The importance of computational modeling in the prediction of the course of the ongoing coronavirus disease 2019 (COVID-19) pandemic, as well as evaluating multiple potential drug leads, has never been more clear. Now, a new study describes a set of designer vaccine antigens that may improve vaccine stability as well as enhance the neutralizing antibody response.
The virus causing this pandemic is the novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which causes an asymptomatic or mild infection in the vast majority, but spreads rapidly and extensively to produce a huge caseload in a relatively short period of time. This has led to overwhelming demands on healthcare services in many severely affected countries.
The only effective way out has been thought to be attempting to achieve population immunity by potent COVID-19 vaccines that can prevent symptomatic disease, at least, and thus reduce the number of hospitalizations and deaths. Over a dozen are currently being used after receiving emergency use authorization (EUA) by the concerned regulatory authorities on different continents.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Most of these vaccines are based on modified SARS-CoV-2 spike protein, stabilized in the prefusion conformation by a couple of amino acid changes in the S2 spike domain. This is supposed to enhance vaccine efficacy.
As new strains emerge, including the variants of concern (VOCs) that seem to reduce the neutralization capacity of some of these vaccines generated against earlier variants, there is a need for next generation vaccines. Not only will this ensure effective protection, but they may solve the logistical problems with the current messenger ribonucleic acid (mRNA) vaccines that are among the most effective vaccines available at present.
The current study focused on engineering the spike by introducing stabilizing mutations outside the S2 domain to solve these issues.
The viral spike engages with the host cell receptor, the angiotensin-converting enzyme 2 (ACE2), via its receptor-binding domain (RBD), to accomplish viral entry. Thus, the most potent neutralizing antibodies bind to the RBD, and this domain, in isolation, can elicit powerful neutralizing antibodies.
Convalescent serum has higher levels of non-neutralizing antibodies, binding to epitopes outside the RBD, compared to neutralizing RBD-targeting antibodies. The reason is speculated to be the unstable structure of RBD neutralizing epitopes, as well as the ‘down’ conformation of the RBD much of the time that hides these epitopes from immune cells.
This is the rationale for stabilizing the RBD in the ‘up’ conformation where its neutralizing epitopes are accessible to immune cells, thus ensuring a more targeted immune antibody response. This would involve either allowing the RBD to be used as the vaccine antigen or using the spike antigen with the RBD stabilized in the ‘up’ conformation.
Novel design and screening approach
The current study, which has been released as a preprint on the bioRxiv* server, reports the use of a novel pipeline for computational design and in vitro screening, called Stabilizer for Protein Expression and Epitope Design (SPEEDesign), for this purpose.
The goal of the designers was fourfold: the antigen should ensure a targeted and powerful neutralizing antibody response; non-neutralizing or immunodominant epitopes should be blocked; the antigen should be heat-stable so that it survives longer in the human body; and the antigen should be exposed in the optimal state, such as the ‘up’ conformation of the RBD, for instance.
The process led to identifying three enhanced antigens, with about nine residue changes compared to the wildtype RBD sequence. All had four residue changes that were common.
These were produced as properly folded proteins with four-fold higher yields compared to the wildtype RBD, which is important for commercial production. Compared to the ”hexapro” spike ectodomain that is considered optimized, or the widely-used double-proline stabilized spike ectodomain, the yields are six-fold and 200-fold higher, respectively.
