A team of scientists from Sweden and India has recently conducted a study to evaluate type I interferon (IFN-I) responses in cells infected with severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). The findings reveal that SARS-CoV-2 infection is associated with a delayed and dysregulated IFN-I response and that pretreatment with IFN-I can reduce the risk of SARS-CoV-2 infection. The study is currently available on the bioRxiv* preprint server.
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative pathogen of coronavirus disease 2019 (COVID-19), is a beta coronavirus that shares many structural similarities with the other two lethal members of the human coronavirus family, including SARS-CoV and Middle East respiratory syndrome coronavirus (MERS-CoV). The upper or lower respiratory tract infection with any of these three beta coronaviruses can lead to severe pulmonary complications, especially in susceptible individuals. A substantial number of studies have shown that the pathogenesis of severe COVID-19 is often associated with aberrant IFN-I signaling, which is primarily caused by SARS-CoV-2-encoded immunomodulatory proteins (IFN antagonists).
IFN-1, including IFNα and IFNβ, is a part of the host's innate immune responses, which play a pivotal role in providing the first line of defense against any viral infection. Recognition of viral RNA by host pattern recognition receptors (TLRs and RLRs) triggers the production of IFN-I, which subsequently binds to specific cell surface receptors and activates a cascade of signaling events, ultimately leading to transcriptional activation of IFN stimulated genes with antiviral functions.
In the current study, the scientists used quantitative proteomics approaches to investigate the mechanism by which SARS-CoV-2 modulates cellular IFN-I signaling. To investigate the immunomodulatory effects, they infected human hepatocyte-derived cellular carcinoma cells (Huh7) with SARS-CoV-2. In a separate set of experiments, they pretreated the cells with IFN-I to explore its therapeutic potential in SARS-CoV-2 infection.
Important observations
The quantitative proteomics findings revealed that SARS-CoV-2 significantly altered IFN-I responses after 48 hours of infection. However, no alteration in IFN-I response was observed after 24 hours of infection. Cellular proteins associated with RIG-I and IFN-I signaling pathways were found to be significantly affected by SARS-CoV-2 infection. Specifically, delayed activation of RIG-I and dysregulation of IFN stimulated genes were observed in cells infected with SARS-CoV-2.
RIG-I is a cytosolic pattern recognition receptor that recognizes SARS-CoV-2 RNA. It triggers the production of IFN-β, leading to activation of IFN stimulated genes and initiation of virus eliminating processes. Given the RIG-I modulating effects of SARS-CoV-2, the scientists next characterized the IFN-β signaling in SARS-CoV-2-infected cells. The findings showed induction in IFN-β mRNA level after 48 hours of infection, which was associated with an induction in RIG-I expression. For further clarification, the scientists treated the virus-infected cells with RIG-I agonist and observed that SARS-CoV-2 could potently inhibit the production of IFN-β in RIG-I activated cells.
Regarding IFN stimulated genes, an increased protein expression of IFN stimulated gene 15 (ISG15) was observed after 48 hours of infection. Mechanistically, ISG15 can conjugate (ISGylation) to host proteins and viral proteins to regulate their functions. Interestingly, a significant reduction and induction in conjugated ISG15 levels were observed after 24 hours and 48 hours of infection, respectively. These observations reveal that SARS-CoV-2 can alter the conjugation status of ISG15 to modulate the cellular microenvironment.
Regarding the therapeutic potential of IFN-I, it was observed that pretreatment of cells with IFN significantly reduced the viral replication. In contrast, no such reduction was observed in cells that were first infected with the virus and then treated with IFN. These observations indicate that a cellular environment with a high IFN level is capable of suppressing the propagation of viral infection; however, once the infection is established, SARS-CoV-2 can modify the cellular environment in its favor by suppressing IFN signaling.
Because older adults are more susceptible to SARS-CoV-2 infection, the scientists investigated the IFN signatures in SARS-CoV-2-infected senescent cells (aging cells). The findings revealed that viral replication, as well as IFN response, increased significantly in these cells. Among various IFN stimulated genes studied, a significant reduction in the mRNA expression of IFIT1, an antiviral protein, was observed after infection. These findings reveal that despite having a heightened IFN response, senescent cells are highly susceptible to SARS-CoV-2 infection, probably because of virus-mediated reduction in IFIT1 level.
A comparison study conducted by the scientists revealed that SARS-CoV-2, SARS-CoV, and MERS-CoV possess distinct IFN signatures and that each of these viruses differentially regulates the transcription of IFN stimulated genes.
