PCR amplification can seem straightforward, but in single-cell workflows even small miscalculations can quickly complicate results and downstream analysis.
Conventional PCR systems require users to set a predetermined number of cycles based on assay- and sample-specific factors. When performing single-cell sequencing experiments, this requires knowledge of the number of cells captured and the RNA content within them.
Different RNA quality and cell numbers necessitate scrupulous curation of PCR cycle numbers for each sample, which entails multiple thermocycler runs, increasing the potential for error and complicating the workflow.
n6 has developed icon96™, representing a crucial change in PCR technology. Its key innovation lies in the use of iconPCR™ technology with AutoNorm™, wherein each PCR reaction is individually monitored and terminated in real time based on fluorescence thresholds. This removes the guesswork inherent in traditional workflows and ensures that every library is optimally amplified.

Figure 1. icon96, the world’s first real-time thermocycler with 96 individually controlled wells. Image Credit: n6
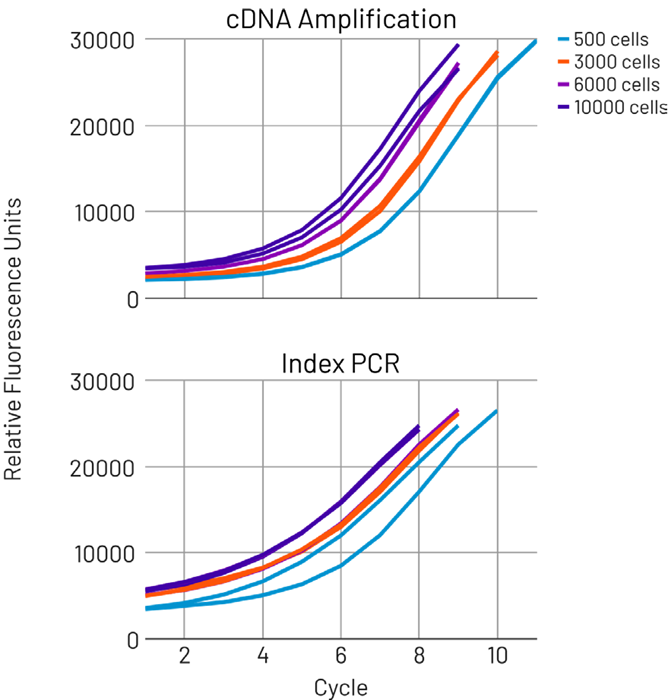
Figure 2. AutoNorm optimizes both cDNA amplification and index PCR reactions, ensuring data quality while eliminating the need to batch samples by cell capture number. Image Credit: n6
icon96 unlocks new single-cell workflows, enabling the simultaneous running of different sample types and input conditions, thereby preventing erroneous cycling conditions and increasing efficiency. To evaluate its performance, a series of 10x Genomics GEM-X Universal 3’ Gene Expression v4 libraries was prepared across a range of cell inputs.
The libraries were prepared using standard protocols or on the icon96 instrument with AutoNorm after cDNA synthesis. icon96 enabled a simplified workflow, while retaining the quality of the single-cell gene expression data.
Experimental design
Using two replicate GEM-X chips, a single culture of cells was captured. Two wells each of 500, 3000, 6000, and 10,000 cells were captured for each chip, for a total of 16 samples. Half of the samples followed the traditional workflow, while the other half used a modified workflow with icon96.
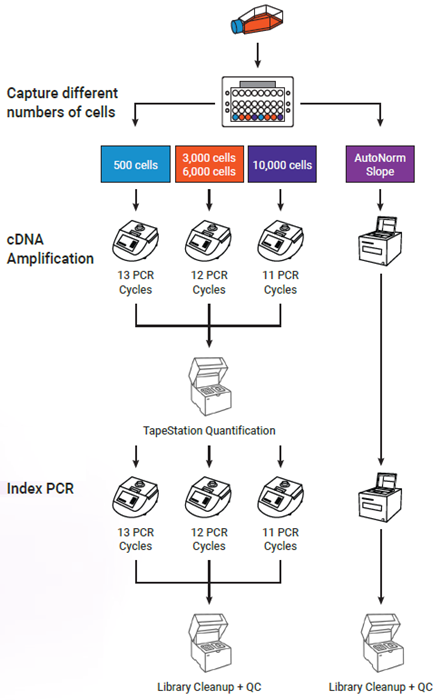
Figure 3. Experimental design showing critical workflow differences. Following cell capture and cDNA generation, samples following the standard workflow (left side) were run across three PCR reactions, in accordance with the standard PCR cycling conditions. Following cDNA amplification, each sample was quantified to determine the appropriate number of cycles for index PCR and was amplified accordingly. During iconPCR (right side), all 8 samples were run together on a single icon96 instrument using the slope method of AutoNorm. Following cDNA amplification, samples were again run together using the slope method of AutoNorm for index PCR. Following cleanup and QC, all libraries were pooled for sequencing. Image Credit: n6
Reduced workflow complexity
The conventional PCR methodology necessitated three separate thermocycler runs for both index PCR and cDNA amplification. On the other hand, all of the samples, regardless of input amount, were run simultaneously during both steps when the icon96 was used. Using AutoNorm, cycle numbers were dynamically determined, with each well stopping when the fluorescence reached the maximum slope of the linear amplification curve.