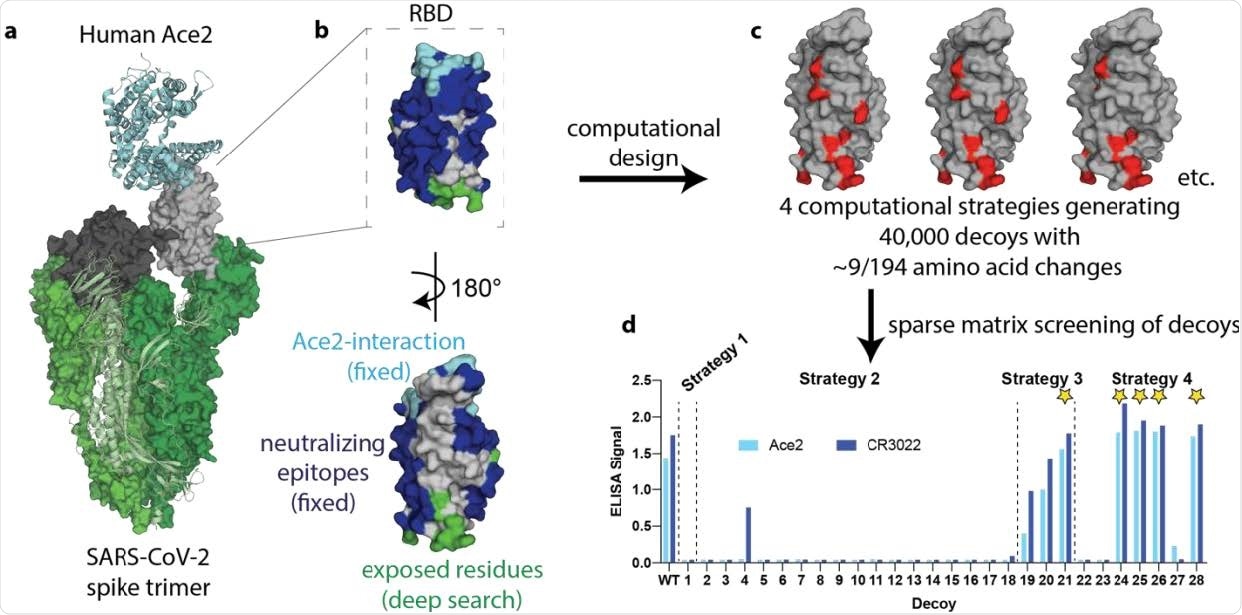
Overview of SPEEDesign pipeline used to create RBD immunogens. a, The SARS-CoV-2 spike trimer (green) binds human Ace2 (cyan) to mediate viral entry. This interaction is mediated by the “up” conformation of the RBD (grey), which can also exist in a down conformation (black). b, The RBD design process retained the Ace2-interaction surface (cyan) and all known SARS neutralizing epitopes (blue). Residues exposed upon isolation of the RBD (green) were heavily designed while all other residues (grey) were designed more conservatively. c, Four computational strategies were used to create 40,000 decoys, each of which has an average of nine amino acid changes (red). d, 28 sequences sampling the top scoring decoys were screened in vitro, identifying 5 lead immunogen candidates (stars).
These were also heat-stable, with a melting temperature 5-10 °C higher than the wildtype RBD. Not only will this result in higher yields, but the half-life will be longer following administration, as well as obviating the necessity for cold-chain temperatures during storage and transport, as well as at the vaccination centers themselves.
Structurally, the changes seem to improve the solubility of the RBD and may promote the ‘up’ conformation. All engineered residues occur distal to the mutations found in the current VOCs, and thus are suitable for the development of next-generation vaccines based on these variants.
In fact, they are expressed even better with the South African variant than with the reference strain, indicating that the engineered immunogens are more stable as a result of the structural and energetic changes resulting from the modified residues. This has occurred without adversely affecting the structure of the antigen.
As a result, neutralizing epitopes are retained intact, with ACE2-binding ability comparable to that of the wildtype RBD in almost all cases. This is important as it ensures that the resulting antibodies will recognize the actual wildtype RBD and hence neutralize the virus.
In vivo mouse studies demonstrated high neutralizing antibody titers with geometric mean titers (GMT) greater than the wildtype RBD, confirming that the modified residues produced favorable biophysical characteristics.
In fact, one of them showed a 30-fold higher GMT than that elicited by the wildtype RBD, compared to the ten-fold increase seen with the earlier 2-P prefusion stabilized spike antigen. This is equal to a five-fold increase in the lifetime of the antibody, and thus of the duration of protection. Not only so, it more than overcomes the 1.5-12 fold reduction in neutralizing capacity observed when earlier vaccine-elicited antibodies are tested against the South African variant, indicating a broad range of protection.
Functional antibody responses involving ACE2 blocking were also up to ten-fold higher.
What are the implications?
The study reports a new approach that may be very helpful in vaccine development. Secondly, the use of a few amino acid changes in the RBD was associated with markedly improved neutralizing responses. This suggests that these changes are suitable for use in next-generation vaccines to ensure a broader and longer protective response against multiple variants of SARS-CoV-2.
The designed immunogens can also be easily adapted to all other vaccine platforms by making amino acid changes to the RBD sequence within the FL-spike or within RBD-only vaccines. The changes identified here do not exclude the existing improvements in spike, they are fully compatible with changes in the S2 domain, and they are compatible with all known changes in emerging SARS-CoV-2 variants of concern.”
The stability-enhanced yield, and amenability to methods already in use to improve immunogenicity make this a very attractive option for the sufficiently large production of vaccines against the pandemic.
The reasons for this dramatically improved performance remain to be known, but it is clear that the non-neutralizing antibody response was not raised anywhere as much as the neutralizing antibody titer. Thus, this platform will be useful in the development of customized vaccine antigens against many pathogens which have epitopes, the structure of which is known.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources