A group of over 120 researchers from various institutions across Europe has performed the first genome-wide association study to reveal host genetic factors that may contribute to respiratory failure in cases of coronavirus disease 209 (COVID-19).
The authors say the genetic variants they have identified could help guide further research into the pathophysiology of COVID-19 and aid the clinical risk profiling of patients.
A pre-print version of the paper is available on the server medRxiv*, while the article undergoes peer review.
The rapid spread of the pandemic
Since the COVID-19 outbreak began in Wuhan, China, late last year, it has rapidly become a pandemic health emergency that has now infected more than 6.39 million people worldwide and killed almost 400,000.
In Europe, Italy and Spain have been among the most severely affected countries, with epidemics peaking during the second half of February and more than 60,000 fatal cases being reported by May 28th.
Most people infected with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) – the causative agent of COVID-19 – only experience mild or even no symptoms.
Mortality rates are mainly driven by patients who are more susceptible to respiratory failure after becoming ill with pneumonia or respiratory distress syndrome. However, for reasons that are not properly understood, this is only the case for less than 10 percent of people who become infected with SARS-CoV-2.
Novel Coronavirus SARS-CoV-2 Colorized scanning electron micrograph of an apoptotic cell (pink) heavily infected with SARS-COV-2 virus particles (green), isolated from a patient sample. Image captured at the NIAID Integrated Research Facility (IRF) in Fort Detrick, Maryland. Credit: NIAID

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Potential factors involved in respiratory failure
The development of severe disease has also been associated with the presence of comorbidities such as cardiovascular disease, obesity, diabetes, and hypertension. However, the role these health problems play in determining the severity of disease risk is unclear.
Some observations of endothelitis and vascular complications have suggested that the disease is systemic and mainly involves the vascular endothelium. Still, these insights into the pathology of severe COVID-19 are only hypothetical.
Performing a genome-wide analysis
To investigate, Tom Karlsen (Oslo University Hospital Rikshospitalet, Norway) and colleagues in Spain, Italy, and Germany, recruited 1,980 COVID-19 patients with respiratory failure from five cities in Spain and Italy.
They conducted a genome-wide association analysis with the aim of identifying any host genetic susceptibility factors that contribute to the development of respiratory failure.
“Using a pragmatic approach with simplified inclusion criteria and a complementary team of clinicians at the European Covid-19 epicenters in Italy and Spain and available German and Norwegian scientists, we were able to perform a complete GWAS for Covid-19 respiratory failure in about two months,” say the researchers.
After considering quality control and potential outliers, the final study population included 835 patients and 1,255 controls from Italy and 775 patients and 950 controls from Spain.
A total of 8,582,968 single-nucleotide polymorphisms (SNPs) were analyzed, and a meta-analysis of the Italian and Spanish cohorts was conducted.
What did the study find?
The team detected a cross-replicating association between SNPs on chromosome 3 and chromosome 9 that reached genome-wide significance.
A cluster of genes that could be relevant to the development of severe COVID-19 was identified on chromosome 3p21. One of these genes – SLC6A20 – encodes a transporter protein that interacts with angiotensin-converting enzyme 2 (ACE2), the host cell receptor that SARS-CoV-2 uses to gain viral entry.
In the lungs, this protein, which is called Sodium/Imino-acid Transporter 1 (SIT1), is mainly expressed in pneumocytes, and the authors think these cells should be investigated for any involvement that SIT1 may have in viral entry.
A lead SNP was also identified on chromosome 9 at the ABO blood group locus, and further analysis showed that A-positive participants were at a 45% increased for respiratory failure, while individuals with blood group O were at a 35% decreased risk for respiratory failure.
The authors say that early clinical reports have suggested the ABO blood group system is involved in determining susceptibility to COVID-19 and has also been implicated in susceptibility to SARS-CoV-1.
"Our data thus aligns with the suggestions that blood group O is associated with lower risk compared with non-O blood groups whereas blood group A is associated with higher risk of acquiring Covid-19 compared with non-A blood groups," the authors state.
