On the 11th of March 2020, the World Health Organization (WHO) announced Coronavirus Disease 2019 (COVID-19) to be a pandemic. This disease is highly infectious and is caused by a novel positive-stranded β-coronavirus, namely, Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Globally, this virus has had a devastating effect on public health and the economy. Scientists believe that only the discovery of effective vaccines and potential drugs can contain the ongoing pandemic.
Despite the commencement of vaccination programs in many countries, there is a need for more immunization strategies as the demand is exceptionally high, and there are certain limitations in the mass production of vaccines.
Many more new vaccines are in the clinical trial stage, but the development of a vaccine is complex. Several detailed evaluations are required at each stage of vaccine production ahead of receiving approval from regulatory bodies such as the United States Food and Drug Administration (FDA). These evaluations focus on a number of factors, such as adverse effects, efficacy levels, dosing procedures, storage requirements, the emergence of SARS-CoV-2 variants, and induction of long-term immunological memory. The above complexities enhance the need for drug repurposing, which deals with the re-utilization of existing drugs where their side effects are well established. This is an operational strategy that could reduce the SARS-CoV-2 viral load and be effectively used for COVID-19 treatment.
While studying the immunopathology of COVID-19, researchers found that it causes non-hemostatic inflammation associated with cytokine storm. The cytokine storm occurs via several mediators such as interleukin-6 (IL-6), which is also considered a biomarker for disease severity. For drug discovery, computational biology and bioinformatics are the two most powerful tools for repurposing approved drugs or drugs under clinical trials. In the current scenario, some of the strategies employed to discover drugs for COVID-19 disease are as follows:
- Identification of host or virus targets, for example, the receptor-binding domain (RBD) present in spike glycoprotein and angiotensin-converting enzyme II (ACE2), which facilitate virus-host cell interaction.
- Identification of proteins or enzymes from virus biosynthesis machinery. For example, main protease (Mpro) and RNA-dependent RNA polymerase (RdRp).
- Reversal of the host gene expression, which is associated with SARS-CoV-2-infection.
A new study published on the bioRxiv* preprint server focuses on the in-silico predictions to repurpose existing drug compounds, that may have the ability to reverse the SARS-CoV-2 gene expression induced in host cells. These drugs could target proteins or enzymes vital for the SARS-CoV-2 life cycle or inhibit the expression of Interleukin 6 (IL-6) that acts as both a pro-inflammatory cytokine and an anti-inflammatory myokine. Thereby, these drugs could be used as potential agents for the treatment of COVID-19 disease.
In this study, the researchers identified 39 repurposed drug candidates that show promise as effective treatments for COVID-19. These drug compounds are potentially capable of reverting the genetic signature of SARS-CoV-2-infected cells and show high affinity towards viral targets (protein or enzymes). Celastrol was found to be one of the best-matched drugs that can restrain the release of the virion particles and the secretion of IL-6 by SARS-CoV-2-infected cells. Currently, both computational and experimental studies have substantiated the findings of a previous study, which had claimed the effectiveness of celastrol as a potent drug for COVID-19 treatment.
