PCR cycle selection is a delicate balancing act. When preparing RNA-seq libraries, choosing too many cycles risks overcycling, which can compromise data quality and distort transcript representation. Choosing too few cycles, on the other hand, can yield insufficient material for sequencing.
This difficulty is particularly acute when working with heterogeneous sample sets, in which RNA quantity, integrity, and transcript complexity can vary widely, as in FFPE-derived material. Legacy fixed-cycle PCR forces researchers to commit to a single cycle number, often leading to excessive handling steps, imbalanced amplification, and suboptimal results across the batch.
icon96™ addresses these difficulties with iconPCR and AutoNorm™ by constantly monitoring real-time amplification and stopping each reaction at its optimal endpoint. Because it adapts to the true amplification kinetics of each RNA library, AutoNorm reduces per-sample optimization efforts, eliminates the need for input-matched batching, and prevents the loss of quality associated with overcycling.

Figure 1. icon96™, the world’s first real-time thermocycler with 96 individually controlled wells. Image Credit: n6
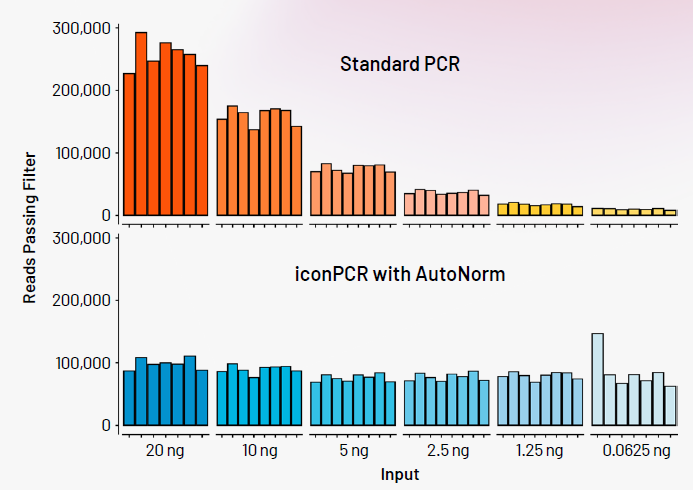
Figure 2. Here we show the significant yield variance when using a single PCR instrument with a fixed number of PCR cycles compared to iconPCR with AutoNorm where each sample is amplified to similar levels. Image Credit: n6
Experimental design
For all of the library preparations in this study, RNA was processed with the E7760L NEBNext® Ultra™ II Directional RNA Library Prep Kit for Illumina®, the E7400X NEBNext® rRNA Depletion Kit v2 (Human/ Mouse/ Rat), and indexed using the E6442S NEBNext® Multiplex Oligos for Illumina®.
The libraries were created using either standard PCR or iconPCR with AutoNorm from 10, 50, and 250 ng of RNA extracted from FFPE (colon, liver) and control (UHR and ERCC) samples. Two cycling conditions were applied to each sample amplified by standard PCR: the lower cycle number recommended by NEB and a higher cycle number deliberately used to induce overcycling.
iconPCR samples were intensified using AutoNorm, which dynamically adjusts cycle numbers in real time based on fluorescence signals, preventing overamplification and optimizing yield across a range of sample qualities and inputs.
