Throughout the ongoing coronavirus disease 2019 (COVID-19) pandemic, antibody detection has been crucial for epidemiological surveillance, examining the natural history of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), and evaluating potential vaccine candidates.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Background
Antibodies specific to the SARS-CoV-2 spike protein and its receptor-binding domain (RBD) have been shown to neutralize viral binding and entrance in serological investigations/ Thus, antibodies are often considered correlates of protection against infection and reinfection. Current COVID-19 vaccines elicit immune responses to the spike protein; thus, vaccine assessment has relied heavily on serological research.
Saliva can be utilized for the detection of anti-SARS-CoV-2 antibodies. However, there remains a lack of research on novel salivary immunoassays for the detection of recent infection.
Saliva primarily contains two antibody types including secretory immunoglobulin A (SIgA) and IgG. Since SARS-CoV-2 has an affinity towards cells in the respiratory tract, localized production of mucosal IgA antibodies could be vital for protection against infection and reducing the rate of transmission, whereas salivary IgG may serve as a surrogate for systemic immunity.
Novel salivary immunoassays to detect SARS-CoV-2 antibodies
In a recent study published on the preprint server medRxiv*, six salivary enzyme-linked immunosorbent assays (ELISA) were developed to evaluate the immune response to SARS-CoV-2 and estimate rates of past infection. The ELISA methodology utilized in this study is based on previous studies.
For the current study, whole saliva samples acquired from the Bristol BioBank were obtained from individuals who had recovered from polymerase chain reaction (PCR) confirmed COVID-19, suspected SARS-CoV-2 infections, and healthy individuals.
In a blinded test accuracy trial, the researchers were interested in determining whether an assay might identify recently confirmed SARS-CoV-2 infection. Herein, they linked antibody levels detected by the assays in paired serum and saliva samples to assess salivary antibody responses and allow for its implementation in cohorts with uncertain infection status.
Finally, the authors evaluated mucosal antibody responses in recently exposed adults and children by field-testing top-performing assays in a home transmission environment. This research establishes a solid foundation for evaluating saliva as a sample for SARS-CoV-2 antibody detection and provides insight on how to characterize mucosal immune responses after infection.
The operating settings of the assay were adjusted to minimize background, maximize selectivity among positive and negative samples, and maintain a wide dynamic range. The greatest background was observed on a high-binding hydrophilic plate, whereas background was reduced and equivalent on the remaining medium binding hydrophobic plates/ Ultimately, the Griener plate provided the best selectivity.
For each antigen, an antigen coating concentration of 10 µg/mL was considered optimal. The optimal sample and secondary antibody concentrations were identified through the utilization of checkerboards. The ELISA signal was unaffected by heat inactivation or freeze-thawing of samples, thus demonstrating that this assay allows for safe and practical sample handling.
How well did the assay perform?
The spike IgA and IgG assays showed the greatest selectivity among known negative and positive samples determined by the combined sample set. Both nucleocapsid (N) protein and RBD IgA had poor selectivity and few known positives displayed reactivity to N-protein, whereas several known negatives displayed reactivity to RBD.
Spike IgG was the most effective assay, followed by N-protein IgG; however, N-protein IgA produced the worst results. The threshold that offered optimal detecting rates of PCR-confirmed cases while sustaining at least 98% specificity invalidation was chosen after examining each assay's overall effectiveness in the combined threshold/validation set. Importantly, thresholds set for 98% specificity sustained this effectiveness in the validation set.
Does combining the assays allow for a better prediction of COVID-19 status?
Response heterogeneity was detected among antigens, as well as a disconnect in the isotype reactivity for the same antigen. Individuals appeared to have mostly SARS-CoV-2 specific IgA or IgG, with only a few having significant amounts of both. The spike IgG and, to a lesser extent, the spike IgA assays ensure the highest selectivity.
The authors hypothesized that combining findings from different assays could enhance sensitivity for recent infection because of the heterogeneity in response. AdaBoost classifiers were used to predict positive and negative individuals through the use of one or a combination of the six assays to test this hypothesis.