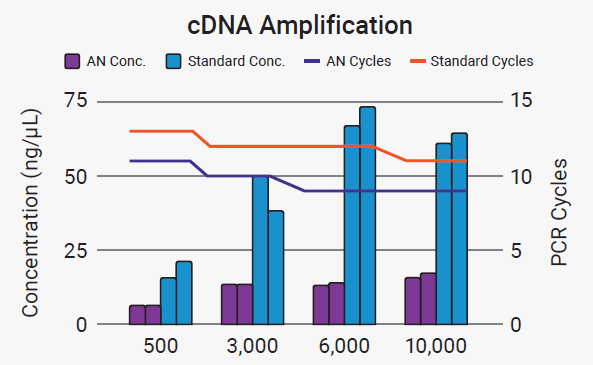
Figure 4. AutoNorm (AN) enables efficient processing of samples across a range of inputs. AutoNorm resulted in a 2-3-cycle decrease compared with standard PCR cycling recommendations (blue vs. orange lines). A wide range of concentrations was observed across the various inputs under standard PCR conditions (blue bars). In comparison, iconPCR with AutoNorm provided a more even final concentration (purple bars). It’s important to note that although the concentration post-cDNA amplification was determined for all samples, this is not a requirement to proceed with index PCR when using AutoNorm. Image Credit: n6
Table 1. AutoNorm simplifies index PCR. Due to differences in concentration after cDNA amplification, the standard protocol required 3 separate PCR runs. Conversely, iconPCR samples were able to be run on a single icon96 instrument. Use of AutoNorm reduced the number of cycles by 2-6 compared to the recommendations in the standard protocol. Source: n6
| Chip |
Input
(cells) |
Input for
Index PCR (ng) |
Index PCR
cycles |
Standard recom.
cycle number |
Cycle
Difference |
| Chip 1 |
500 |
65.5 |
10 |
12-14 |
2-4 |
| 3000 |
136.5 |
9 |
12-14 |
3-5 |
| 6000 |
135 |
9 |
12-14 |
3-5 |
| 10000 |
159 |
8 |
12-14 |
4-6 |
| Chip 2 |
500 |
66. |
9 |
12-14 |
3-5 |
| 3000 |
137.5 |
9 |
12-14 |
3-5 |
| 6000 |
141 |
8 |
12-14 |
4-6 |
| 10000 |
176.5 |
8 |
12-14 |
4-6 |
Table 2. Summary of PCR cycling. iconPCR with AutoNorm (AN) resulted in a 5-6-cycle decrease in PCR cycles compared to the standard PCR protocol. In total, the standard protocol required 6 separate thermocycler runs, while using iconPCR reduced that to 2, showcasing its ability to simplify complex workflows. Source: n6
| Chip |
Method |
Input |
cDNA Amp
Cycles |
Index PCR
Cycles |
Total
Cycles |
| Chip 1 |
AutoNorm |
500 |
11 |
10 |
21 |
| AutoNorm |
3000 |
10 |
9 |
19 |
| AutoNorm |
6000 |
9 |
9 |
18 |
| AutoNorm |
10000 |
9 |
8 |
17 |
| Standard |
500 |
13 |
13 |
26 |
| Standard |
3000 |
12 |
12 |
24 |
| Standard |
6000 |
12 |
11 |
23 |
| Standard |
10000 |
11 |
11 |
22 |
| Chip 2 |
AutoNorm |
500 |
11 |
9 |
20 |
| AutoNorm |
3000 |
10 |
9 |
19 |
| AutoNorm |
6000 |
9 |
8 |
17 |
| AutoNorm |
10000 |
9 |
8 |
17 |
| Standard |
500 |
13 |
13 |
26 |
| Standard |
3000 |
12 |
12 |
24 |
| Standard |
6000 |
12 |
11 |
23 |
| Standard |
10000 |
11 |
