The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which is the causal agent of the coronavirus disease 2019 (COVID-19), has a polybasic cleavage motif in the spike (S) protein. This is recognized by the hosts’ furin protease. The S protein is activated by proteolytic cleavage, which, in turn, affects the cellular entry pathway and cell tropism of SARS-CoV-2.
Background
Proinflammatory cytokines like interleukin 6 (IL-6) play a vital role in the hyperinflammatory response during the acute phase of infection, which is often associated with the severity of the disease. SARS-CoV-2 variants with adaptive mutations in their S gene have been detected in many countries, which has raised concerns over the transmissibility and immune escape against ancestral SARS-CoV-2.
The S protein, which is located on the virion surface, is a homotrimeric glycoprotein that binds to entry receptors and plays an important role in the entry of SARS-CoV-2 into host cells. The S1/S2 boundary has a discriminative polybasic cleavage motif that is recognized by the host furin protease.
Although previous research has characterized the SARS-CoV-2 mutants well, the role of the furin cleavage site in pathogenesis and cell tropism has yet to be established. Scientists have noted that the loss of the cleavage site leads to attenuation in the pathogenicity of SARS-CoV-2 in animal models.
About the study
The current study uses a hamster model to investigate the in vivo growth and pathogenicity of S gene mutants with deletions or substitutions at the furin cleavage sites of the S proteins. Scientists used histopathological and cytokine expression analyses to study the muted and mild inflammatory response post-infection with S gene mutants.
The hamsters that were infected with the S gene mutants developed neutralizing antibodies that cross-reacted with different lineages of SARS-CoV-2. This led scientists to study whether primary infection with S gene mutants protected the animals from reinfection with both the parental SARS-CoV-2 strain, as well as the emerging variants belonging to the lineages B.1.1.7 and P.1.
Study findings
Hamsters develop pneumonia and experience body weight loss following infection with SARS-CoV-2. In the present study, the histopathological findings and cytokine gene expression levels revealed that hamsters infected with S gene mutants showed no bodyweight loss and only slight lung damage as compared to clinical strains. This shows the attenuated virulence of S gene mutants in hamsters.
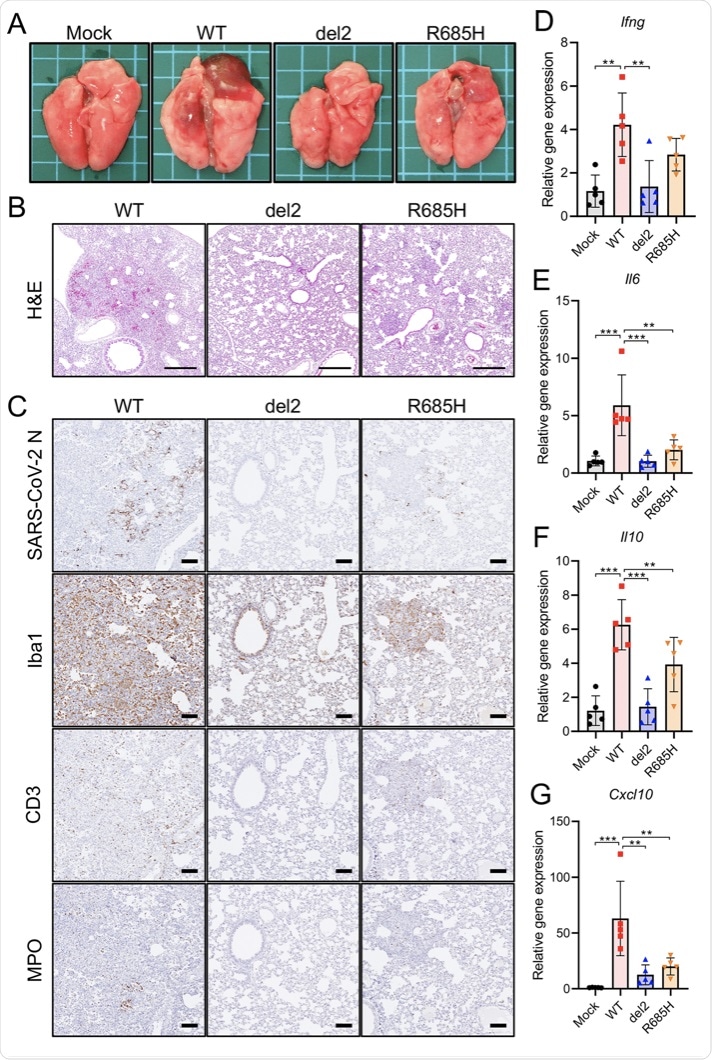 Pathological changes and immune response in the lung tissues of hamsters infected with SARS-CoV-2 S gene mutants. (A) Gross pathological images of the lungs of hamsters infected with WT or S gene mutants at 4 days postinfection (dpi). (B) Histopathological images of the lungs of hamsters infected with WT or S gene mutants at 4 dpi with H&E staining. Scale bars = 500 μm. (C) Immunohistochemistry for SARS-CoV-2 N protein, macrophage (Iba1), T cell (CD3), and neutrophil (MPO) markers. Cell nuclei were counterstained with hematoxylin. Scale bars = 100 μm. (D to G) Cytokine gene expression profile in lung tissues from hamsters at 4 dpi. Relative gene expression levels of the indicated cytokines in the lungs compared with those of lungs from mock-infected hamsters were examined using qRT-PCR. Data were normalized to β-actin. One-way analysis of variance with Tukey’s test was used to determine the statistical significance of the differences. **, P < 0.01; ***, P < 0.001.
