The quality of a piece of fruit is hidden beneath its skin. Who knows what lies behind each perfectly shining apple?
In the quest for RNA biomarkers indicative of fruit maturity, a cost-effective, reliable workflow is crucial. Accurate harvest timing is critical for shelf life and apple quality, but at present, growers rely on their eye and a simple starch test. As a result, roughly 30% of harvested fruit is discarded in storage due to quality issues.
Molecular biomarkers could help overcome these limitations, but identifying them would require the sequencing of hundreds of apple samples. RNA extraction has historically been difficult in plants, and, in a farming context, costs must be low.
I’m at a genomics institute with genome sequencers in my own lab, and still a major bottleneck for every apple biomarker project is data generation.”
Dr. Alex Harkess, HudsonAlpha Institute for Biotechnology
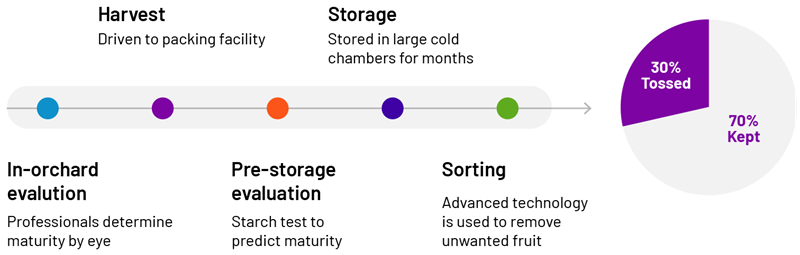
Image Credit: n6
RNA normalization: The hidden labor drain
Before the HudsonAlpha team can search for biomarkers by sequencing the fruit-extracted RNA samples, they must undertake several processing steps to ensure they get the most out of each sequencing run. Each step takes time and money and increases the risk of errors that may affect data quality.
- Isolate RNA from hundreds of apple samples
- Quantify each sample (concentration can vary five-fold or more)
- Manually normalize them to equal concentrations
- Prepare sequencing libraries with consistent amplification
- Normalize library pools before sequencing
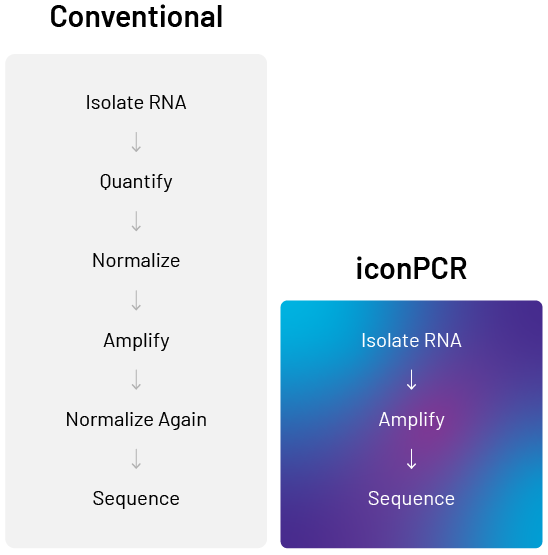
Image Credit: n6
We have RNA inputs ranging from 0.3 to 1.5 micrograms – a fivefold difference. Normally, we’d want these identical before sequencing. This is labor-intensive and often insufficient.”
Dr. Alex Harkess, HudsonAlpha Institute for Biotechnology
Introducing intelligent amplification with icon96™ and AutoNorm™
icon96 with AutoNorm technology has a huge potential impact. The system monitors each well in real time, rather than setting a fixed PCR cycle count in advance. This means that all libraries reach identical amplification targets, regardless of the starting RNA quantity.