Data from the N-protein, RBD, and spike IgG assays were used to train the best model. This model outperformed the models trained with the individual assays by a significant margin.
The model trained with the spike IgA and IgG assays outperformed those trained with the individual spike IgA and spike IgG assays by a small margin. When N-protein IgG and spike IgG assays were combined, the results were comparable to when both spike assays were combined.
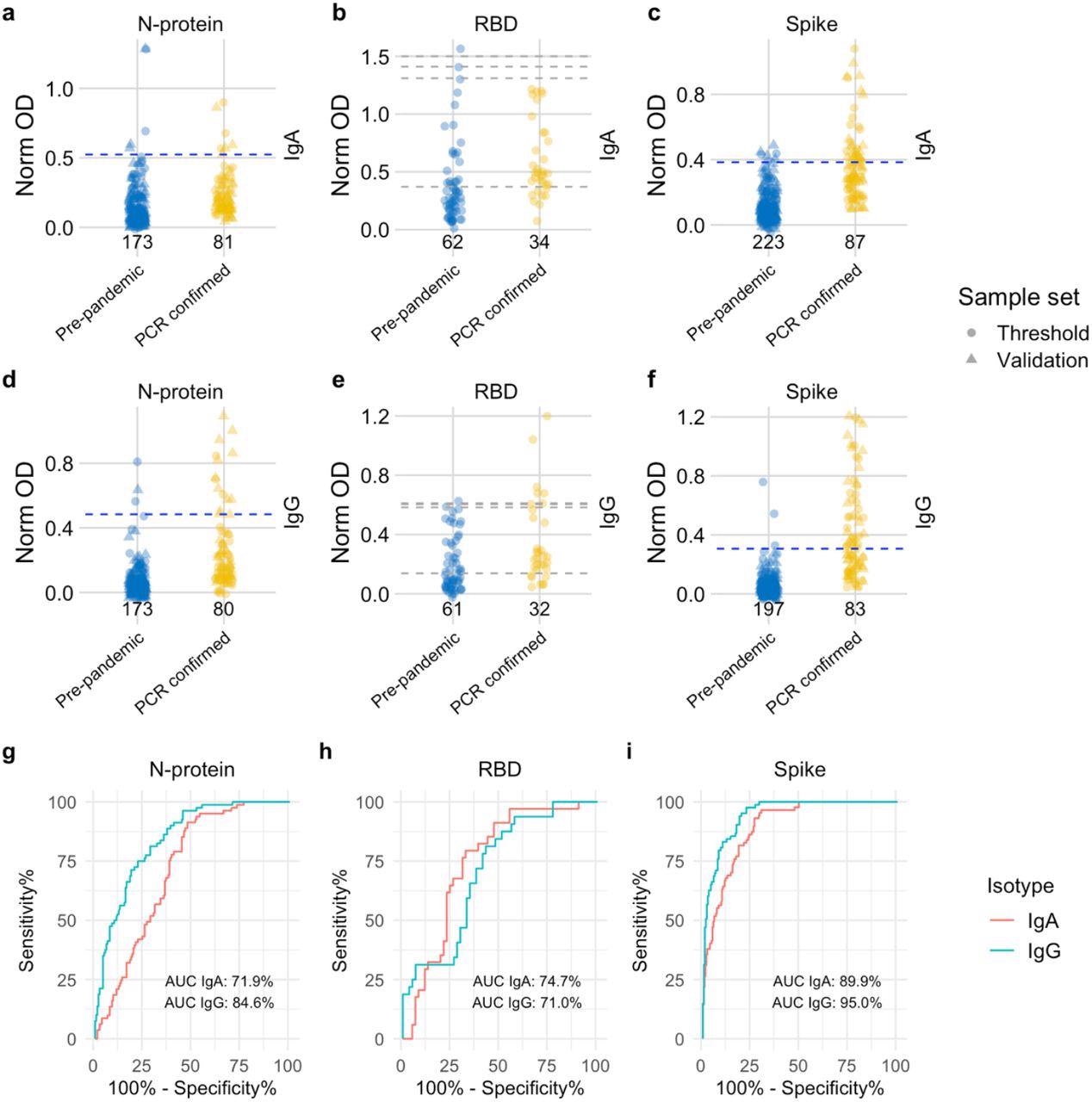 Distribution of semi-quantitative titres for each assay and corresponding ROC curves. a-c, Dotplots show the scatter of values for SARS-CoV-2 specific IgA responses and specific IgG responses in d-f. Data are presented for known negative and known positive samples, the number of samples tested in each group is shown under the corresponding data points. For N-protein and Spike, proposed thresholds (97th – 99th percentile and Youden’s index) are shown as dashed lines, with the final selected threshold in blue; thresholds were derived in the threshold setting phase (threshold samples shown as circles, validation samples shown as triangles). RBD was not taken forward to full evaluation, so performance in the threshold set and corresponding proposed thresholds are shown as dashed grey lines. g, i, ROC curves for N-protein and Spike represent assay performance determined on all threshold and validation set samples combined. h, The ROC curve for RBD represents assay performance in threshold sample set. ROC = receiver operating characteristic. AUC = area under the curve. Norm OD = Normalised OD (a relative ratio to the serum standard). N-protein = Nucleocapsid protein. RBD = Receptor binding domain.
Distribution of semi-quantitative titres for each assay and corresponding ROC curves. a-c, Dotplots show the scatter of values for SARS-CoV-2 specific IgA responses and specific IgG responses in d-f. Data are presented for known negative and known positive samples, the number of samples tested in each group is shown under the corresponding data points. For N-protein and Spike, proposed thresholds (97th – 99th percentile and Youden’s index) are shown as dashed lines, with the final selected threshold in blue; thresholds were derived in the threshold