Since its emergence in 2019, the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), which is the virus responsible for the coronavirus disease 2019 (COVID-19), has had a significant influence on human life around the planet. The rapid discovery and widespread dissemination of effective vaccinations, as well as breakthroughs in the development of small molecule and monoclonal antibody-based treatments, have improved the understanding of how to control the current COVID-19 pandemic.
Background
However, on November 24, 2021, a new SARS-CoV-2 Omicron (B.1.1.529) variant was identified in South Africa. The World Health Organization (WHO) Technical Advisory Group on SARS-CoV-2 Virus Evolution recognized B.1.1.529 as the fifth variation of concern (VOC) and named it the Omicron variant on November 26, 2021.
Among all SARS-CoV-2 variants, Omicron has by far the most mutations. To this end, there are 32 mutations in this variant’s spike protein, which is the key viral component that determines the infectivity and antigenicity of the virus. Furthermore, 15 of the 32 mutations were found in the spike protein’s receptor-binding region (RBD).
These mutations include almost all key mutations that were identified in prior SARS-CoV-2 VOCs, such as K417N, E484A, and N501Y, as well as other known mutations that change the sensitivity of the virus to neutralization by protective antibodies. It has been proposed that the spike protein’s complicated mutations may allow immunity induced by prior infection or vaccination to be bypassed, thereby resulting in a large number of breakthrough infections or re-infections with mutated viral strains.
In a recent study, a team of researchers from the National Institutes for Food and Drug Control used vesicular stomatitis virus (VSV) vector and Omicron spike protein containing all 32 mutations to create the pseudotyped SARS-CoV-2 Omicron variant (PV-Omicron). The neutralization sensitivity of PV-Omicron was tested against serum samples from a panel of COVID-19 convalescent patients, with PV-D614G (S protein from SARS-CoV-2 D614G strain) serving as the baseline neutralization reference.
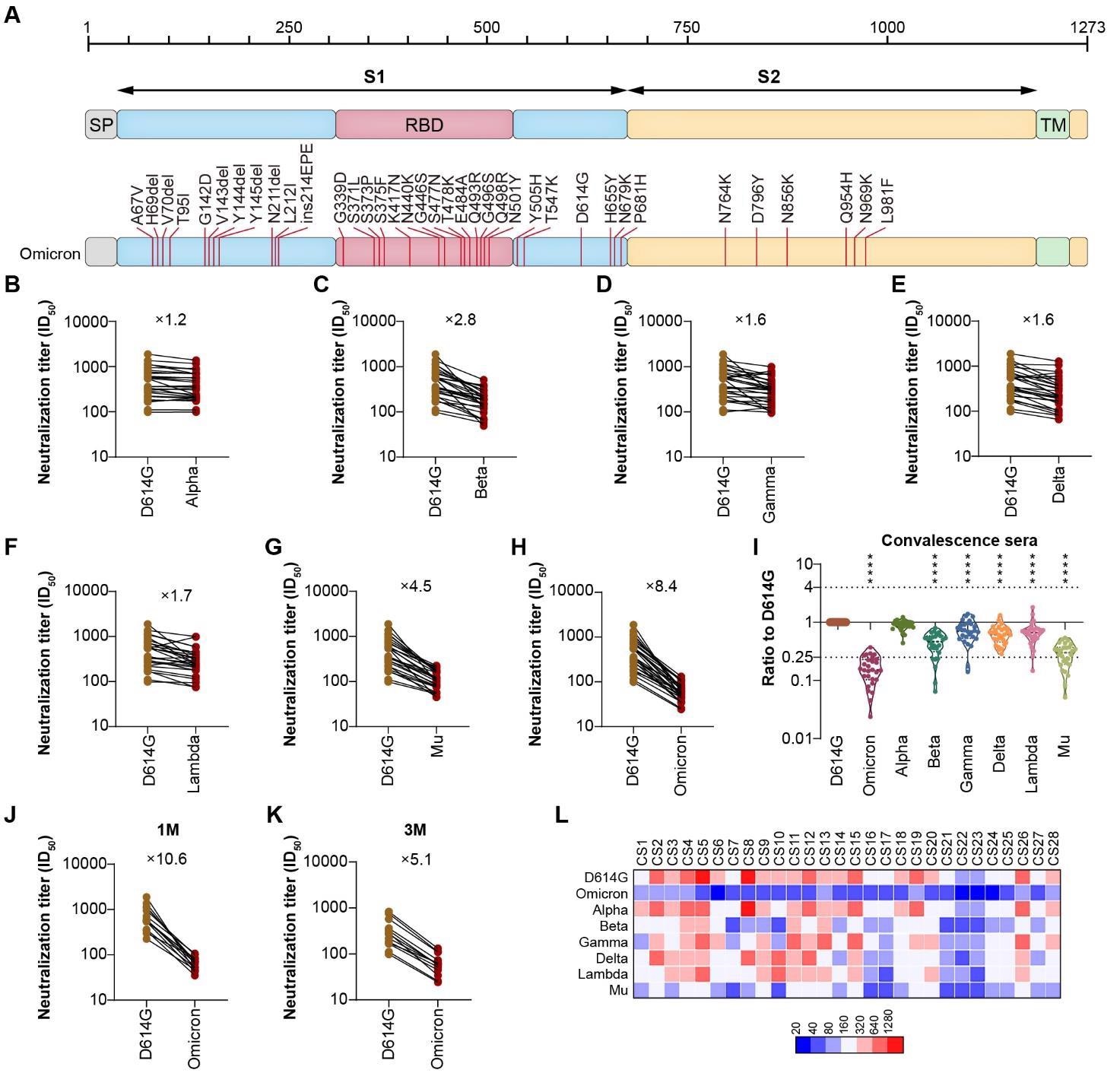 A. Schematic Illustration of Omicron Spike. All of the 32 mutations were located on Omicron Spike, which was used to construct the pseudotyped Omicron virus. B.H. The neutralization analysis of convalescent sera against Omicron, VOCs and VOIs. The neutralization activity of 28 convalescent sera from COVID-19 patients was tested. The neutralization ED50 and ratio compared to the reference strain D614G was also displayed as indicated. Data represented ED50 of three independent experiments. I. The comparison of neutralization activity against different VOCs and VOIs. J-K. The neutralization sensitivity of sera collected from convalescent patients at different time point against Omicron. The neutralization ED50 and ratio compared to the reference strain D614G was also displayed as indicated. Data represented ED50 of three independent experiments. J sera were collected 1-month after recovery; K, sera were collected 3-month after recovery; L. The heatmap of individual neutralization data. Data represented ED50 of three independent experiments. The Red to blue colour represented ED50 high to low as shown in the scale bar. CS1-CS15 sera were collected 1-month after recovery; CS16-28 sera were collected 3-month after recovery.