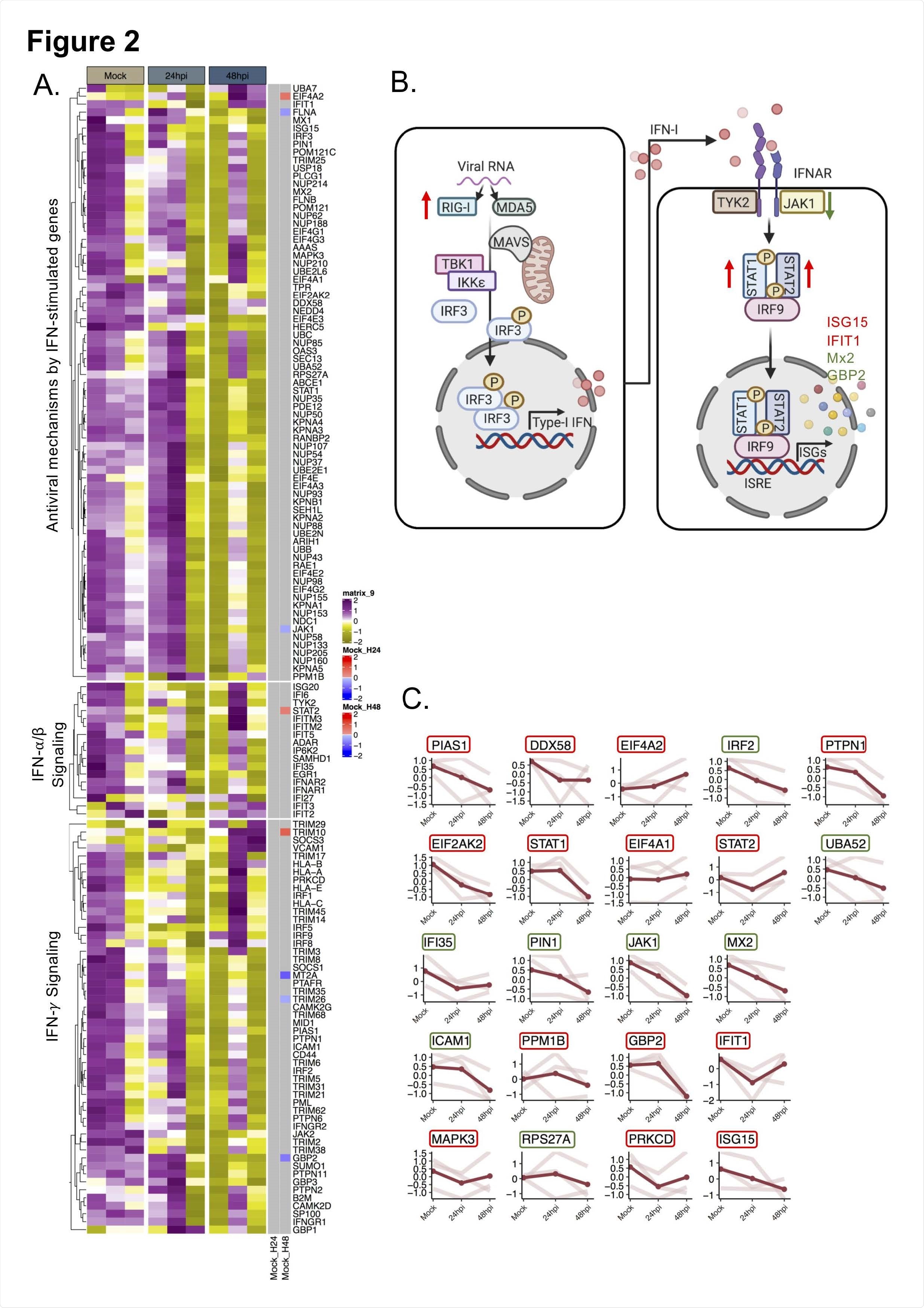
SARS-CoV-2 induced transcriptional changes in the IFN-signaling genes in transcriptomics data. A) Heatmap of IFN-stimulated transcripts before infection and at 24 and 48hpi. Data were log2 normalized and Z-score transformed. Lower values are represented in yellow and higher values in purple. Significant differential expressed genes between time points are indicated in blue if downregulated and in red if up regulated. B) The scheme graph of the type I interferon signaling pathways, in which the regulated genes expression level trend is noted. The significantly changed proteins observed in the proteomics data are denoted by green arrows or letters (downregulated) or red arrows or letters (upregulated) C) Dot plot for each transcript that were detected as significantly altered in proteomics. For each gene, the scaled values in triplicates are represented in mock, 24hpi and 48hpi and linked by light red line, average value is displayed in red. The name of the genes is indicated in colored box based on the proteomics data. The genes corresponding to increased protein levels are in red boxes and to decreased protein levels in green boxes.
Study significance
The study provides an in-depth overview of IFN response during SARS-CoV-2 infection and reveals the therapeutic importance of interferons in COVID-19 patients.
Journal reference:
- Type-I interferon signatures in SARS-CoV-2 infected Huh7 cells Xi Chen, Elisa Saccon, K. Sofia Appelberg, Flora Mikaeloff, Jimmy Esneider Rodriguez, Beatriz Sá Vinhas, Teresa Frisan, Ákos Végvári, Ali Mirazimi, Ujjwal Neogi, Soham Gupta, bioRxiv, 2021.02.04.429738; doi: https://doi.org/10.1101/2021.02.04.429738, https://www.biorxiv.org/content/10.1101/2021.02.04.429738v1