Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the causative agent of the ongoing coronavirus disease 2019 (COVID-19) pandemic, is a virus with a ribonucleic acid (RNA) genome.
Now a new preprint research paper posted to the bioRxiv* server dismisses the possibility, raised by that study, that the virus inserts its genetic material into the host cell’s genome.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Background
This virus lacks a reverse transcriptase enzyme, which is required to convert the RNA into its complementary deoxyribonucleic acid (DNA) strand. Therefore, SARS-CoV-2 does not incorporate its genetic material into human genomic DNA during its infection lifecycle.
This is a crucial assumption that underlies the current diagnostic techniques used for this infection and for its potential long-term consequences. There are other viruses, such as the human immunodeficiency virus 1 (HIV-1) and hepatitis B virus (HBV), which do integrate with DNA and cause cancerous transformation or chronic infection.
All mammalian cells have LINE-1 (L1) retrotransposons in their genomes. This gives rise to messenger RNA (mRNA) that encodes two proteins required for the movement of L1. These are represented as ORF1p and ORF2p.
The ORF2p acts as an endonuclease as well as a reverse transcriptase (RT), with a strong tendency to catalyze reverse transcription of the L1 mRNA. It can also mobilize cellular RNA, which carries a polyadenylated tail, including RNA from other retrotransposons and from genes encoding proteins.
The latter activity is infrequent, however, probably because it is suppressed by the cell. In fact, less than a single cellular RNA insertion should occur per 2,000 L1 insertions.
Both types of insertions are marked by target site duplications (TSDs) and a 3′ polyA region, occurring at the L1 sequence 5′-TTTT/AA. These signs help to identify true insertions.
Earlier evidence
Earlier, SARS-CoV-2-infected cells with overexpressed L1 were shown to contain DNA identical to viral RNA. This was coupled with other weaker evidence to prove the integration of SARS-CoV-2 RNA into the host cell genome.
This was followed by the identification of 63 supposed sequences integrated from SARS-CoV-2 by the same researcher, using a sequencing technique called Oxford Nanopore Technologies (ONT) long-read sequencing. Only one of these was spanned by an ONT read, on the X chromosome, which showed potential TSDs on either side, and this one lacked the expected 3′ polyA tract.
Moreover, a 28 kb deletion seemed to have occurred from the viral sequence. Exons were enriched 26-fold in these integrants, a finding that cannot be explained by the preference of the L1 enzyme.
The second study was followed by still another, using Illumina short-read sequencing. This was claimed to show supposed junctions where SARS-CoV-2 sequences were joined to the host genome in cells lacking high levels of L1.
Other interpretations
However, the reads did not span the integrated sequence. Again, the preparation of a library for Illumina sequencing is known to tend towards artifact generation. These facts leave the conclusion in doubt.
Earlier, however, the scientists responsible for the current preprint used ONT reads to detect L1-dependent reverse transcription of viral genomes. One example is the incorporation of two hepatitis C virus (HCV) sequences in a liver tumor sample.
Moreover, the cell platform used for ONT sequencing, namely, HEK293T cells, has many features that may make it easy to assess the occurrence of such reverse-transcribed viral sequences. For instance, these cells express the L1 ORF1p, have been used to accept inserted L1 retrotransposons, and serve to support productive SARS-CoV-2 infection.
Study aim
Using ONT sequencing, the current study attempted to analyze genomic DNA from this cell line after infecting them with SARS-CoV-2 at multiple doses. Control cells were the tumor and non-tumor cell lines from a liver tumor positive for the hepatitis B virus.
The ONT sequencing data from the earlier studies were added. As negative controls, the previously identified HCV-positive liver carcinoma cells were used, along with normal liver cells.
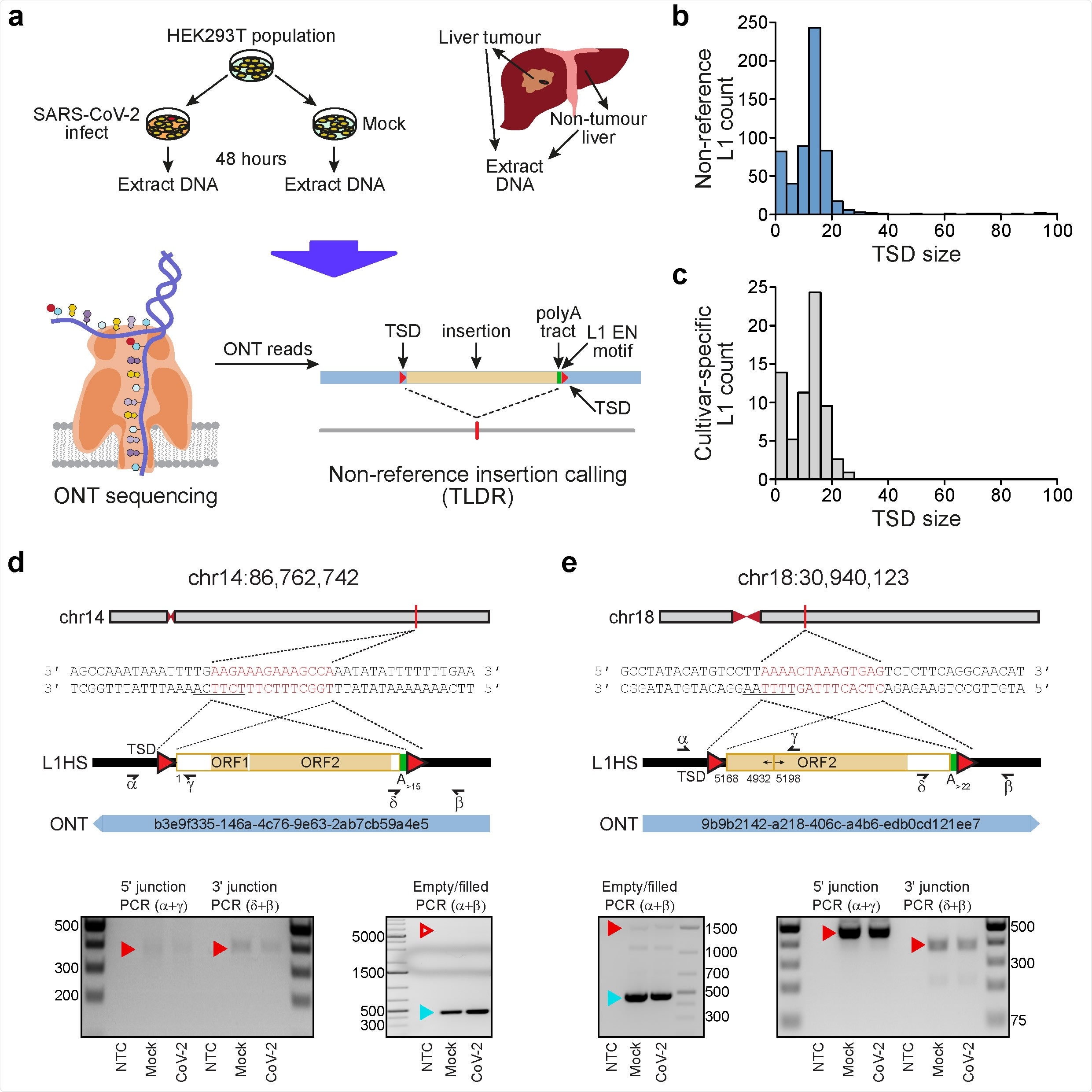
