A recent study by scientists at Celentyx Ltd, the University of Birmingham, and the University of Warwick and posted to the bioRxiv* preprint server illustrated a cell-based severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) spike (S) protein attachment assay.
A crucial step for SARS-CoV-2 entrance into human cells is the attachment of the SARS-CoV-2 S protein with the angiotensin-converting enzyme 2 (ACE2) receptor. Therefore, preventing this engagement is a significant determinant in the therapeutic effectiveness of serum antibodies induced by the coronavirus disease 2019 (COVID-19) vaccination and therapeutic monoclonal antibodies.
Moreover, the appearance of SARS-CoV-2 variants like Omicron and Delta has made it necessary to develop flexible assays that could assess the efficacy of COVID-19 therapeutics. Arising SARS-CoV-2 variants might show resistance to or evasion from neutralization by COVID-19 monoclonal antibodies or antibodies produced by vaccination centered on the initial SARS-CoV-2 strain. Indeed, understanding these constraints guides political and health strategies during the societal challenges brought on by the COVID-19 pandemic.
About the study
The goal of the present study was to establish a reductionist cell-based framework suited for high-throughput assessment that measures inhibition of SARS-CoV-2 S protein adherence to mammalian cells appropriate to aid immune tracking and development of therapeutics.
Initially, the team explored the attachment of a recombinant His-tagged SARS-CoV-2 S protein (S A) to lung epithelial A549 cell lines, either over-expressing ACE2 (A549-ACE2), wild-type (A549-WT), or over-expressing ACE2 and transmembrane protease, serine 2 (TMPRSS2) (A549-ACE2-TMPRSS2). The His-tagged S protein was evaluated utilizing a mouse immunoglobulin G (IgG) anti-His antibody, a secondary anti-mouse IgG antibody coupled to Alexa Fluor 488 (AF488) dye, and high-content imaging measuring the intensity of S protein labeling per cell. Subsequently, the attachment of several recombinant S proteins to A549-ACE2 and A549-WT cells was analyzed.
The effects of various widely available anti-S antibodies on the labeling of ACE2-A549 cells by S A and S F were evaluated to examine the effectiveness of the currently designed assay for monoclonal antibody testing. Further investigations were proceeded using three antibodies. The efficacy of receptor-binding domain 8 (RBD8), RBD1, and RBD7 antibodies to neutralize SARS-CoV-2 infection of Vero cells was explored to gauge the translational significance of the assay.
