New high-throughput protein profiling tools are opening fresh possibilities for precision medicine, but this review shows why clinical laboratories must balance innovation with rigorous measurement, validation, and real-world diagnostic reliability.
Review: Next-generation proteomics in medical laboratories: metrologically sound quantitative protein tests vs. innovative and personalized proteome patterns. Image Credit: Shutterstock
A recent review published in the journal Clinical Chemistry and Laboratory Medicine (CCLM) highlights a key dilemma in clinical proteomics: whether to prioritize precise, metrologically robust protein quantification or adopt high-throughput profiling approaches. While affinity-based assays using deoxyribonucleic acid (DNA) barcoding and next-generation sequencing (NGS) enable scalable multi-omics integration, mass spectrometry (MS) remains unmatched at resolving specific proteins and their proteoforms. Balancing analytical rigor with innovation will be critical to advancing precision medicine.
In the era of precision medicine, researchers are leveraging protein phenotypes and advanced analytical tools to improve diagnostic accuracy and support patient management. Advances in scalable proteomic platforms have renewed focus on plasma proteomics, enabling faster, large-scale profiling. Designed to integrate with machine learning, these technologies support population-level analyses and broader insights. While conventional immunoassays and targeted MS remain central to clinical practice, emerging methods, including aptamer-based assays, ultrahigh-resolution MS, and activity-based nanosensors, are driving more comprehensive proteomic profiling, although many remain primarily discovery tools rather than routine clinical diagnostics.
In this review, Yuri E.M. van der Burgt and Christa M. Cobbaert of the Leiden University Medical Center, The Netherlands, examine the role of proteomics in precision medicine, outlining established clinical technologies alongside emerging high-sensitivity, high-throughput platforms that may help shape future diagnostics.
Current Proteomic Tools in Precision Medicine
Precision medicine increasingly relies on molecular characterization to uncover disease pathways and tailor therapies to individual patients rather than population averages. While genomics has long served as a cornerstone for assessing disease risk, its limited ability to predict downstream transcriptomic and proteomic states has driven the integration of complementary omics approaches. Among these, proteomics provides a direct readout of physiological status, with protein signatures reflecting dynamic biological processes and treatment responses.
MS-based proteomics has played a central role in linking molecular phenotypes to clinical outcomes, enabling systems-level insights into human biology. Targeted MS and conventional antibody-based assays, including enzyme-linked immunosorbent assays (ELISA) and multiplex platforms such as Luminex, remain widely used in clinical laboratories due to their sensitivity and reproducibility. However, these approaches face notable limitations. Antibody-based assays are constrained by cross-reactivity, limited multiplexing, and an inability to resolve distinct proteoforms, while MS-based methods, despite their analytical precision, require complex workflows, extensive validation, and remain low throughput.
Moreover, the translation of proteomic biomarkers into routine clinical practice has been slower than anticipated. This reflects not only technical challenges but also systemic barriers, including lengthy validation processes and limited incentives for clinical implementation. Importantly, emerging evidence suggests that single biomarkers are insufficient, with multi-protein panels, and increasingly, proteoform-resolved data, offering greater diagnostic value. Together, these challenges highlight the need for more scalable, efficient proteomic platforms to fully realize the potential of proteomics in precision medicine.
Emerging High-Throughput Proteomics Technologies
To address the limitations of conventional approaches, proteomics is advancing toward scalable, high-capacity, systems-level analysis. Affinity-based technologies integrated with NGS have emerged as powerful tools for protein profiling.
For example, proximity extension assays such as those developed by Olink use paired antibodies linked to DNA oligonucleotides that generate unique barcodes upon target binding, enabling the simultaneous measurement of thousands of proteins with high sensitivity and dynamic range. Similarly, aptamer-based platforms like SomaLogic employ engineered SOMAmer reagents to capture proteins based on structural complementarity, producing multiplexed proteomic readouts that can be more readily integrated with genomic and transcriptomic data.
These platforms offer clear advantages over traditional immunoassays and MS, including greater throughput, improved scalability, and simplified data outputs that readily support machine learning-driven pattern recognition. As a result, they are particularly well suited for population-scale studies and biomarker discovery. However, unlike MS-based methods, binding-based approaches typically measure relative protein levels, often reflect mixtures of proteoforms rather than precisely defined measurands, and do not resolve distinct proteoforms, which may limit biological interpretation. Their performance may also be influenced by other binding partners or autoantibodies.
Beyond affinity approaches, emerging strategies such as protease-activated nanosensors introduce a functional dimension, detecting enzyme activity in vivo. These systems release mass-encoded reporters measurable by targeted MS, enabling sensitive detection of disease-associated protease activity. Although still in early development, such innovations highlight a shift toward dynamic, activity-based proteomics. Collectively, these advances are expanding the scope of precision medicine, while also underscoring the need for careful validation, standardization, and scrutiny of pre-analytical and assay-specific limitations before widespread clinical adoption.