Detection of endogenous L1-mediated retrotransposition in human cells. a, Experimental design. HEK- 293T cells were divided into two populations (cultivars), which were then either SARS-CoV-2 infected or mock infected. DNA was extracted from each cultivar, as well as from hepatocellular carcinoma patient samples, and subjected to ONT sequencing. ONT reads were used to call non-reference L1 and virus insertions with TLDR, which also resolves TSDs and other retrotransposition hallmarks. TSDs: red triangles; polyA tract: green rectangle; ONT read: blue rectangle. Note: some illustrations are adapted from Ewing et al.23. b, TSD size distribution for non-reference L1 insertions, as annotated by TLDR. c, As for b, except showing data for L1 insertions found only in either our HEK293T cells infected with SARS-CoV-2 or our mock infected cells. d, Detailed characterisation of an L1 insertion detected in SARS-CoV-2 infected HEK293T cells by a single spanning ONT read aligned to chromosome 14. Nucleotides highlighted in red correspond to the integration site TSD. Underlined nucleotides correspond to the L1 EN motif. The cartoon indicates a full-length L1HS insertion flanked by TSDs (red triangles), and a 3ʹ polyA tract (green). Numerals represent positions relative to the L1HS sequence L1.346. The relevant spanning ONT read, with identifier, is positioned underneath the cartoon. Symbols (α, β, δ, γ) represent the approximate position of primers used for empty/filled site and L1-genome junction PCR validation reactions. Gel images display the results of these PCRs. Ladder band sizes are as indicated, NTC; non-template control. Red triangles indicate the expected size of L1 amplicons (empty triangle: no product observed; filled triangle: product observed). Blue triangles indicate expected empty site sizes. e, As for d, except for a 5ʹ inverted/deleted L1HS located on chromosome 18.
No insertions found
All of these supposedly integrated sequences were analyzed to identify all L1 insertions specific for human cells, spanned by at least one ONT lead aligned to a unique locus. Not a single insertion was detected from the SARS-CoV-2, HCV or HBV genomes.
However, the program did identify 575 L1 insertions with the flanking TSDs. Seventy-eight were unique to the SARS-CoV-2-infected cells or mock-infected controls, the majority spanned by a single read.
Among these single-spanning insertions, six, which were chosen at random for manual checking and confirmation by a polymerase chain reaction (PCR) test, were confirmed to show all three features of a true L1 insertion.
Secondly, the researchers found that the program could recover a single HBV integrated sequence.
However, it did not show the presence of any ONT reads that could align to the SARS-CoV-2 or HCV genomes, showing that retrotransposons and insertions from exogenous viruses are reliably distinguished.
This was followed by the discovery that the ONT data from the first study contained 555 reads that could be aligned to the SARS-CoV-2 genome. One of these reads was the putative integrated sequence on the X chromosome that did not have a 3’ polyA tract.
The mean length of these reads was too short, however, compared to the overall dataset. Over half of these reads were composed of SARS-CoV-2 sequence, a high proportion relative to L1 sequences in the reads that were aligned to L1.
The program failed to call supposed integrants from the SARS-CoV-2 genome. Thus, despite the presence of reads that could be aligned to the viral genome in the dataset of the earlier study, these reads were too short compared to the typical read. Thus, the researchers postulate that these could be molecular artifacts wrongly interpreted as SARS-CoV-2 integrated sequences.
These results point to the ability of single spanning ONT reads to detect retrotransposons and to show endogenous L1 insertions in this cell line that lacked overexpressed L1.
What is the conclusion?
“In sum, we do not observe L1-mediated SARS-CoV-2 genomic integration in HEK293T cells, despite the availability of the L1 machinery.”
The reason is that the L1 clearly prefers to reverse transcribe its own mRNA rather than SARS-CoV-2. Thus, although possible, this is probably a rare phenomenon, as with other poly-A-bearing cellular RNAs.
“That we found no evidence of SARS-CoV-2 integration suggests such events in vivo are highly unlikely to drive later oncogenesis or explain post-recovery detection of the virus.”

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources