Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a positive-sense single-stranded (ssRNA) virus that belongs to the Coronaviridae family. It is also closely related to the Middle East respiratory syndrome coronavirus (MERS-CoV) and SARS-CoV-1. SARS-CoV-2 infection causes coronavirus disease 2019 (COVID-19) and develops into severe respiratory distress in a subgroup of patients. Severe COVID-19 causes sepsis-like manifestations, intense lung injury, and multi-organ failure, all associated with the overproduction of pro-inflammatory cytokines. This "cytokine storm" directly correlates with poor prognosis in this subgroup of patients.
This suggests that an overactive immune response may be responsible for unleashing virus-dependent immune pathology and inducing the defensive antiviral state mediated by type I IFN release. Thus, inborn errors in production and amplification of type I IFN or pre-existing auto-antibodies against the IFN family of cytokines were also found to be associated with poor prognosis.
Analyzing the SARS-CoV-2 genome to identify activators of endosomal TLR7/8 and MyD88 pathway
SARS-CoV-2-associated molecular patterns (SAMPs) and innate sensors have not been clearly defined so far. Researchers from the UK and Italy recently identified ssRNA fragments from the SARS-CoV-2 genome as direct activators of the endosomal TLR7/8 and MyD88 pathways. These same sequences induced DC activation in humans in terms of phenotype and functions, such as the production of IFN and cytokines and polarization of Th1. The study is posted to the preprint server, bioRxiv*.
"This study characterizes the first SARS-CoV-2 associated molecular patterns (SAMPs) and identifies the TLR7/8/MyD88 axis as a crucial pathway in the activation of human pDCs and cDCs."
The researchers performed a bioinformatic scan of the viral genome and identified hundreds of fragments potentially activating TLR7/8. This suggests that products of virus endosomal processing can possibly activate the IFN and inflammatory responses downstream.
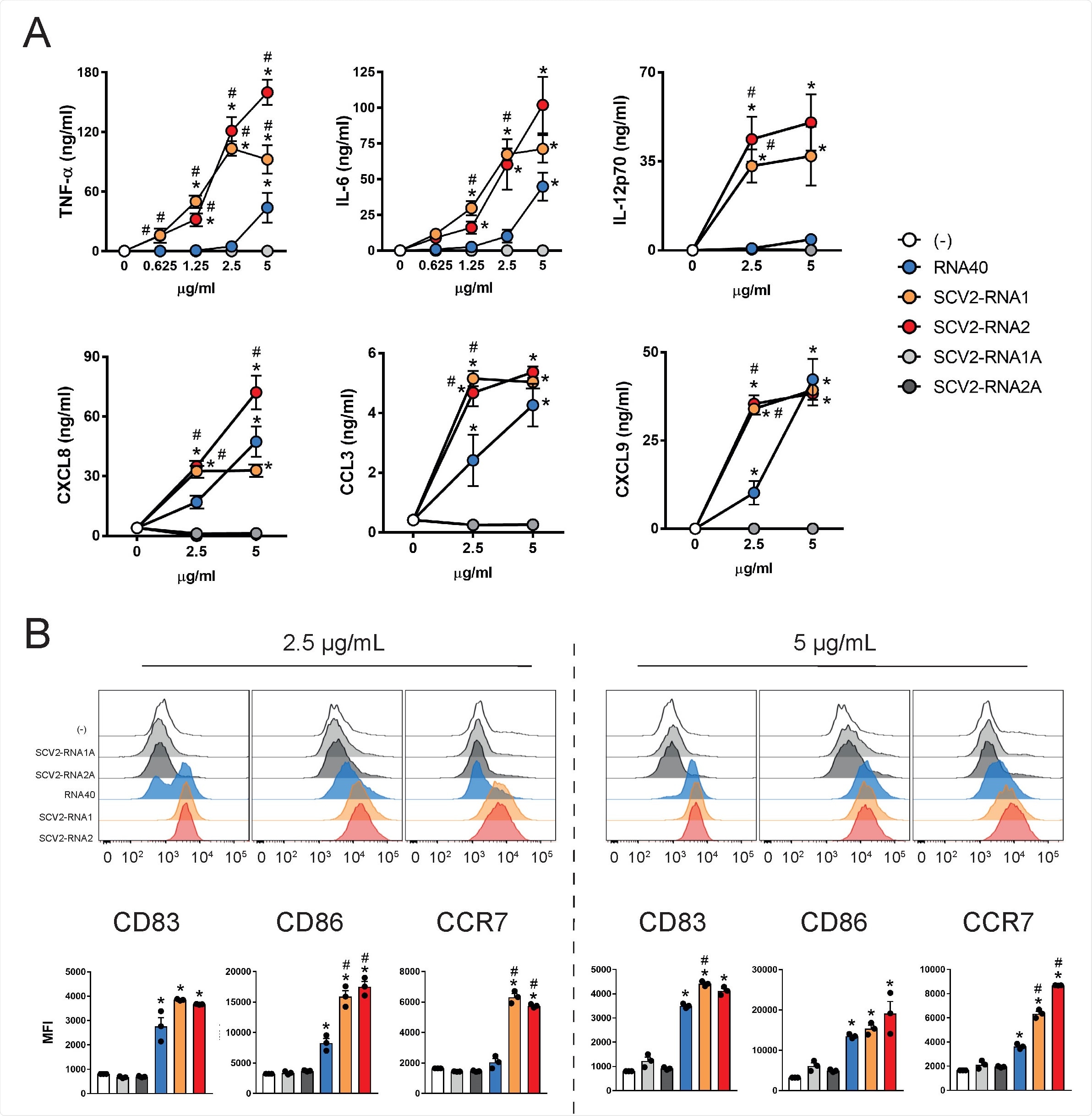
SAMPs activate cytokine secretion and phenotypical maturation of moDCs. (A) moDCs (2x106/ml) were stimulated with increasing concentrations of the indicated viral RNAs or with vehicle alone (-) for 24 hours. The production of TNF-α, IL-6, IL-12p70, CXCL8, CCL3 and CXCL9 was evaluated by ELISA in cell-free supernatants. Data are expressed as mean + SEM (n=3). Results of SCV2- RNA1A and SCV2-RNA2A are superimposed in all graphs. (B) moDCs were stimulated as described in (A) and the surface expression of CD83, CD86 and CCR7 evaluated by FACS analysis. Data are expressed as representative cytofluorimetric profiles (upper panels) or as the mean + SEM (n=3) of the Median of Fluorescence Intensity (MFI) (lower panels). (A-B) *P< 0.05 versus (-) by one-way ANOVA with Dunnet’s post-hoc test; # P< 0.05 versus RNA40 by paired Student’s t test.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
TLR7 and TLR8 are important cellular sensors of ssRNAs involved in host resistance and COVID-19 pathogenesis
