Jun 28 2017
Bioorthogonal labeling of human prostate cancer tissue slice cultures to identify marker glycoproteins
Glycosylated proteins are often overexpressed in tumor cells and thus could serve as tumor markers, especially those with the interesting molecule sialic acid as their sugar moiety. In the journal Angewandte Chemie, American scientists now report on a bioorthogonal labeling test for sialylated glycoproteins based on a glycoproteomics approach. This assay not only assesses the level of sialylated glycans in the tumor cell membranes, but also identifies up- or downregulated proteins directly in the prostate cancer tissue.
Tumor cells are characterized by an accelerated metabolism with some proteins up- and some downregulated compared to cells in normal healthy tissue. As the quantity and quality of proteins in the cells can be assessed by a proteomics approach, scientists seek to use a proteomics test system to identify and explore the proteins typical for cancer metabolism. Carolyn R. Bertozzi and her research team from Stanford University have chosen a bioorthogonal labeling strategy to identify sialylated glycoproteins, which are especially interesting because the sialic acid sugar moiety helps cells to evade the immune system. Their labeling approach further applies the cancer tissue, not cell cultures, thus it provides direct assess to the tumor metabolism in its natural environment.
In bioorthogonal labeling, a label, usually a fluorescent molecule, is chemically attached to target molecules, which can then be identified by bioimaging or mass spectrometry. One of the key aspects is that there is as little interference with the normal cell metabolism as possible. "Accurate models of human biology are particularly important for research at the intersection of glycoscience and human health," the authors argue. Therefore, they chose tissue slice cultures as a form of live human tumor tissue, because "prostate tissue slice cultures ... allow direct comparisons of cancerous and normal tissue from the same patient source."
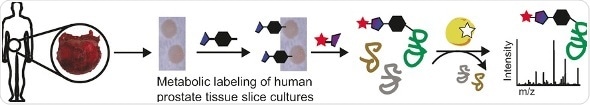
In their approach, the tissue slice cultures were treated with an azide-modified sialic acid, which was readily integrated into the tumor cell metabolism. Then, a fluorescent label was chemically attached to the azide group. After the labeling, the scientists inspected the tissue slices either directly by imaging or by mass spectrometry after cell lysis. They observed clear differences between the cancer tissue and the healthy one and found characteristic proteins up- or downregulated in the cancer tissue. Merging this platform with existing glycoproteome analysis techniques are future options, the authors propose, setting the stage for addressing further questions related to the roles of sialic acid, glycoproteins, and cancer.