The high infectivity and rapid transmissibility of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection has called for various tests. Early in the coronavirus disease 2019 (COVID-19) pandemic, caused by the SARS-CoV-2 infection, scientists tested everywhere for live virus contamination hazards – skin, food, water, door handles, surfaces made of copper, stainless steel, cardboard, and plastic.
However, there is still a paucity of data regarding the survival and infectivity of the SARS-CoV-2 on surfaces in closed environments. Also, the environmental viability of SARS-CoV-2 is still not fully understood.
While qRT-PCR (quantitative reverse transcriptase-polymerase chain reaction) studies showed the presence of viral RNA indoors for up to several weeks, the lack of viral RNA detected in an oncology ward housing patients with COVID-19 is surprising. Contradictory observations undermine the significance of the presence and infectivity of this virus on environmental surfaces.
In this context, an interdepartmental team from the University of California Davis, USA, analyzed the surface swabs for SARS-CoV-2 RNA and the infectivity in a hospital setting by both qRT-PCR and a viral culture assay. They also determined the suitability for sequence analysis and phylogenetically identifying the source of the virus.
The team claims this is the first report on recovering near-complete SARS-CoV-2 genome sequences directly from environmental surface swabs.
They found a low likelihood of the SARS-CoV-2 contamination on hospital surfaces with an infectious virus, disputing the importance of ‘fomites’ in COVID-19 transmission. Also, there was no detectable infectious virus from the surfaces; this is in agreement with the previous studies.
However, the researchers found almost complete genome sequences from two surfaces, suggesting that some viral genomes are present even though not infectious. The researchers amplified the viral sequences from several samples, which appeared negative by qRT-PCR. Interestingly, this highlights the potential to obtain viral sequences in some PCR negative samples.
The phylogenetic analysis helped the team identify the source of the positive SARS-CoV-2 genomes - they were derived from hospitalized patients. Notably, the environmental contamination was linked to a single lineage of the virus, most likely from a single patient.
For this study, the team collected the samples during the two waves of COVID-19 at the University of California, Davis Medical Center, in COVID-19 patient serving and staff congregation areas. They collected the first set of samples in April 2020 and the second set between late July and early August 2020.
They investigated the qRT-PCR positive samples in Vero cell cultures for cytopathic effects and then phylogenetically assessed the samples by whole genome sequencing. The complete protocol used here is made available online.
Even though we recovered near-complete genome sequences in some, none of the positive samples (11 of 224 total) caused cytopathic effects in cultured cells suggesting this nucleic acid was either not associated with intact virions, or they were present in insufficient numbers for infectivity.”
In this study, the researchers found improved cleaning and patient management practices between April and August 2020 were associated with decreased recovery of SARS-CoV-2 RNA from hospital surface samples. A substantial reduction of SARS-CoV-2 qRT-PCR positivity (from 11% to 2%) in hospital surface samples is observed.
