The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), responsible for the current COVID-19 (Coronavirus Disease 2019) pandemic, has left public health systems worldwide in tatters.
An intensive effort has been made internationally to develop prophylactic vaccines and neutralizing antibodies (NAb) against the virus. There is, however, limited information about the SARS-CoV-2 virus-induced B cell immune response. Thus, understanding the principles of the B-cell response to the virus is essential for developing antiviral vaccines and NAbs.
In a new study published in the journal Human Immunology, scientists sought to better understand how B-cell immune repertoire changes over time during SARS-CoV-2 infection by comprehensively characterizing the dynamics of immunoglobin heavy chain (IGH) repertoire in COVID-19 patients.
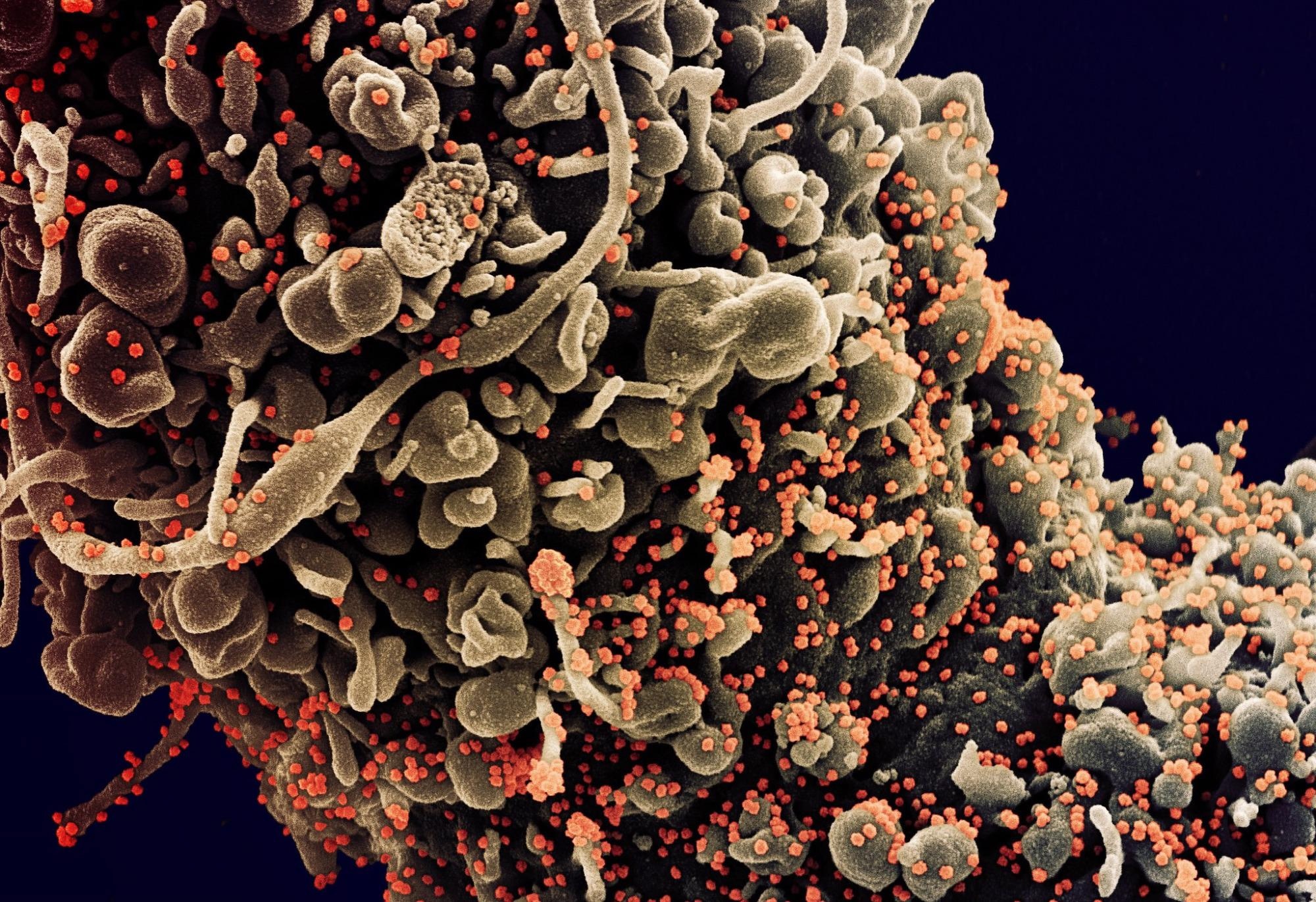 Study: Landscapes and dynamic diversifications of B-cell receptor repertoires in COVID-19 patients. Image Credit: NIAID
Study: Landscapes and dynamic diversifications of B-cell receptor repertoires in COVID-19 patients. Image Credit: NIAID
Background
B-cell surfaces have immunoglobin molecules called B-cell receptors (BCRs). They recognize and bind foreign antigens. B-cells become activated, multiple, and may differentiate into different cells on encountering their specific antigen. B-cells may get differentiated into short-lived effective antibody-secreting plasma cells or long-lived plasma and memory cells. Scientists have stated that analyzing BCR sequences is of great interest, as repertoires may have similar features on exposing individuals to the same pathogen, thereby, giving rise to convergent antibodies.