How icon96 works
- Real-time monitoring tracks amplification curves for each of the samples
- User-defined fluorescence threshold catalyzes the stop-cycle automatically
- Even if the inputs are chaotic, the output is perfect
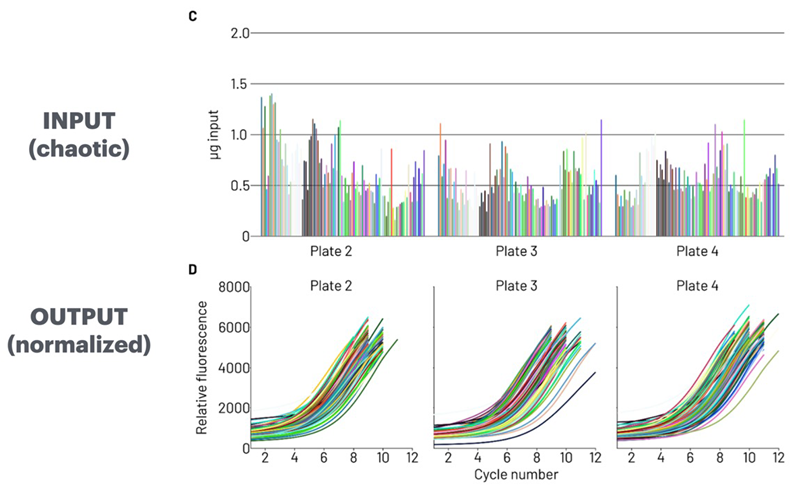
Figure 1. Plant-derived RNA samples ranging from 0.5-1.5 micrograms (INPUT) were amplified on icon96. The stop cycle for each sample was dynamically selected using AutoNorm based on the slope of the amplification curves (OUTPUT). Image Credit: n6
The outcome: A practical solution for RNA-based biomarker testing
In a pilot test, the HudsonAlpha team sequenced 384 apple peel samples after library preparation using icon96. They identified 14 biomarkers for fruit maturation that showed strong correlation with internal ethylene concentration (IEC), the industry standard for assessing maturity.
11 of the 14 biomarkers consistently predicted harvest timing, demonstrating their utility in real-world conditions. They also calculated that their sequencing costs were approximately 30 % lower, demonstrating that icon96 is both impactful and practical.
Key metrics
- 384 apple peel samples processed
- Five-fold variation in input RNA controlled
- Roughly 30% reduction in RNA-seq costs
- Strong correlation between fruit maturity metrics and RNA-seq biomarkers
- 11 of 14 biomarkers consistently predict harvest timing
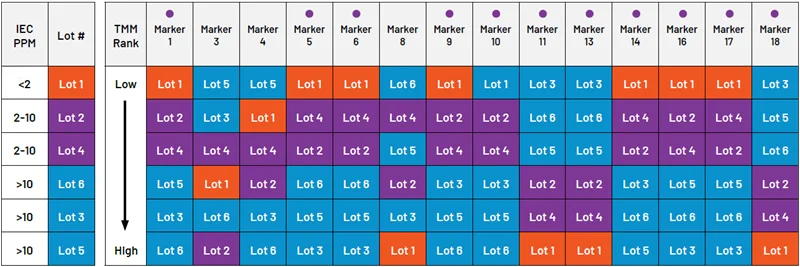
Figure 2. Ranking six lots by relative expression of maturity biomarkers measured by RNA-seq. 11 of 14 biomarkers ( ) consistently correlate with ethylene concentration. Image Credit: n6
) consistently correlate with ethylene concentration. Image Credit: n6
From laboratory bench to grower’s toolkit
This article established a blueprint for scalable biomarker discovery in agriculture. The USDA and HudsonAlpha team collaborated to demonstrate that combining icon96 with automated normalization enables the use of transcriptional biomarkers in the field to help growers make data-driven decisions about their harvests, thereby improving fruit quality across the supply chain and reducing waste.
Key takeaways
- Reproducible NGS data generation can be scaled up
- Molecular biomarkers can predict fruit maturity reliably
- Workflow efficiency directly affects costs and research timelines
- The approach extends beyond apples: any variable RNA input can benefit from this method
Icon96+ AutoNorm: Transforming RNA workflows
This apple biomarker project showcases icon96’s power in RNA-seq workflows beyond agriculture. Whether preparing libraries from single-cell suspensions, cell-free DNA (cfDNA), FFPE tissues, or difficult plant samples, AutoNorm eliminates the manual guesswork that cycle optimization can require, resulting in faster decision making, scalable workflows, and cleaner data.
The ultimate result to you is that we’ve achieved about a 30 % reduction in the cost of generating this RNAseq data for apple and pear biomarker projects.”
Dr. Alex Harkess, HudsonAlpha Institute for Biotechnology
About n6
n6 proudly introduces icon96, a pioneering advancement in the genomics field with the world’s first real-time thermocycler with 96 individually controlled wells. This breakthrough technology promises to revolutionize DNA amplification and sequencing by offering unmatched simplicity and flexibility, setting a new standard for genomic research and diagnostics.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices, and treatments.