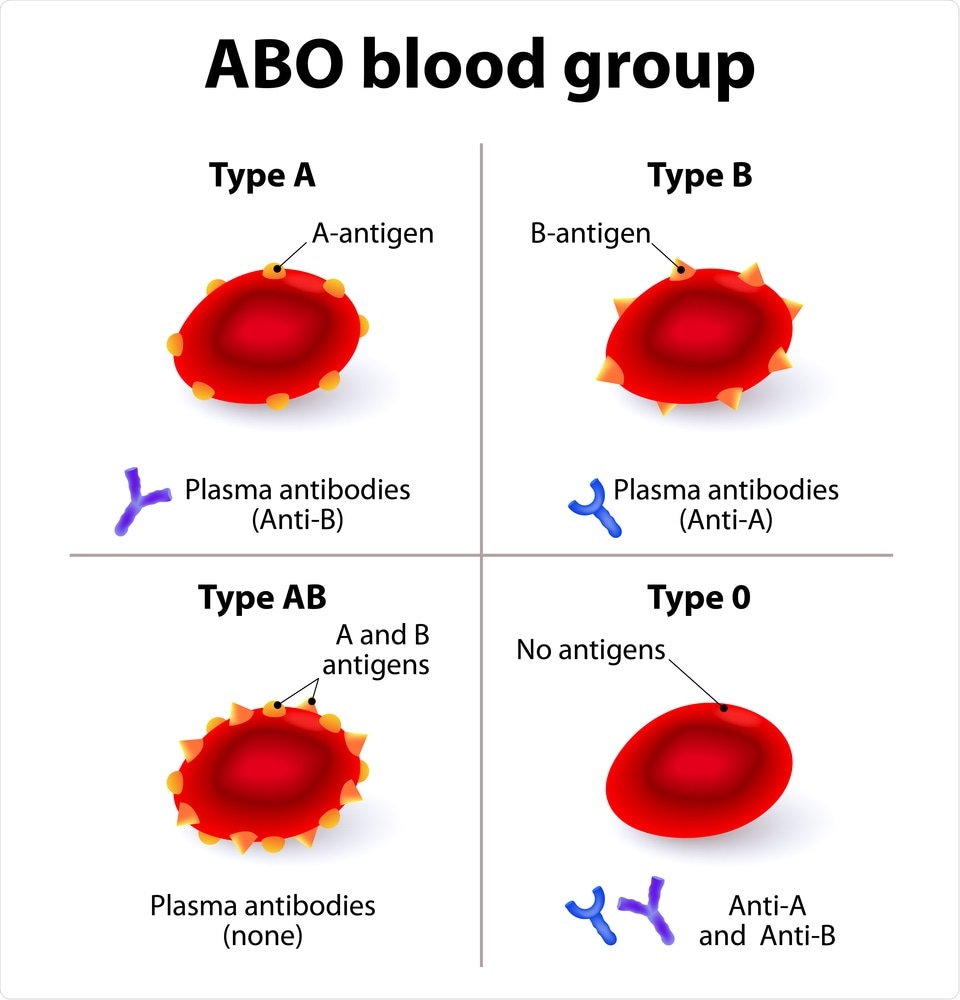
Study: The ABO blood group locus and a chromosome 3 gene cluster associate with SARS-CoV-2 respiratory failure in an Italian-Spanish genome-wide association analysis. Image Credit: Designua / Shutterstock
“Further exploration” of the findings is “now warranted”
“We herein report the first robust genetic susceptibility loci for the development of respiratory failure in Covid-19. Identified variants may help guide targeted exploration of severe Covid19 pathophysiology,” say Karlsen and team.
“Further exploration of current findings, both as to their utility in clinical risk profiling of Covid-19 patients and mechanistic understanding of the underlying pathophysiology, is now warranted,” they conclude.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Karlsen T, et al. The ABO blood group locus and a chromosome 3 gene cluster associate with SARS-CoV-2 respiratory failure in an Italian-Spanish genome-wide association analysis. medRxiv 2020. doi: https://doi.org/10.1101/2020.05.31.20114991
- Peer reviewed and published scientific report.
Ellinghaus, David, Frauke Degenhardt, Luis Bujanda, Maria Buti, Agustín Albillos, Pietro Invernizzi, Javier Fernández, et al. 2020. “Genomewide Association Study of Severe Covid-19 with Respiratory Failure.” New England Journal of Medicine, June. https://doi.org/10.1056/nejmoa2020283, https://www.nejm.org/doi/10.1056/NEJMoa2020283