After binding, the S1 and S2 subunits are thought to dissociate, releasing S2, which helps membrane fusion of the virus to the host. However, before this can happen, S2 needs to change its structure to the fusion intermediate (FI) form to allow membrane fusion.
The FI has three N-terminal fusion peptides, which are inserted into the host cell membrane. Then the membranes are refolded, helping fusion, and the virus genome enters via fusion pore.
However, not much is known about the FI, in particular, its structure. One challenge is the short lifetimes of the FI. The lack of experimental data makes it difficult to do any computational studies of the intermediate. A better understanding of how the fusion process works in SARS-CoV-2 may help develop better therapeutics to combat emerging variants.
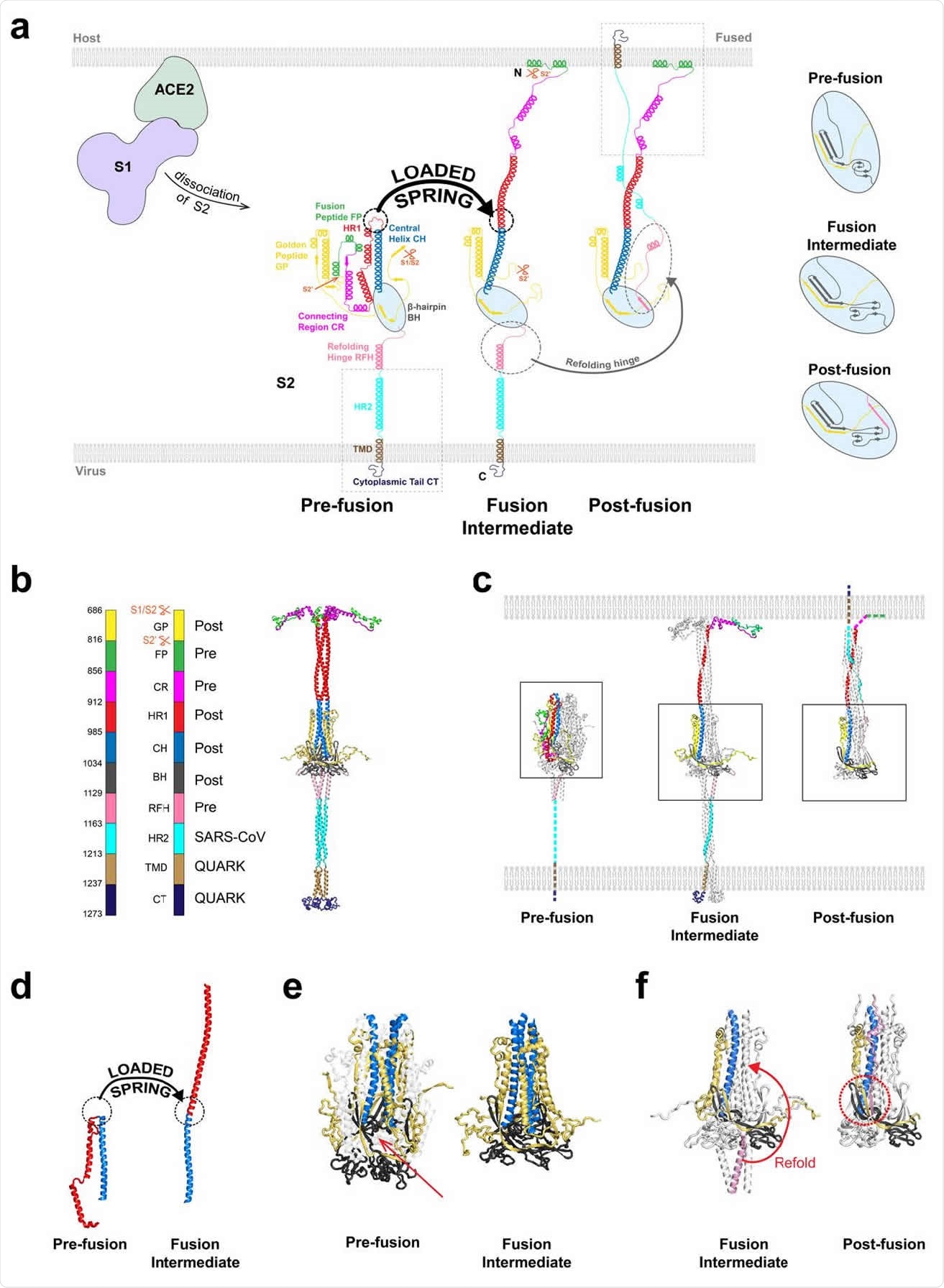
Model of the SARS-CoV-2 spike protein fusion intermediate (a) Model of the fusion intermediate (FI) of the CoV-2 spike (S) protein, schematic. One protomer of the S trimer is shown. Schematics of the pre-and postfusion S2 states from the known structures, other than unknown regions (boxed). Details of BH and adjacent domains, at right. Following dissociation of S2 from S1, the unstructured pre-fusion HR1 loops become helical (loaded spring release), giving the HR1-CH backbone that thrusts the FPs toward the host cell membrane. The FI subsequently refolds into the postfusion structure, driven by structural changes in RFH and chaperoned by GP, with RFH and HR2 providing leashes that pack the HR1-CH backbone grooves. (b) Model of the FI of the CoV-2 spike (S) protein, exact structure. Source structures for each S2 subunit domain are indicated, either CoV-2 prefusion (PDB: 6XR8), CoV-2 postfusion (PDB: 6XRA) or HR2 of CoV (PDB: 2FXP). Transmembrane domain (TMD) and cytoplasmic tail (CT) structures predicted by QUARK. (c) Comparison between predicted FI structure and known crystal structures of the prefusion (PDB: 6XR8) and postfusion (PDB: 6XRA) CoV-2 S2 subunit. One protomer highlighted in color. Dashed lines: missing domains from partially solved crystal structures. (d) Details of loaded spring transition. (e) β-hairpin (BH) domains in the known prefusion and predicted FI structures (boxed regions of (c)). The RFH domain is omitted from the FI BH for clarity. Following the loaded spring transition, HR1, CR and FP (shown faint in the prefusion BH) vacate their prefusion locations in BH. The resultant cavity (arrow) would presumably be unstable. We assume the FI adopts the more compact postfusion BH structure (right). (f) The golden peptide (GP) chaperones refolding of the fusion intermediate (FI) into the postfusion structure. Blowups of boxed regions in (c) are shown. Refolding of the refolding hinge (RFH) domain is guided by GP. RFH forms a parallel β-strand with GP (red circle), the RFH unstructured portion packs the CH-GP groove, and RFH helices interact with two GP helices. Colored BH and CH belong to one protomer; colored RFH belongs to a different protomer.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
A model for the fusion intermediate
With this aim, researchers from Columbia University in New York constructed a model for the SARS-CoV-2 FI using available data and computationally explored its behavior. The research is published on the bioRxiv* preprint server prior to peer review.