The performance of the S binding assay in plasma from individuals was analyzed with anti-S IgM/G/A measured by enzyme-linked immunosorbent assay (ELISA) to increase the potential utility of this assay. In addition, a score ≥ 1.0 was regarded as positive in this evaluation. Besides, the performance of RBD8, RBD14, and RBD7 was compared against the SARS-CoV-2 Omicron and Delta S protein labeling of A549-ACE2 cells.
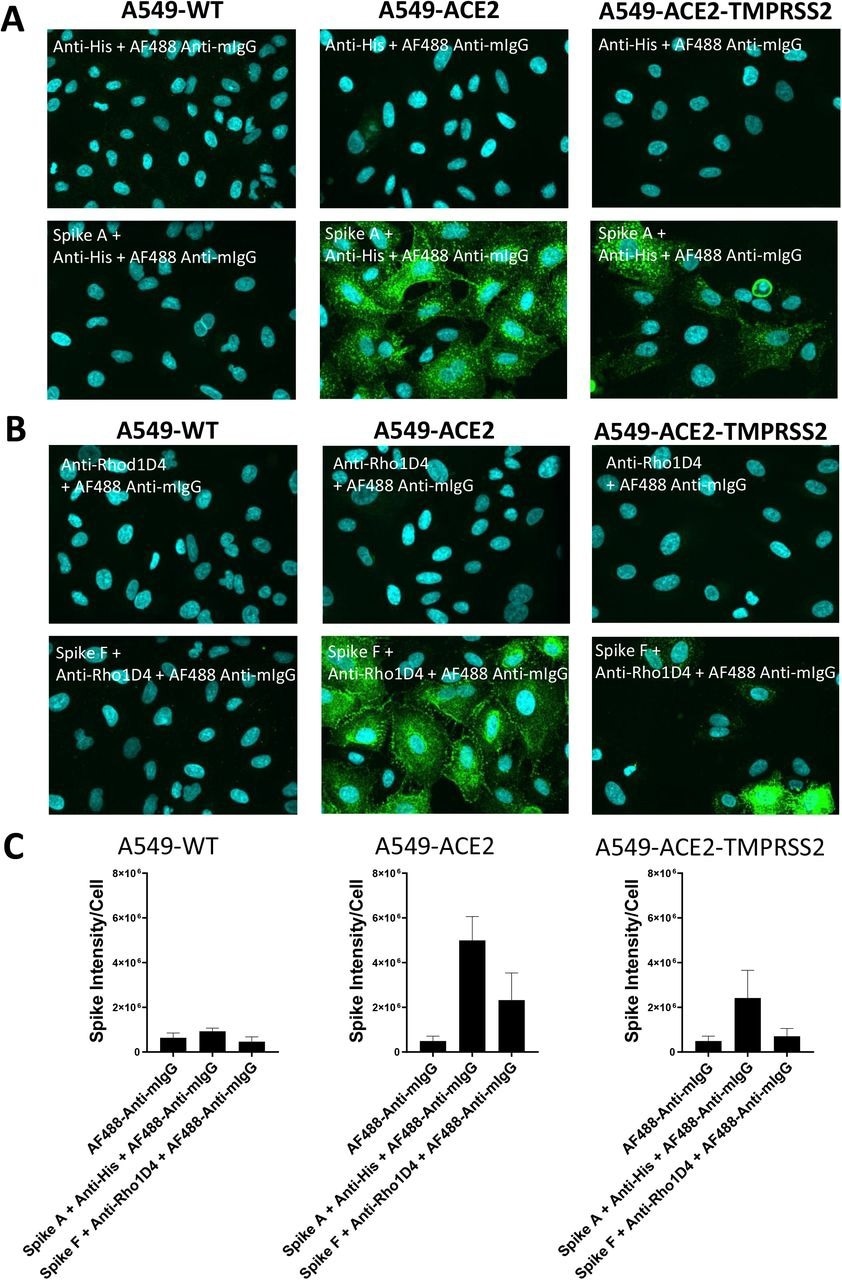
Labeling of ACE2 expressing A549 cells by recombinant spike proteins (A) Representative confocal images showing labeling of wild type (A549-WT), ACE2-overexpressing (A549-ACE2) and ACE2 and TMPRSS2-overexpressing (A549-ACE2-TMPRSS2) A549 cells by recombinant spike protein A (‘Spike A’) and anti-His + AF488-conjugated anti-mIgG detection antibodies. (B) Representative confocal images showing labeling of A549 cells (A549-WT), ACE2-overexpressing (A549-ACE2) and ACE2 and TMPRSS2-overexpressing (A549-ACE2-TMPRSS2) A549 cells by recombinant spike protein F (‘Spike F’) and anti-His + AF488-conjugated anti-mIgG detection antibodies (green; nuclei in cyan). (C) Quantification of spike protein labeling intensity from images acquired by high content confocal microscopy for cells labeled in (A) and (B). Data expressed as mean + SEM from three independent experiments for A549-ACE2 and from two independent experiments for A549-WT.
Results and conclusions
The study results showed that while A549-WT cell labeling with S protein was similar to background concentrations, both A549 cells overexpressing ACE2 and TMPRSS2 and those overexpressing ACE2 had punctate labeling of S protein. The S protein labeling was less intense on A549-ACE2-TMPRSS2 cells than on A549-ACE2, which might be due to different levels of ACE2 expression in the cell lines. An anti-mouse IgG secondary antibody coupled to AF488 and an anti-RhoD1A4 antibody showed clear labeling upon subsequent use of an S protein inside a detergent micelle (S F), which presumably comprises the trimeric S in its native structure.
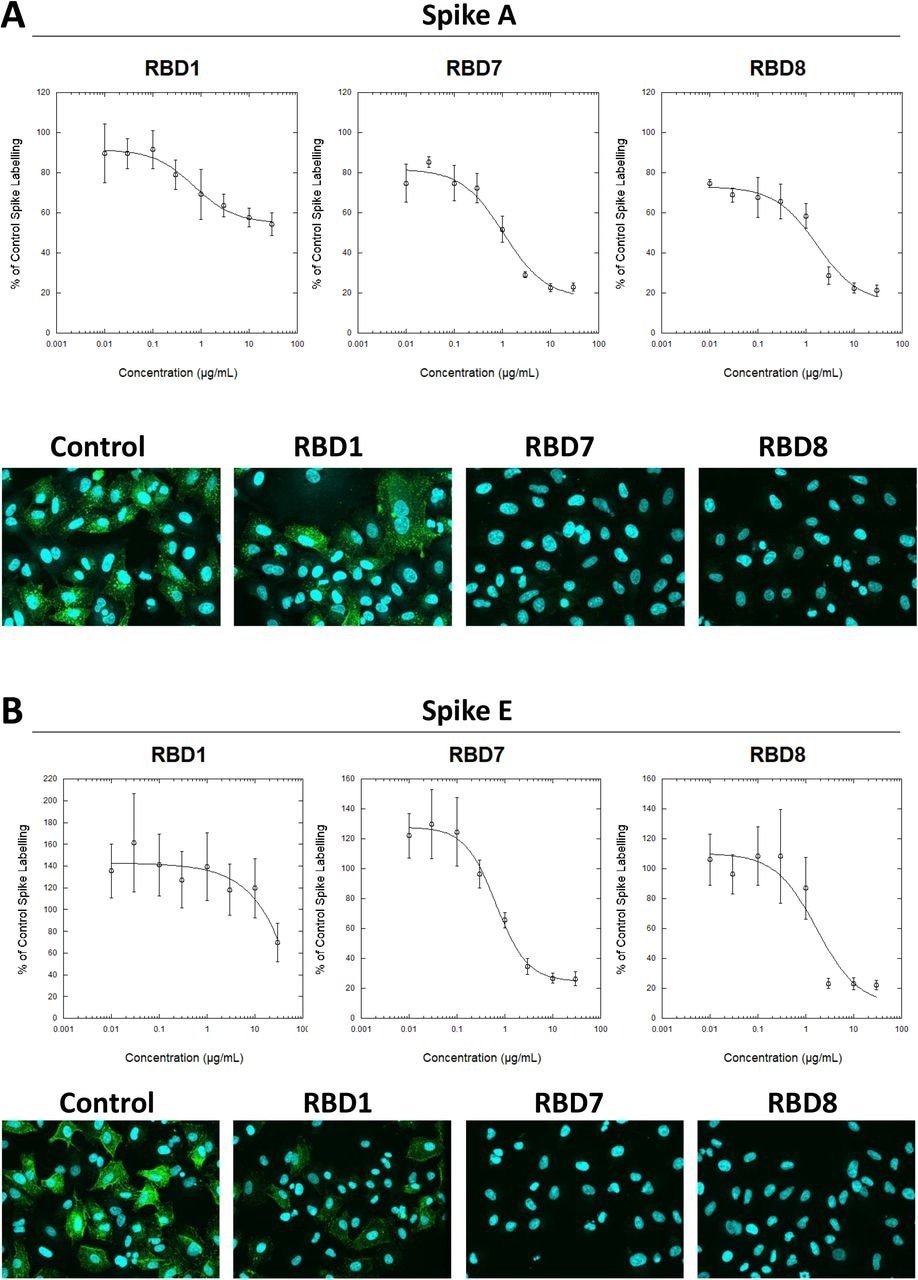
Characterization of spike blocking antibodies Labelling of A549-ACE2 cells by recombinant spike protein A (A) or E (B) that had been pre-incubated in the absence or presence of the indicated concentrations of antibodies RBD1, RBD7 or RBD8, followed by detection using anti-His + AF488-conjugated anti-mouse IgG antibodies, and visualization and quantification of spike protein labeling intensity by confocal microscopy. Plots show data expressed as mean ± SEM of three independent experiments. A solid line represents nonlinear regression using a 4-parameter logistic equation. Representative confocal images show labeling of cells under control conditions or with spike protein pre-incubated with 10 μg/mL RBD1, RBD7 or RBD8.