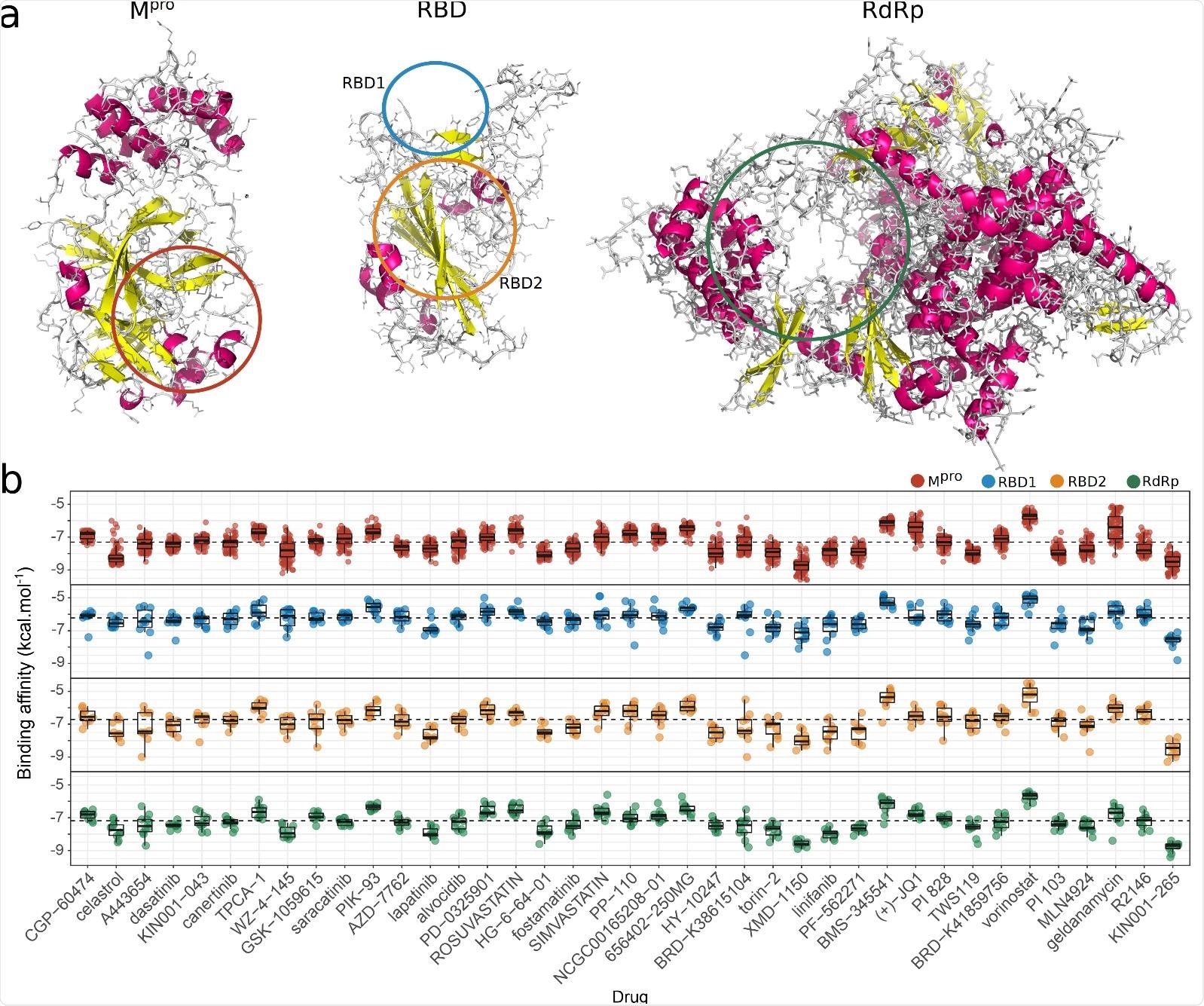
Docking analysis of molecular interactions between viral targets and drugs capable of reversing SARS-CoV-2 genetic signature. (a) Representative structures of the SARS-CoV-2 molecular targets (Mpro, RBD, and RdRp), where atoms are represented as lines and secondary structures as a cartoon with helices highlighted in magenta and sheets in yellow. Colored line circles (red - Mpro; RDB1 - blue; RBD2 - orange; RdRp - green) indicate the binding sites used for docking related to known inhibition of critical regions from each viral target 19,24–27. (b) Boxplots illustrate the affinity binding energies (kcal.mol-1) obtained from docking analysis between several structural conformations of 39 drugs and each viral site (Mpro = 83; RDB1= 10; RBD2 = 10; and RdRp = 10 structures). The reverser drugs were sorted based on decreasing order of Qscore that indicated their potential to revert the genetic signature of SARS-CoV-2 infection. Dotted lines indicate median affinity binding energies defined for each viral target (Mpro = -7.3; RDB1= -6.2; RBD2 =-6.7; and RdRp = -7.2 kcal.mol-1) considering all investigated drugs.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This drug possesses anti-inflammatory and antiviral properties. Celastrol is a triterpene that is also effective for HIV-1 Tat-induced inflammatory responses. Previous research has revealed that celastrol blocks the production of pro-inflammatory chemokines, e.g., CXCL1. It can also suppress replication of the Dengue virus serotype by activating IFN-α expression and triggering downstream antiviral responses. The in-silico predictions have also detrained other drugs, such as CGP-60474, dasatinib, canertinib, geldanamycin, etc., that have the potential to treat COVID-19 disease.
