Feb 7 2019
Chemical and topographical single-cell imaging at nanoscale resolution by near-field desorption mass spectrometry
How are chemicals distributed in a cell? Chinese scientists have developed a combined mass spectrometry and biological imaging device that enables direct, label-free detection, and high-resolution mapping of chemicals inside a biological cell. As demonstrated in their publication in the journal Angewandte Chemie, the distribution and accumulation of the disinfectant proflavine around the cell organelles could be visualized directly, based on the mass signal from the molecule.
Ultrafine optical methods, such as STED and PALM microscopy, are well-established techniques to identify gene expression and localize molecules in cell compartments at molecular resolutions. But these are indirect methods, which usually monitor the fluorescence generated when a dye binds to target molecules.
A direct method to identify molecules is mass spectrometry, which senses the chemical mass of a molecule that has been desorbed from a surface and ionized by a laser beam. However, mass spectrometry poses inherent diffraction problems when combined with high-resolution imaging processes. In addition, biological cells usually have rough surfaces, which give rise to signal artifacts. Considering all these challenges, Wei Hang and colleagues at Xiamen University, Xiamen, China, have now constructed a time-of-flight mass spectrometer with a desorption–ionization-imaging method that accounts for both the special surface conditions of biological cells and the high resolution demanded in such a system.
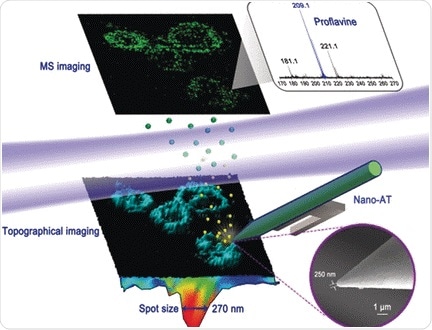
They developed an elaborate setup called “near-field desorption postionization time-of-flight mass spectrometer” (NDPI–TOFMS) and used it to detect and map chemical molecules in HELA cells—a human cell line and workhorse in cell biology. The dried cells were put on a stage and an ultraprecise laser scanned the surface by etching craters a few tenths of a micrometer in size. The desorbed molecules were ionized by another laser beam and then identified in the mass spectrometer.
As the authors pointed out, the advantage of this method is that the cells can be imaged at the same time as sample acquisition, thus enabling “co-registered chemical and topographical imaging within an individual cell.” Indeed, their 3D reconstructed images revealed the signals for proflavine, a drug that was added to the cells, exactly where they were expected: in the cytoplasm and around the organelles. The three-dimensional information was gathered to account for the uneven surface.
In contrast to the available mass spectrometry imaging techniques, this “hybrid technique,” which combined scanning probe microscopy and mass spectrometry “provides undistorted high-resolution chemical mapping of irregular surfaces,” says Hang. Given the compact nature of the device, the authors recommend its implementation in diverse mass spectrometry imaging setups, but especially where biological samples are concerned.
However, some fine-tuning is still needed. Although this first test showed that chemical mapping was possible on the submicrometer scale, the authors aim to go further down the scale and, in addition, improve the processing conditions for the cells. This would set the stage for direct, label-free chemical mapping of drug molecules within biological cells.