Pathological changes and immune response in the lung tissues of hamsters infected with SARS-CoV-2 S gene mutants. (A) Gross pathological images of the lungs of hamsters infected with WT or S gene mutants at 4 days postinfection (dpi). (B) Histopathological images of the lungs of hamsters infected with WT or S gene mutants at 4 dpi with H&E staining. Scale bars = 500 μm. (C) Immunohistochemistry for SARS-CoV-2 N protein, macrophage (Iba1), T cell (CD3), and neutrophil (MPO) markers. Cell nuclei were counterstained with hematoxylin. Scale bars = 100 μm. (D to G) Cytokine gene expression profile in lung tissues from hamsters at 4 dpi. Relative gene expression levels of the indicated cytokines in the lungs compared with those of lungs from mock-infected hamsters were examined using qRT-PCR. Data were normalized to β-actin. One-way analysis of variance with Tukey’s test was used to determine the statistical significance of the differences. **, P < 0.01; ***, P < 0.001.
The cellular entry mode of S gene mutants accounts for the low growth capacity in respiratory airways in hamsters. During viral entry, the S protein is primed by the host’s TMPRSS2 or cathepsin, which facilitates membrane fusion.
The entry of S gene mutants is different in that it is triggered by the cathepsin-dependent endosome pathway. The direct fusion pathway enables rapid entry and escapes from innate immune restrictions by interferon (IFN)-induced transmembrane proteins.
These observations led scientists to conclude that S gene mutants exhibit low infectivity in some cell lines, including the human lung-derived Calu-3 cells. The present study also documented that the mild infection is sufficient to induce protective immunity against SARS-CoV-2 in hamsters.
The usage of attenuated virus strains in vaccines is not new. Previous examples where this has been done are yellow fever virus 17D strain, measles, poliovirus, and Sabin strain, to name a few.
Study takeaways
The current study demonstrates that the SARS-CoV-2 S gene mutants can also induce protective immunity in hamsters. The infection with S gene mutants inhibited viral growth in both the nasal and lung tissues of hamsters that were reinfected with a pathogenic clinical strain of SARS-CoV-2. This finding is quite important and highlights the benefit of vaccination.
Scientists also noted that the S gene mutant-induced protective immunity cross-reacted with emerging SARS-CoV-2 variants that belong to the B.1.1.7 and P.1 lineages. These variants have the capacity to evade neutralization with some monoclonal antibodies, as they possess certain mutations (K417T, E484K, and/or N501Y) at the receptor-binding domain in the spike protein. The results in hamster models are encouraging and the next step will be to establish the pathogenicity of S gene mutants in humans.
Conclusion
The current study has some limitations. The hamsters were challenged with SARS-CoV-2 relatively quickly; therefore, the long-term immunity needs to be tested. The pathogenicity of S gene mutants in humans has also yet to be studied.
However, the broad neutralizing activity of S gene mutants across different lineages hints at their potential use as immunogens in live-attenuated vaccines.