11 |
22 |
No loss of data quality
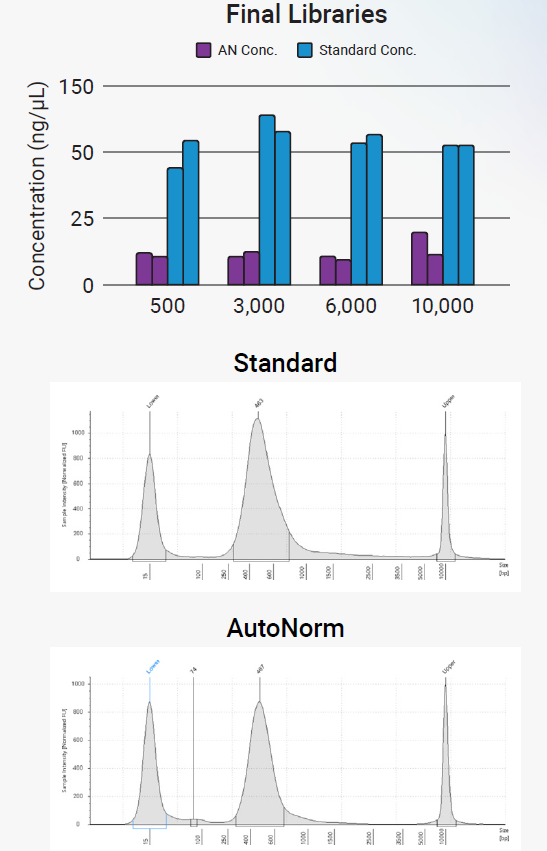
Figure 5. Final library quality control. AutoNormalized libraries provided sufficient quantities of final library for sequencing (top) and maintained the same quality as determined by Tapestation (bottom). Image Credit: n6
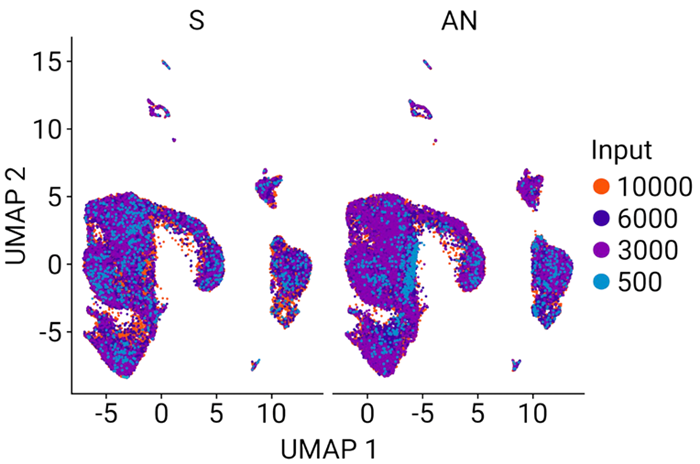
Figure 6. Clustering of all samples reveals common gene signatures. Clustering of all 16 samples revealed no differences between the standard PCR method (S) and AutoNorm (AN), showcasing that the change in workflow does not negatively affect data quality. Image Credit: n6
Advantages of iconPCR
Conventional PCR
- Fixed cycle count (e.g., 16 cycles)
- Under/over-amplification is common
- One-size-fits-all amplification
- Manual normalization and quant needed
- Varied library quality
- Extra costs for reruns and QC
- More hands-on time
- Greater level of reagent waste
iconPCR (with AutoNorm)
- Fluorescence monitoring in real time
- Per-well cycle control is based on signal, rather than guesswork
- Uniform, balanced libraries across every well
- True-to-biology amplification leads to less overcycling and reduced dropouts
- Fully automated normalization
- Fewer failed libraries and less reagent waste
- Reduction in hands-on time of 40-60 %
- Fewer rescue/ QC steps lead to faster turnaround
Conclusion
icon96 has redefined single-cell library preparation by removing the limitations of conventional PCR workflows. Thanks to its innovative per-well AutoNorm mechanism, icon96 provides optimized PCR cycling regardless of cell input. Samples of differing cell qualities or numbers no longer need to be split across thermocyclers, and experiments are no longer at risk when the number of captured cells is different from the expected number. Employing icon96 reduces workflow complexity, enabling higher sample throughput with a lower risk of errors, without sacrificing data quality.
About n6
n6 proudly introduces icon96, a pioneering advancement in the genomics field with the world’s first real-time thermocycler with 96 individually controlled wells. This breakthrough technology promises to revolutionize DNA amplification and sequencing by offering unmatched simplicity and flexibility, setting a new standard for genomic research and diagnostics.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices, and treatments.