SAMPs induced MyD88-dependent lung inflammation in vivo, characterized by the accumulation of pro-inflammatory and cytotoxic mediators, immune cell infiltration, and splenic DC phenotypical maturation. These results show that TLR7 and TLR8 are important cellular sensors of ssRNAs that are encoded by SARS-CoV-2 and are involved in host resistance and COVID-19 pathogenesis.
"DCs play a crucial role as activators of both inflammation, and adaptive immune responses and pDCs are the major producers of type I IFNs in response to viral infections."
This study identifies two short sequences in the SARS-CoV-2 ssRNA genome that induce the production of type I IFNs and the ability of human DCs to activate T cells by triggering endosomal TLR7 and TLR8. These sequences represent prototypical examples of the hundreds of other potential TLR ligands that have been identified with the help of a SARS-CoV-2 genome scan.
Findings are in line with results of previous works that analyzed SARS-CoV-2 nucleic acids
The study's findings are in agreement with previous study results that demonstrate a 20-fold higher density of GU-rich fragments in the closely related SARS-CoV-1 virus compared to HIV-1. The results also agree with a recent bioinformatic study that showed SARS-CoV-2 encodes many such fragments even larger than SARS-CoV. Therefore, endosomal processing of SARS-CoV-2 nucleic acids may produce multiple fragments that have the ability to trigger innate immune activation.
"ssRNA-sensing TLRs are also expressed by other innate immune cells, such as macrophages, as well as by peripheral tissues such as lung, bronchus, rectum, and cerebral cortex."
To summarize, this study demonstrates that SARS-CoV-2 is a potential source of immunostimulatory nucleic acid fragments and identifies the SARS-CoV-2-specific molecular patterns that can promote inflammation and immunity triggering TLR7 and TLR8. The authors believe that these findings fulfill a gap in understanding host-pathogen interaction in SARS-CoV-2 infection based on previous studies demonstrating the role of type I IFNs and TLR7 against life-threatening SARS-CoV-2 infection and the in vitro pDC activation by SARS-CoV-2.
"Since the magnitude of TLR activation differs in individuals, such as elderly people, differences in TLR activation may help explain differences in the quality of the antiviral immune response independently of SAMP potency."

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
SARS-CoV-2-associated ssRNAs activate inflammation and immunity via TLR7/8, Valentina Salvi, Hoang Oanh Nguyen, Francesca Sozio, Tiziana Schioppa, Mattia Laffranchi, Patrizia Scapini, Mauro Passari, Ilaria Barbazza, Laura Tiberio, Nicola Tamassia, Cecilia Garlanda, Annalisa Del Prete, Marco A. Cassatella, Alberto Mantovani, Silvano Sozzani, Daniela Bosisio, bioRxiv 2021.04.15.439839; doi: https://doi.org/10.1101/2021.04.15.439839, https://www.biorxiv.org/content/10.1101/2021.04.15.439839v1
- Peer reviewed and published scientific report.
Salvi, Valentina, Hoang Oanh Nguyen, Francesca Sozio, Tiziana Schioppa, Carolina Gaudenzi, Mattia Laffranchi, Patrizia Scapini, et al. n.d. “SARS-CoV-2–Associated SsRNAs Activate Inflammation and Immunity via TLR7/8.” JCI Insight 6 (18): e150542. Accessed April 24, 2022. https://doi.org/10.1172/jci.insight.150542. https://insight.jci.org/articles/view/150542.