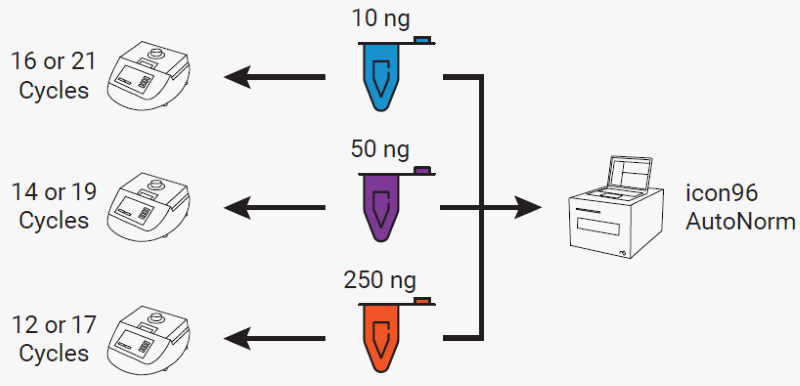
Figure 3. Experimental design illustrating sample distribution across workflows. On the left, samples were split by input amount and subjected to either the recommended number of cycles or the recommended cycles plus five. On the right, all samples were processed together in a single run using AutoNorm on icon96. Image Credit: n6
AutoNorm prevents overcycling
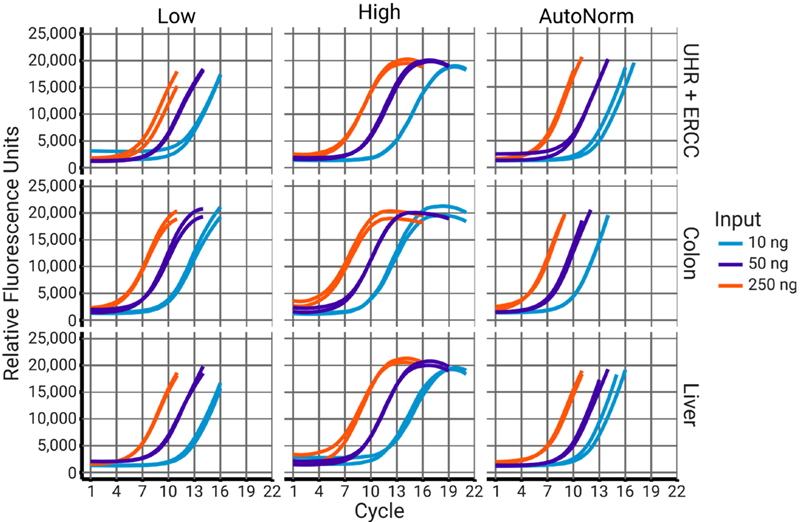
Figure 4. iconPCR enables optimized workflows with built-in safeguards against overamplification. Libraries amplified under standard low-cycle conditions were terminated near or before the start of the plateau phase. Samples with higher cycle numbers were overamplified, as expected. In contrast, iconPCR with AutoNorm accurately terminated all reactions during the linear amplification phase, preventing overcycling and preserving library quality. Image Credit: n6
Table 1. AutoNorm dynamically and accurately controls cycle numbers across RNA samples. This table shows the average stop cycle for each sample in the study, illustrating how AutoNormalization™ adjusts amplification cycles in real time to match the specific input and quality of each RNA sample. By tailoring cycle numbers individually, AutoNormalization prevents under- or overamplification, ensuring consistent library quality within a single PCR run. Source: n6
| |
Standard Low |
Standard High |
AutoNorm |
| Input |
All Samples |
All Samples |
FFPE Colon |
FFPE Liver |
UHR + ERCC |
| 10 ng |
16 |
21 |
14 |
16 |
17 |
| 50 ng |
14 |
19 |
12 |
13 |
14 |
| 250 ng |
11 |
16 |
9 |
11 |
10 |
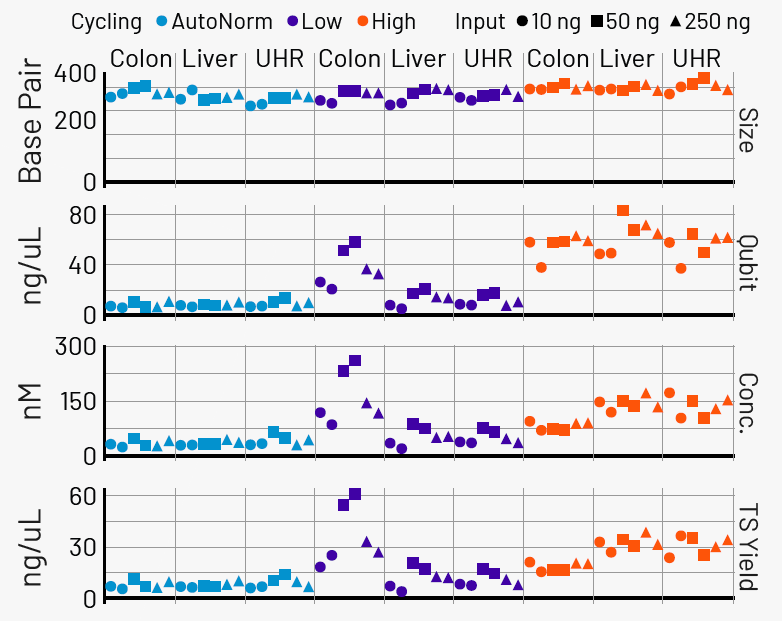
Figure 5. iconPCR enables effective normalization across input amounts. After library preparation, samples were individually SPRI-purified and quantified. Libraries amplified with iconPCR showed consistent quantities across all samples and input levels, with no detectable changes in library size distribution. Image Credit: n6
Data quality
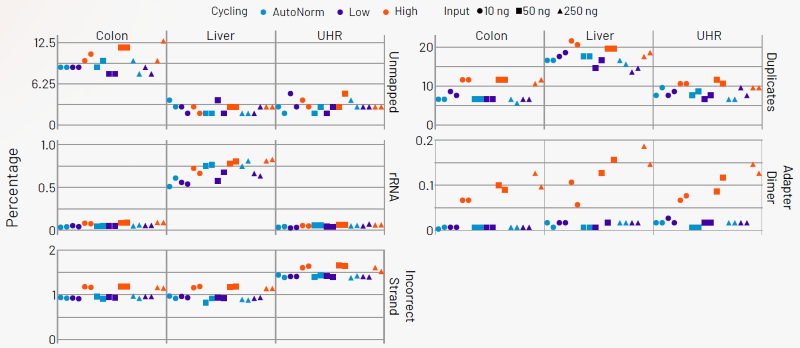
Figure 6. AutoNorm ensures optimal data quality. Samples amplified using the lower fixed cycle numbers and AutoNorm displayed consistent, optimal quality metrics. In contrast, overcycled samples showed increased dimer formation, elevated PCR duplicate rates, and more reads assigned to the incorrect strand. Additionally, colon samples exhibited a notable increase in unmapped reads. Image Credit: n6
Workflow advantages of iconPCR
Conventional PCR
- Fixed cycle count (e.g., 16 cycles)
- Extra costs and time needed for repeat runs and QC
- Variable library quality leads to inconsistent data