Recent developments in high-throughput sequencing have enabled characterization of the IGH repertoire in large numbers of samples. Researchers are using these tools to gain insights into the humoral responses in healthy individuals and a wide range of diseases. They are now able to understand how the antibody repertoire could change in response to disturbance arising from a viral infection, viral evolution, and vaccination. BCR repertoire analysis has been applied in the case of different viruses (e.g., influenza virus, human immunodeficiency virus, etc.), but the dynamics of antibody response elicited by SARS-CoV-2 infection remain to be determined.
A New Study
In the current study, scientists sought to understand how B-cell immune repertoire changed over time during SARS-CoV-2 infection. To this end, IGH repertoires from the peripheral blood samples were collected multiple times from five COVID-19 patients (longitudinal data).
First, the sequences were classified into clones by lineage clustering analysis. The next step was to track the evolution of certain characteristics, such as the unique number of CDR3, Shannon index, the number of high-frequency clones, cumulative frequency of the top 100 clones, and V, J-gene segment usage. In addition to the above, researchers also performed clonotype overlap, lineage expansion, and CDR3 sequence network structure. This was done to study the similar and dissimilar trends of IGH repertoire status over time.
Key Findings
Scientists found that a period of 2–3 weeks after symptom onset is of great importance to B-cell immune response. It was observed that each individual’s frequency of different V and J gene segment usage stayed relatively stable over time, hinting at the fact that V, J segment use might not be markedly altered by SARS-CoV-2 infection. The use of V components showed a differentiated preference which might signify that the human immune system is quite variable across individuals but relatively stable over time within a given individual. This result is in line with a previous study showing that IGHV3-23 was highly represented in patients and healthy controls. Contrary to the above findings, another reported that IGHV3-23 was over-represented in patients compared to the control group. Researchers also documented that IGHV4-34 is more frequently used in COVID-19 patients compared to healthy donors.
Bulk BCR sequencing could be more suitable for BCR gene usage frequency assessment, and this is one advantage of large amounts of sequencing data. However, the current study considered only five patients; therefore, this small sample size could most certainly limit the power of the analysis.
The current study reported that all patients’ BCR repertoire composition, in 2–3 weeks of onset, was quite different from that in other periods.
There were many more common clones and evident lineage expansion. Additionally, it was observed that each patient’s average degree and diameter indicators peaked around the said time frame.
It must be noted that this 2-3 week timeline is consistent with an antigen-induced B-cell differentiation and maturation. Scientists opined that the above observations could be due to strong neutralizing antibody responses and a significant expansion of virus-specific B-cell clones.
Collaboration among scientists from various countries has contributed to the progress in the development of COVID-19 therapeutics.
Research from the scientists of the current study revealed that there is a clear preference for sequence homology between clones and antibodies with neutralizing potential.
Clone expansion was observed early in one patient (the only severe case and aged 85 years). This led to the, yet to be tested, hypothesis that the earlier clone expansion might be correlated with the severity of clinical symptoms and age of the patients under consideration.
Conclusion
In this study, scientists have provided an extensive B cell deep immune profiling of COVID-19 patients.
The findings from this research should help further our understanding of the immune response of COVID-19 patients and provide the necessary basic knowledge for the development of novel vaccines and nAbs.