setting phase (threshold samples shown as circles, validation samples shown as triangles). RBD was not taken forward to full evaluation, so performance in the threshold set and corresponding proposed thresholds are shown as dashed grey lines. g, i, ROC curves for N-protein and Spike represent assay performance determined on all threshold and validation set samples combined. h, The ROC curve for RBD represents assay performance in threshold sample set. ROC = receiver operating characteristic. AUC = area under the curve. Norm OD = Normalised OD (a relative ratio to the serum standard). N-protein = Nucleocapsid protein. RBD = Receptor binding domain.
Implications
Because of the fast-rising complexity of COVID-19 epidemiology throughout the world, new tools are needed to guide challenging policy decisions, particularly on vaccination and in countries with insufficient data on population immunity. Antibody assays should be tested in the populations to which they are deployed, which is more important in regions with high rates of infection and where pre-existing immunity varies widely.
The best diagnostic selectivity could be gained through the implementation of salivary immunoassays. Moreover, this method could be a vital tool in secluded and rural parts of the world where access to diagnostic equipment is limited. Furthermore, this method would provide significant benefits in the remote regions if it was manufactured to be low cost and high throughput.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Thomas, A. C., Oliver, E., Baum, H. E., et al. (2022). Evaluation of isotype specific salivary antibody assays for detecting previous SARS-CoV-2 infection in children and adults. medRxiv. doi:10.1101/2022.04.11.22273690. https://www.medrxiv.org/content/10.1101/2022.04.11.22273690v1.full-text.
- Peer reviewed and published scientific report.
Thomas, Amy C., Elizabeth Oliver, Holly E. Baum, Kapil Gupta, Kathryn L. Shelley, Anna E. Long, Hayley E. Jones, et al. 2023. “Evaluation and Deployment of Isotype-Specific Salivary Antibody Assays for Detecting Previous SARS-CoV-2 Infection in Children and Adults.” Communications Medicine 3 (1): 1–14. https://doi.org/10.1038/s43856-023-00264-2. https://www.nature.com/articles/s43856-023-00264-2.