admission for six weeks, while the remaining individuals were sampled during their convalescence.
Considerable dynamic fluctuations in early T-cell cytokine response to SARS-CoV-2 specific antigens were reported in the three hospitalized individuals. Peak T-cell response was observed in the first 22 to 28 days, after which it reached a constant level. T-cell responses were also observed against the SARS-CoV-2 structural proteins in infected individuals.
About 33% of subjects responded to the nucleocapsid (N) protein, 29% responded to the S1 protein, 24% responded to the S2 protein, and 19% responded to the membrane (M) protein. No response was observed for the envelope (E) protein.
Individuals with moderate and severe disease were found to have higher frequencies of cytokine-producing SARS-CoV-2-specific T-cells as compared to those with mild disease. Most of the infected individuals showed GzmB responses as their strongest cytokine response that was mainly against N and S1/S2.
The majority of the cytokine response was contributed by CD4+ T-cells. Moreover, 50% of the uninfected and pre-COVID-19 individuals showed cross-reactive responses to SARS-CoV-2 proteins, along with low levels of cytokine responses.
The cytokine responses were found to decline over one year for all SARS-CoV-2 antigens. Among the three individuals who received vaccination during the follow-up period, one showed increased responses to the spike but not N and M protein post-vaccination, one showed continued decay of responses even after vaccination, and one exhibited only increased IFN-γ responses to S1 but no further changes against the other proteins.
The researchers also reported that the immune response variables peaked at 36 days for IL-2, six days for IFN-γ, and seven days for GzmB. The study also predicted that severe disease was associated with a reduced IFN-γ and GzmB, but increased IL-2 production rates.
The cytokine responses were found to be different based on sex. To this end, faster stimulation and slower degradation rates were observed for IFN-γ in males as compared to females, while the opposite was observed in the case of GzmB.
Stimulation and degradation rates were observed to be similar for IL-2. Additionally, the half-life of CD4+ T-cells was found to be 139 days.
Cross-reactive T-cell proliferative responses were observed in both infected and healthy individuals, although it was weaker in healthy individuals. These responses from infected individuals were found to be driven mostly by CD4+ T-cells. The results further indicated that the convalescent individuals developed a T-cell memory response against SARS-CoV-2.
The study also identified 35 T-cell epitopes that belonged to open reading frame 1 ab (ORF1ab) (1 epitope), N (13 epitopes), M (14 epitopes), and S (7 epitopes). The strongest response was reported from the M epitopes followed by N, S, and ORF1ab, while no response to E was observed.
The majority of epitopes were found to favor CD4+ T-cell responses over CD8+ T-cell responses. Other epitopes common to the alpha and beta coronavirus families included the M154-156 region, N76-N88 region, and the M160-163 region of the ORF1ab. Moreover, different T-cell clones with specific epitopes were observed to have different HLA restrictions.
The SARS-CoV-2-specific CD4+ and CD8+ T-cell clones were also found to kill peptide-pulsed autologous B-cell line target cells. SARS-CoV-2 CD8+ T-cell clones with lower effector to target ratios were found to possess high cytotoxic capabilities as compared to SARS-CoV-2 CD4+ T-cell clones. Moreover, both the T-cell clones elicited strong IFN-γ and GzmB responses on stimulation with their peptide of interest.
Conclusions
Taken together, the current study demonstrates the presence of SARS-CoV-2-specific memory T-cells, even after one year from symptom onset. These cells were found to generate IFN-γ, IL-2, and GzmB responses against SARS-CoV-2 structural proteins, which might influence the disease course in case of re-infection with a variant.
The presence of these T-cells might also explain the superior responses to vaccines. However, further research needs to be carried out for the development of T-cell vaccines against SARS-CoV-2.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Lin, J., Law, R., Korosec, C. S., et al. (2022). Longitudinal Assessment of SARS-CoV-2 Specific T Cell Cytokine-Producing Responses for 1 Year Reveals Persistence of Multi-Cytokine Proliferative Responses, with Greater Immunity Associated with Disease Severity. bioRxiv. doi:10.1101/2022.01.18.476864. https://www.biorxiv.org/content/10.1101/2022.01.18.476864v1.
- Peer reviewed and published scientific report.
Lin, Jonah, Ryan Law, Chapin S. Korosec, Christine Zhou, Wan Hon Koh, Mohammad Sajjad Ghaemi, Philip Samaan, et al. 2022. “Longitudinal Assessment of SARS-CoV-2-Specific T Cell Cytokine-Producing Responses for 1 Year Reveals Persistence of Multicytokine Proliferative Responses, with Greater Immunity Associated with Disease Severity.” Edited by Mark T. Heise. Journal of Virology 96 (13). https://doi.org/10.1128/jvi.00509-22. https://journals.asm.org/doi/10.1128/jvi.00509-22.