The abundance and distribution of microRNAs vary vastly across different sample types, indicating their unique regulatory needs and physiological functions. In human samples, most reports suggest that microRNA makes up approximately 0.01 to 2 % of total RNA, depending on cell tissue, type, and development stage.
This biological variability makes library preparation complicated, because uniformly applying the same PCR conditions across different samples can sometimes cause the over-amplification of libraries from miRNA-rich tissues and insufficient amplification of others. This poses difficulties for downstream normalization and risks an imbalance in representation in multiplexed sequencing runs.
The normalization of small RNA libraries has generally been achieved by manipulating library preparation steps, such as empirically varying cycle numbers for each sample type, modifying cleanup strategies, and adjusting adapter concentrations. These approaches can reduce variability slightly but often introduce new sources of bias, require more optimization, and are hard to scale up for processing large sample sets.
n6’s icon96™ system offers an alternative that monitors PCR amplification in real time to adaptively balance libraries at the single-well level. Rather than defining a fixed number of cycles in advance, the system stops amplification for each sample when a user-defined fluorescence threshold is reached. This equalizes library output across different input types.
This feature, known as AutoNorm™ technology (n6), is different to traditional normalization because it actively monitors DNA synthesis during each reaction rather than applying a fixed external adjustment. By setting a fluorescence threshold, the system brings all libraries to a comparable endpoint regardless of microRNA composition or starting material.
In practice, this means that samples containing copious ligation products stop amplifying earlier, which stops PCR duplicates from accumulating. Samples with reduced complexity or lower input, however, continue until they reach the same normalized endpoint.
Methods
The performance of the icon96 system (n6) was evaluated in combination with the NEXTFLEX™ Small RNA-Seq Kit v4. To ensure that the range of microRNA variability observed across human samples was covered, the experiment used total RNA from three samples: skeletal muscle, brain, and placenta (BioChain).
Brain tissue has one of the largest proportions of microRNA content of any human tissue. In brain samples, approximately 1-2 % of total RNA is microRNA, with miR-124 and miR-9 accounting for most of it. Skeletal muscle contains a lower overall proportion, usually around 0.1-0.5 % of total RNA, while the placenta contains intermediate levels, generally 0.5-1 % of the total RNA.
Libraries were prepared from the three sample types at two input amounts (1 ng and 10 ng) according to the manufacturer’s instructions. Amplification was performed using the icon96 system (n6) using either standard conditions or AutoNorm™ technology (n6), which set a fluorescence threshold of 6000 relative fluorescence units. Duplicates of all of the conditions were prepared.
The final libraries were quantified by employing a Qubit® fluorometer (Thermo Fisher Scientific). This was pooled equimolarly and loaded at 9 pM with 3 % PhiX spike-in on an Illumina® Miseq™ platform using 1×50 bp paired-end sequencing, aiming for 1M reads per sample. A custom Revvity script was used to perform small RNA analysis, and alignment was undertaken using mature miRNA sequences from miRBase v22.1.
Results
PCR cycles
10 ng libraries were amplified for 20 cycles under standard conditions, whereas 1 ng libraries required 24 cycles. When the same samples were processed using an icon96™ system (n6), the number of cycles needed to reach endpoint decreased to an average of 23.3 for 1 ng inputs and 18.7 for 10 ng inputs. This adjustment changed the cycle number, bringing it closer to the actual amplification requirements of each sample type and minimizing unnecessary extension in the miRNA-rich libraries (Figure 1).
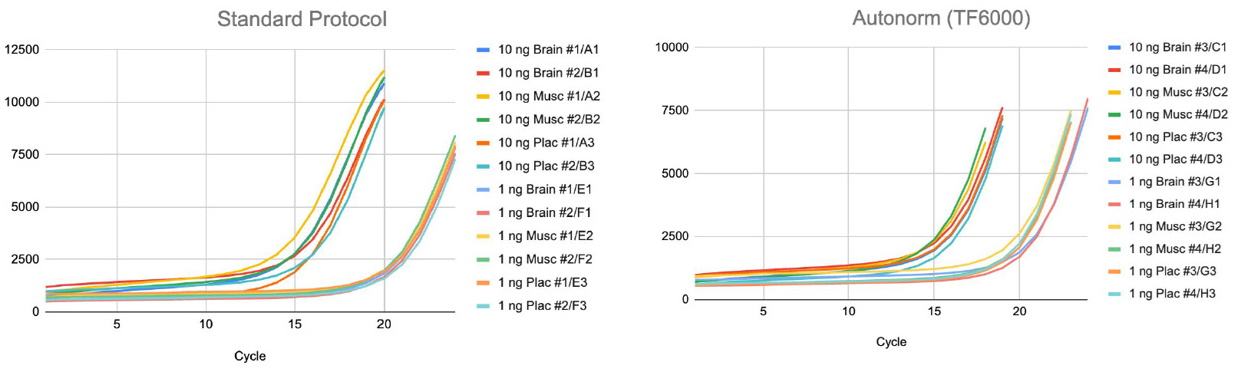
Figure 1. Amplification curves of small RNA libraries on the iconPCR™ system (n6) using standard protocol or with AutoNorm™ technology (n6). Image Credit: n6
Final library yield and average fragment size
Libraries generated using the standard method had an average concentration of 5.08 ng/μL and a CV = 48.5 % across different sample inputs and types. Contrastingly, the libraries amplified with AutoNorm™ technology showed a concentration of 2.53 ng/mL and less variability across different inputs and sample types, with a CV of 25.6 % (Figure 2).