Based on how the influenza hemagglutinin, another viral fusion protein, behaves, the team thought the transition to the FI starts with changing all the heptad repeat 1 (HR1) domains into alpha-helices, so that the unstructured loops in HR1 become helical, like springs, becoming continuous with the central helix.
Using this and the structures of other components, the team came up with a model for FI. They tested their model using molecular dynamics simulations and found it to be stable. They found the regions below the β-hairpin domain to be highly flexible, allowing the FI to tilt.
The prefusion state likely changes to the FI state by a loaded spring release mechanism, where the loop-to-helix transitions become straight, extending the HR1 domain. Simulations of the fusion peptide determined an N-terminal helix with 10 residues that first contacts the membrane.
The author posits that the spring-loaded release occurs first. Then any refolding of the refolding hinge domain occurs after a delay, which allows the FI to change conformation to destabilize a refolding hinge domain into an unstructured loop.
Kinetics of the fusion intermediate
When the authors ran simulations to capture the dynamics of the FI, they found the FI showed significant configuration changes, sweeping a volume of about 25,000 nm3. They identified three hinge regions at the base of the FI, which allows the FI to be highly flexible so that it can cover a large volume of space looking for the membrane for fusion.
The hinges could have several important roles. They increase the accessible area available to the fusion peptides on the FI terminal, capturing the host cell membrane. The flexibility of the FI may also allow many FIs to capture the membrane at different locations and present the fusion peptides directly to the target membranes.
The researchers also found that secondary structures at the base of the FI were also highly dynamic. The three refolding hinge domain helices were spread out like an inverted tripod, buffering the longitudinal changes in the FI backbone. The spread-out helices also allowed upstream refolding hinge domains more flexibility.
Studies that have used isolated fusion peptides show a much rapid binding rate than seen in the FI model. Compared to what previous studies predict, about a 15 μs delay time for the virus to capture the membrane, the researchers did not see any membrane capture up to about 300 μs. This suggests binding of the fusion peptides is much slower when they are attached to the FI.
In addition, the team found that the golden peptide, a disconnected fragment between the S1/S2 site and the S2 site, may chaperone refolding. Monoclonal antibodies that target a region in the refolding hinge helix, which engages the golden peptide, have been seen before. This suggests the golden peptide or the refolding hinge may be potential targets for developing therapeutics against COVID-19. Unveiling the mechanisms governing the CoV-2 S2 fusion intermediate will play a vital role in the search for robust and pan-coronavirus antiviral drugs.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.