A. Schematic Illustration of Omicron Spike. All of the 32 mutations were located on Omicron Spike, which was used to construct the pseudotyped Omicron virus. B.H. The neutralization analysis of convalescent sera against Omicron, VOCs and VOIs. The neutralization activity of 28 convalescent sera from COVID-19 patients was tested. The neutralization ED50 and ratio compared to the reference strain D614G was also displayed as indicated. Data represented ED50 of three independent experiments. I. The comparison of neutralization activity against different VOCs and VOIs. J-K. The neutralization sensitivity of sera collected from convalescent patients at different time point against Omicron. The neutralization ED50 and ratio compared to the reference strain D614G was also displayed as indicated. Data represented ED50 of three independent experiments. J sera were collected 1-month after recovery; K, sera were collected 3-month after recovery; L. The heatmap of individual neutralization data. Data represented ED50 of three independent experiments. The Red to blue colour represented ED50 high to low as shown in the scale bar. CS1-CS15 sera were collected 1-month after recovery; CS16-28 sera were collected 3-month after recovery.
About the study
There are 32 mutations in the spike protein of the Omicron variant, including A67 V, H69del, V70del, T95I, G142D, V143del, Y144del, Y145del, N211del, L212I, ins214EPE, G339D, S371L, S373P, S375F, K417N, N440 K, G446S, S477N, T478 K, E484A, Q493R, G496S, Q498R, N501Y, Y505H, T547 K, D614G, H655Y, N679 K, P681H, N764 K, D796Y, N856 K, Q954H, N969 K, and L981F. The PV-Omicron used in the current study was constructed to contain all 32 mutations.
In addition to the Omicron variant, the researchers also studied and reported were pseudotyped viruses expressing spike proteins from the other current VOCs including the Alpha, Beta, Gamma, and Delta variants, as well as two variants of interest (VOIs) including the Lambda and Mu strains. The PV-Omicron was tested against a panel of human sera from convalescent COVID-19 patients who had been infected with the original SARS-CoV-2 strain to determine the neutralization sensitivity of the Omicron variant and how it has evolved as compared to the other SARS-CoV-2 variants.
In comparison to the reference strain PV-D614G, the mean neutralization (ED50) titer of PV-Omicron was 66, which represented an 8.4-fold reduction in neutralization.
Meanwhile, the authors tested various VOCs and VOIs for neutralization sensitivity against the same panel of human serum samples. The neutralizing activity of the Delta variant, which is currently the dominant circulating strain in 99% of, has only decreased 1.6-fold. The ED50 of Alpha, Beta, and Gamma variants is reduced by about 1.2, 2.8, and 1.6 times, respectively, as compared to the baseline reference strain PV-D614G, whereas neutralization is reduced by 1.7 and 4.5 times for Lambda and Mu variants, respectively.
Individual variations in patient sera against the Omicron variant or other VOCs were also discovered. In the case of PV-Omicron, the ED50 has dropped below 80 in the majority of patient sera.
Although the average ED50 of the one-month group is nearly two-fold greater than that of the three-month group after recovering from COVID-19, both titers were reduced to the same level when the serum was collected at one-month and three-month after recovery. As a result, the one-month group's lower neutralization sensitivity to PV-Omicron is more noticeable at 10.6-fold as compared to the three-month groups.
Implications
More factors, such as the infectivity of the Omicron variant in the human populations as compared to other variants and the viral fitness of Omicron after humans are infected, may influence the exact impact on human protection. To properly establish the global impact of Omicron on the containment of the COVID-19 pandemic, more population studies are needed, including those that determine the amount of immune protection and symptoms among those infected with Omicron.
The current study confirmed the Omicron variant's heightened immune escape, thus demonstrating the ability of this variant to have potentially significant consequences for public health planning around the globe.
Journal reference:
- Zhang, L., Quianqian, L., Liang, Z., et al. (2021). The significant immune escape of pseudotyped SARS-CoV-2 Variant Omicron. Emerging Microbes & Infections. doi:10.1080/22221751.2021.2017757.