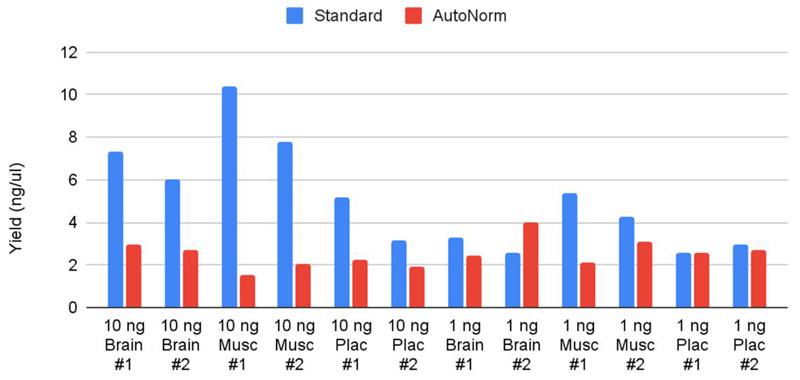
Figure 2. Final yield obtained for each of the conditions tested. Yield variation in libraries amplified with AutoNorm™ technology is lower than in those amplified with standard conditions. Image Credit: n6
The peak library sizes were comparable, measuring 159 ±4 bp for the standard group and 156±3 bp for the AutoNorm™ group.
Sequencing data analysis
Annotation of the mapped reads was performed according to RNA class. Average values were worked out for both groups (standard and AutoNorm™), and the RNA composition profiles demonstrated no substantial differences (Figure 3).
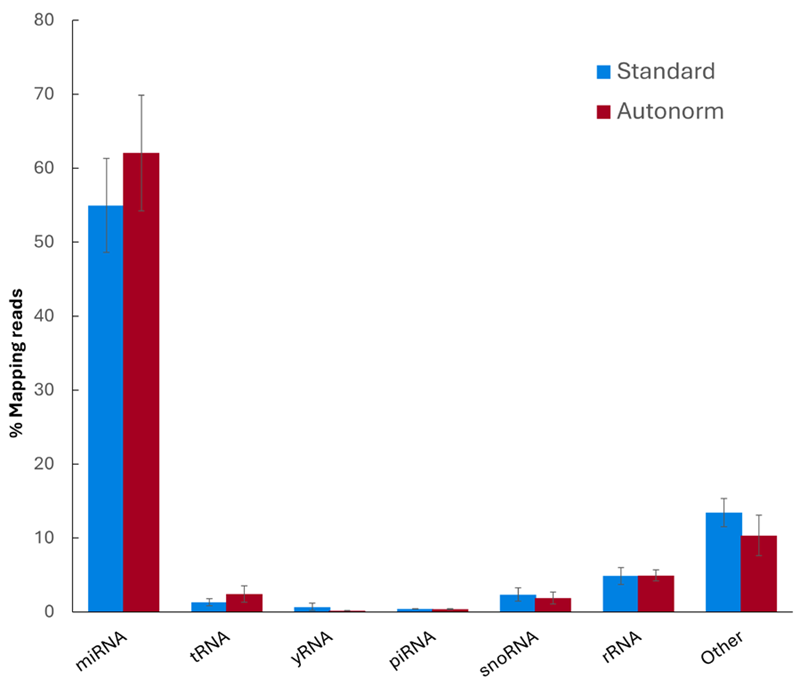
Figure 3. Average RNA type distribution for the two groups of samples (standard and AutoNorm™). Image Credit: n6
Conclusion
These results underscore the benefits of combining the icon96™ system (n6) with NEXTFLEX Small RNA-Seq v4 chemistry for small RNA studies, especially those that require different inputs and/or sample types.
Conversely, conventional normalization strategies that require empirical workflow adjustments, such as AutoNorm™ technology (n6), automate the process at the amplification step, reducing the number of PCR cycles required, improving reproducibility, and streamlining pooling for sequencing, without introducing detectable bias in library composition.
About n6
n6 proudly introduces icon96, a pioneering advancement in the genomics field with the world’s first real-time thermocycler with 96 individually controlled wells. This breakthrough technology promises to revolutionize DNA amplification and sequencing by offering unmatched simplicity and flexibility, setting a new standard for genomic research and diagnostics.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices, and treatments.