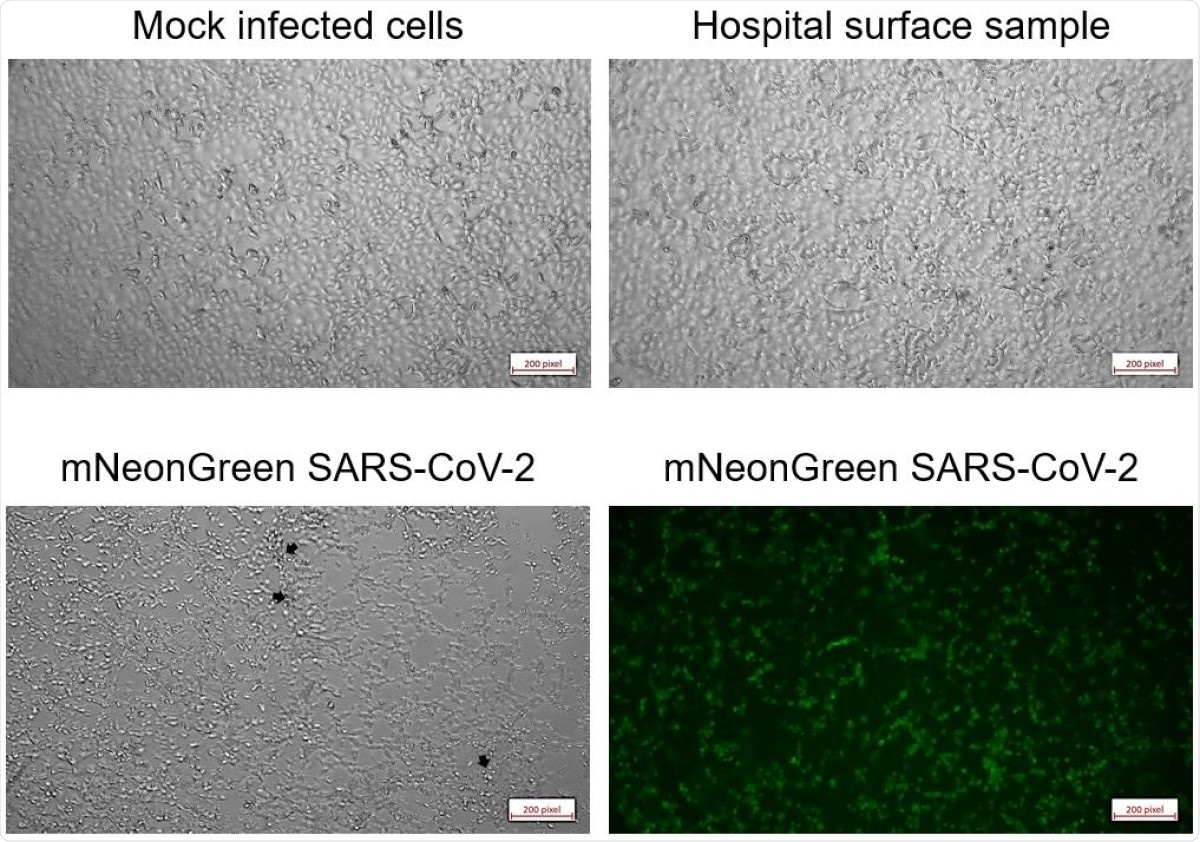
Micrographs of Vero E6 cells five days after inoculation. Cells were either mock-infected (upper left), inoculated with swab samples (representative of all five tested samples, upper right), or infected with one PFU of mNeonGreen SARS-CoV-2 (phase contrast, lower left; mNeonGreen lower right).
The improved patient management of respiratory secretions included earlier intubation, rapid sequence ventilation, and changes in the management of high O2 flow nasal cannulas.
The researchers highlighted that in mid-August, 2020, when the second surge of COVID-19 cases was admitted, substantially increasing the number of patients in the hospital.
However, the improved cleaning practices and patient management helped reduce the environmental presence of the virions.
They also observed that the SARS-CoV-2 RNA from the hospital surfaces did not exhibit infectious nature in a Vero cell culture model in vitro.
The researchers reported that the whole-genome PCR and sequencing yields more effective detection of SARS-CoV-2 than the qRT-PCR. The genomic sequences isolated from qRT-PCR negative samples indicate a superior sensitivity of viral detection by sequencing.
From two different patient rooms (from the floor and a soiled linens basket lid), they also recovered two near-complete genomes.
Phylogenetic analysis suggested that the SARS-CoV-2 genomes of the positive samples were derived from hospitalized patients. The researchers noted that the recovered genome sequences were from the clade 19B - that may have originated from a single patient or multiple patients infected with similar viruses.
In this study, the researchers reported that the 11% of samples collected at the UC Davis Medical Center in April 2020 were positive for SARS-CoV-2, whereas a larger follow-up experiment in August found only 2% of swabs positive. This is likely due to improved cleaning protocols and improved management of patient respiratory secretions. Some viral genomes are present but infectious. This study highlights the potential to obtain viral sequences in some PCR negative samples.
In the ongoing fight against the pandemic, this study throws light on the importance of protocols and procedures in research studies. The results from the unique observations will help the healthcare workers, and researchers adopt better SOPs.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
David A. Coil, Timothy Albertson, Shefali Banerjee, Greg Brennan, A.J. Campbell, Stuart H. Cohen, Satya Dandekar, Samuel L. Díaz-Muñoz, Jonathan A. Eisen, Tracey Goldstein, Ivy R. Jose, Maya Juarez, Brandt A Robinson, Stefan Rothenburg, Christian Sandrock, Ana M. M. Stoian, Daniel G Tompkins, Alexandre Tremeau-Bravard, Angela Haczku. SARS-CoV-2 detection and genomic sequencing from hospital surface samples collected at UC Davis. medRxiv 2021.02.23.21252022; doi: https://doi.org/10.1101/2021.02.23.21252022, https://www.medrxiv.org/content/10.1101/2021.02.23.21252022v1
- Peer reviewed and published scientific report.
Coil, David A., Timothy Albertson, Shefali Banerjee, Greg Brennan, A. J. Campbell, Stuart H. Cohen, Satya Dandekar, et al. 2021. “SARS-CoV-2 Detection and Genomic Sequencing from Hospital Surface Samples Collected at UC Davis.” Edited by Binod Kumar. PLOS ONE 16 (6): e0253578. https://doi.org/10.1371/journal.pone.0253578. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0253578.