- Upfront normalization and quantification are necessary
- Under/over-amplification is common
- One-size-fits-all amplification
- More hands-on time for technicians
- More reagent waste because of failed or overamplified libraries
iconPCR (with AutoNorm)
- Fluorescence monitoring in real time
- Optimal amplification for each sample
- Uniform library quality across all wells
- Less reagent waste and fewer failed libraries
- Fully automated normalization
- Per-well cycle control is based on signal, rather than guesswork
- Less hands-on time leads to a 40-60 % reduction in technician labor
- Faster turnaround with fewer rescue/ QC steps
Conclusion
iconPCR eliminates the limitations of fixed-cycle PCR, bringing a new level of control to RNA-seq library preparation. The per-well AutoNorm system adapts amplification to each sample’s needs, preventing overcycling that can compromise data quality. This approach enables researchers to process RNA samples with vastly different quantities and integrity in a single run, improving consistency and simplifying workflows.
By reducing batch variability and maintaining optimal library quality, iconPCR supports more reproducible and accurate RNA-seq results, which enhance the reliability of both clinical and research-led transcriptomic studies.
By integrating the robust chemistry and depletion technology of NEBNext kits with the real-time, automated amplification control of iconPCR and its per-well AutoNorm capability, this workflow provides reproducible, reliable RNA-seq libraries across varying sample qualities and input amounts.
iconPCR counteracts the challenges of fixed-cycle PCR by adapting amplification to each sample’s needs, preventing overcycling that can impair data quality. This combined approach minimizes batch variability and the risk of overamplification, reduces hands-on time, and increases efficiency, all of which enable researchers to process RNA samples with vastly different integrity and quantity in a single run.
Correspondingly, the generated libraries are consistently high-quality, supporting more reproducible and accurate transcriptomic studies for clinical and research applications alike.
About n6
n6 proudly introduces icon96, a pioneering advancement in the genomics field with the world’s first real-time thermocycler with 96 individually controlled wells. This breakthrough technology promises to revolutionize DNA amplification and sequencing by offering unmatched simplicity and flexibility, setting a new standard for genomic research and diagnostics.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices, and treatments.