Previous studies as well as the current research, have identified several genes where expressions are altered during SARS-CoV-2 infection. The production of pro-inflammatory cytokines (IL-1β, IL-6, TNF-α, etc.), chemokines (CXCL1 and CCL20), and genes related to the NF-κB pathway are modified post-infection. These modifications are directly correlated with the progression and severity of COVID-19. The present research further validated the in-silico findings with in vitro experiments. Researchers have reported that celastrol can reduce the viral load of SARS-CoV-2 infected cell lines.
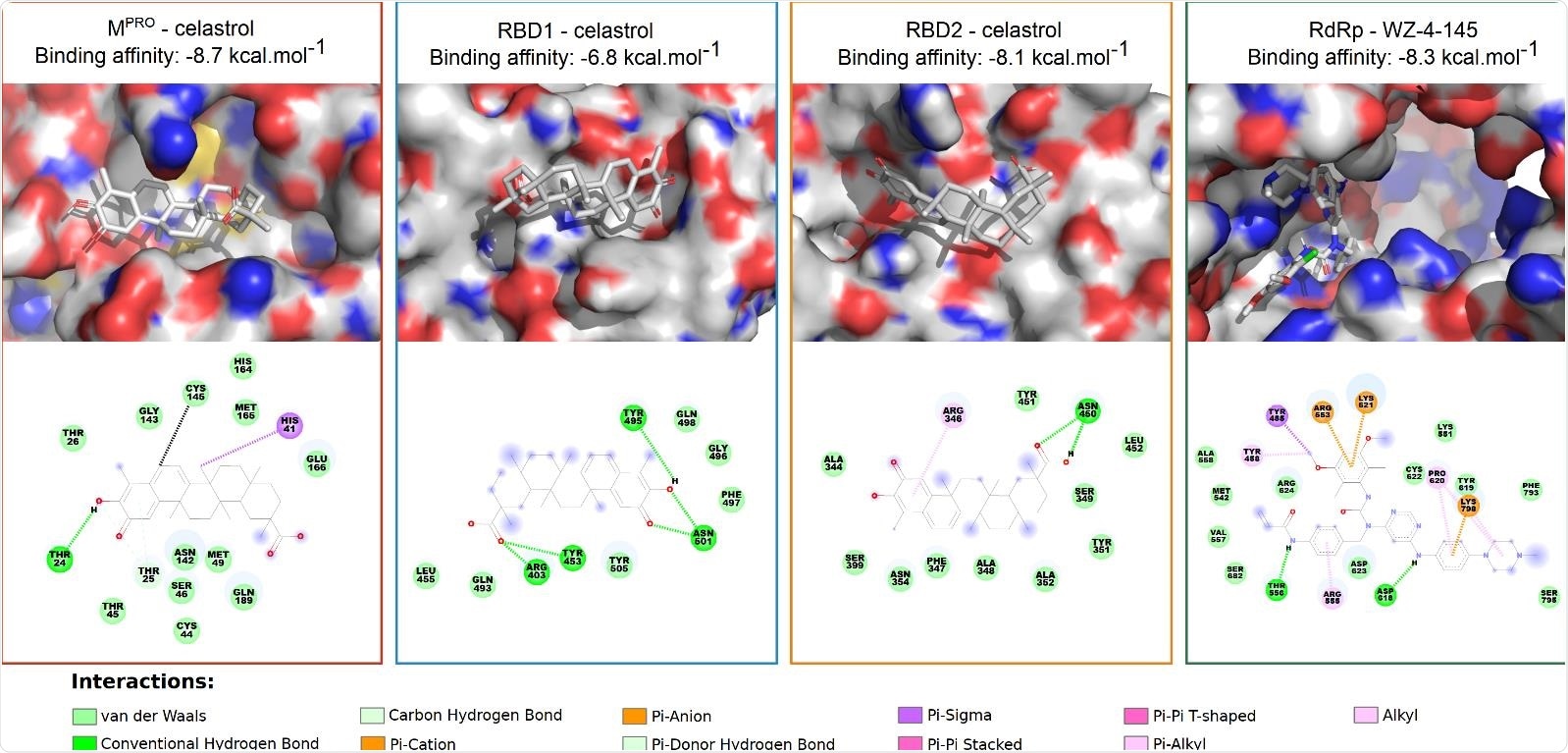
Detailed chemical interactions between the best-ranked drugs and inhibition sites of SARS-CoV-2 molecular targets. 2D target-drug interaction diagrams for the best structural configurations of viral molecular targets was determined using the Discovery Studio® software (version-2020). Celastrol had the most attractive ΔGbind median values of affinity energy to three targets (Mpro, RBD1, and RBD2) and WZ-4-145 to one (RdRp). The distance between the B-ring C6 and the sulfur atom of the Cys145 residue, which may be related to a possible Michael adduct formation for the best energy poses in each Mpro structure, ranged from 0.43 to 1.33 nm, with average value of 0.63 nm (black dashed line).
Scientists have revealed that SARS-CoV-2 infection induces IL-6 production. Such occurrences were spotted in both COVID-19 patients and in vitro SARS-CoV-2 infection models. Scientists have reported that celastrol could down-regulate the infection-induced IL-6 production, which causes a decrease in the SARS-CoV-2 load. Previous research has also indicated that this drug candidate has many functions. For example, it can down-regulate gene expression in PC-3 prostate carcinoma