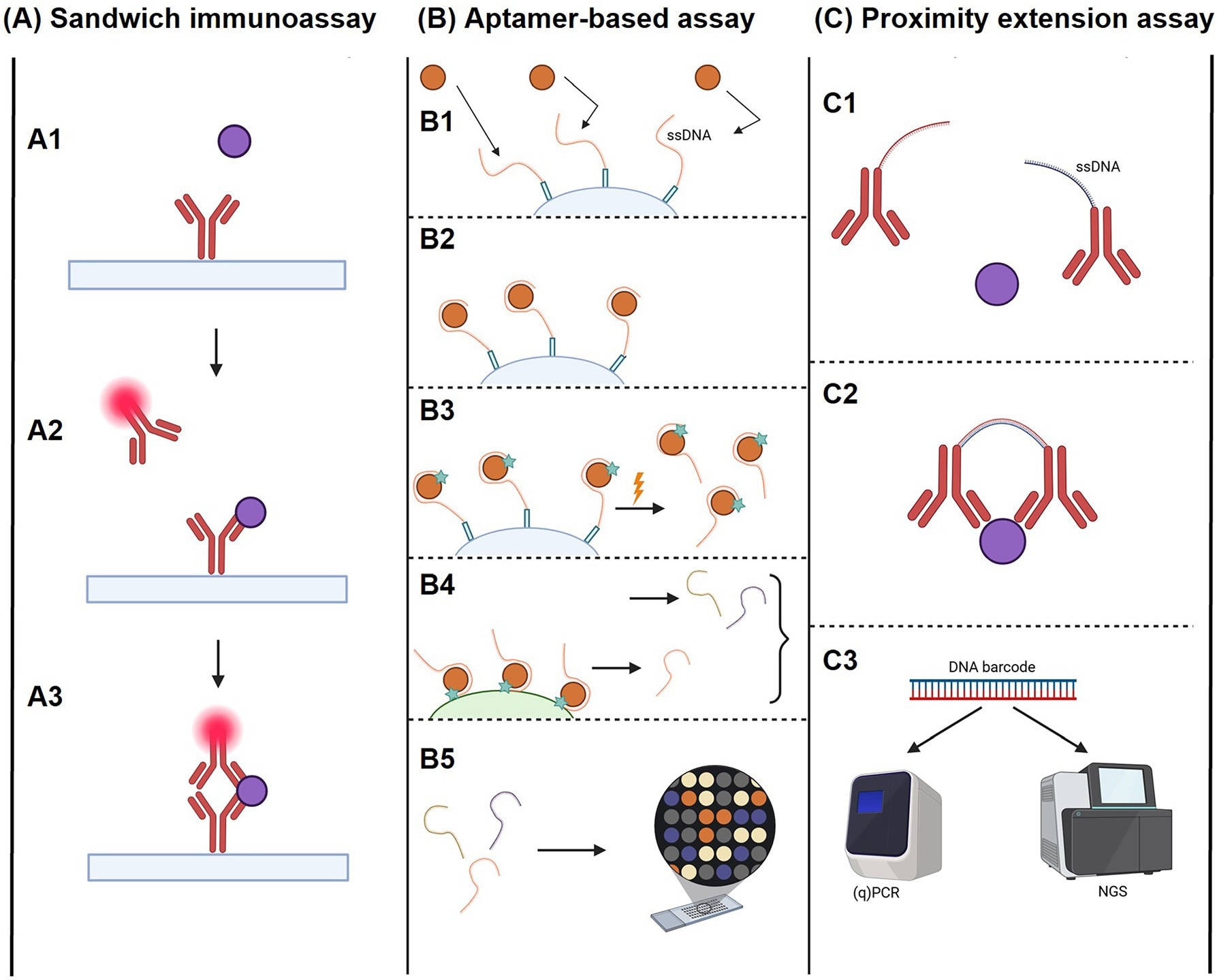
High-throughput affinity-based assays for proteome analyses.
Future Clinical Proteomics Directions
Based on the findings, protein diagnostics now stands at a pivotal crossroads, balancing the precision of quantitative assays with the promise of personalized proteome profiling powered by machine learning. While validated biomarkers will remain essential for many clinical applications, growing evidence suggests that subtle, longitudinal changes in proteotypes may inform earlier, more tailored interventions. A hybrid approach, or intermediate path, integrating targeted, metrologically robust assays with high-dimensional proteomic patterns, may offer the most practical path forward.
Advances in MS, including intact protein analysis, alongside scalable affinity-based platforms, are expanding the diagnostic toolkit. However, translating these innovations into routine care will require rigorous standardization, reproducibility, robustness, and quality assurance. As proteomics converges with other omics and digital health strategies, it offers an opportunity to enhance disease detection, enable proactive health monitoring, and support a more sustainable and personalized healthcare system.