Subsequently, the analysis of various recombinant S proteins in A549-ACE2 and A549-WT cells displayed that adherence to A549-WT was typically poor. Nevertheless, different labeling intensities were visible in A549-ACE2 cells, with some recombinant S proteins showing a weak signal. These results demonstrated that some recombinant S proteins permitted clear labeling of A549-ACE2 cells.
The labeling of both S A and S F was decreased consistently by antibodies RBD5, RBD1, RBD8, and RBD7. Titration of RBD7, RBD1 and RBD8 antibodies demonstrated concentration-reliant blockage of S protein A and E labeling, enabling quantification of each antibody's half-maximal inhibitory concentration (IC50) values. RBD1 had the lowest strength to neutralize viral infection, mirroring the S labeling results with S A; however, RBD8 and RBD7 successfully stopped the viral infection, with RBD8 having higher potency than in the S binding experiment. Additionally, human plasma samples with anti-S IgA/M/G levels above three showed concentration-dependent S blocking action; on the other hand, those with anti-S IgA/M/G levels less than three did not. These findings imply that the present S binding assay could characterize monoclonal antibodies and track neutralizing antibody function in plasma.
Furthermore, the authors discovered that while RBD14, RBD7, and RBD8 significantly decreased labeling by the Delta S protein, just RBD14 showed a similar efficiency against the Omicron S protein.
In summary, the team has created a high-content imaging technique to measure SARS-CoV-2 S protein attachment to target cells expressing ACE2 in the current investigation. The investigators envisage that the cell-based S protein assay established in this research would supplement existing pseudovirus and biochemical assays in COVID-19 therapeutic development. It could also be modified to work with other cell kinds to explore the potentially ACE2-autonomous adherence of the SARS-CoV-2 S protein.

 *Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
*Important notice: bioRxiv publishes preliminary scientific reports that are not peer-reviewed and, therefore, should not be regarded as conclusive, guide clinical practice/health-related behavior, or treated as established information.
Journal reference:
- Preliminary scientific report.
A cell-based, spike protein binding assay highlights differences in antibody neutralising capacity for SARS-CoV-2 variants; Neale Harrison, Lauren Richardson, Chiara Pallini, Ines Morano, Elizabeth Jinks, Jamie Cowley, Hujo Chan, Harriet J Hill, Cristina Matas de las Heras, Ana Teodosio, Andrea S Lavado, Timothy R Dafforn, Dimitris K Grammatopoulos, John Gordon, Catherine A Brady, Lawrence S Young, Nicholas M Barnes, Zania Stamataki, Omar S Qureshi. bioRxiv preprint 2022, DOI: https://doi.org/10.1101/2022.06.24.496409, https://www.biorxiv.org/content/10.1101/2022.06.24.496409v1