cells, suppress LPS-induced IL-6 production in RAW264, and express mRNA in influenza A-infected MDCK cells. In addition to these, scientists developed an experimental model of acute respiratory distress syndrome. They reported that celastrol could decrease lung injury by producing IL-6 and pro-inflammatory mediators into the pulmonary airways.
Although several studies have pointed out the effectiveness of celastrol as a therapeutic agent, it has some limitations. One of the limitations is its low solubility that results in poor bioavailability. The cytotoxic studies using the mouse model have shown that it is safe even at a higher concentration. However, more studies are required to assess its toxicity to humans. At present, celastrol is under clinical trials for the treatment of a variety of diseases, such as neurodegenerative disorders, different types of cancer, and inflammatory conditions (rheumatoid arthritis, psoriasis, and Crohn's disease).

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Fuzo, A.C. et al. (2021). Drug repurposing to face Covid-19: Celastrol, a potential leading drug capable of inhibiting SARS-CoV-2 replication and induced inflammation, bioRxiv 2021.04.20.439992; doi: https://doi.org/10.1101/2021.04.20.439992, https://www.biorxiv.org/content/10.1101/2021.04.20.439992v1
- Peer reviewed and published scientific report.
Fuzo, Carlos A., Ronaldo B. Martins, Thais F. C. Fraga‐Silva, Martin K. Amstalden, Thais Canassa De Leo, Juliano P. Souza, Thais M. Lima, et al. 2022. “Celastrol: A Lead Compound That Inhibits SARS‐CoV‐2 Replication, the Activity of Viral and Human Cysteine Proteases, and Virus‐Induced IL‐6 Secretion.” Drug Development Research 83 (7): 1623–40. https://doi.org/10.1002/ddr.21982. https://onlinelibrary.wiley.com/doi/10.1002/ddr.21982.