Researchers from Qatar have demonstrated that antibody repertoires against endemic human coronaviruses are qualitatively different in children when compared to the general adult population, as well as healthy adult blood bank donors – and they may also provide cross-protection against epidemic/pandemic strains. Their paper is currently available on the bioRxiv* preprint server.
These include severe acute respiratory syndrome coronavirus (SARS-CoV), Middle East respiratory syndrome coronavirus (MERS-CoV), and now severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) – the etiological agent of the ongoing coronavirus disease 2019 (COVID-19) pandemic.
MERS Coronavirus Particles Colorized scanning electron micrograph of Marburg virus particles (blue) both budding and attached to the surface of infected VERO E6 cells (yellow). Image captured and color-enhanced at the NIAID Integrated Research Facility in Fort Detrick, Maryland. Credit: NIAID
Thus far, several studies have suggested that endemic human coronaviruses can induce broadly cross-reactive T cell responses, affecting clinical outcomes of acute infections with the phylogenetically related epidemic viruses (namely MERS-CoV and SARS-CoV-2).
Nevertheless, a systematic appraisal to elucidate immunodominant B cell antigen determinants of endemic human coronaviruses, their degree of representing cross-reactive antigenic sites, as well as the variation in the cellular and humoral immune responses among humans of different age has not yet been pursued.
Recently, the researchers from Sidra Medicine, Qatar University, Weill Cornell Medical College in Qatar and Hamad Bin Khalifa University in Doha, Qatar, conducted a comprehensive retrospective analysis of coronavirus-specific antibodies in a large number of samples from children and adults.
Collecting antiviral antibody repertoires
To obtain a more in-depth insight into human antibody responses to endemic human coronaviruses, the researchers performed Phage-Immunoprecipitation Sequencing (PhIP-Seq) on serum or plasma samples collected before COVID-19 pandemic from a large number of individuals in three different cohorts.
More specifically, these cohorts included healthy male adult blood donors with diverse ethnic backgrounds, adult male and female participants of a national cohort study known as the Qatar Biobank, as well as pediatric inpatients and outpatients. They were tested for metabolic conditions unrelated to infection, malignancies, or chronic diseases.
In short, with the use of PhIP-Seq, comprehensive antiviral antibody repertoires were gathered across individuals in these three human cohorts by phage display of oligonucleotide-encoded peptidomes, which was followed by immunoprecipitation and massive parallel sequencing.
The seroprevalence of endemic coronaviruses were assessed in the three cohorts separately by imputing species score values via counting significantly enriched peptides for a given coronavirus species that share less than seven amino acids linear sequence identity.
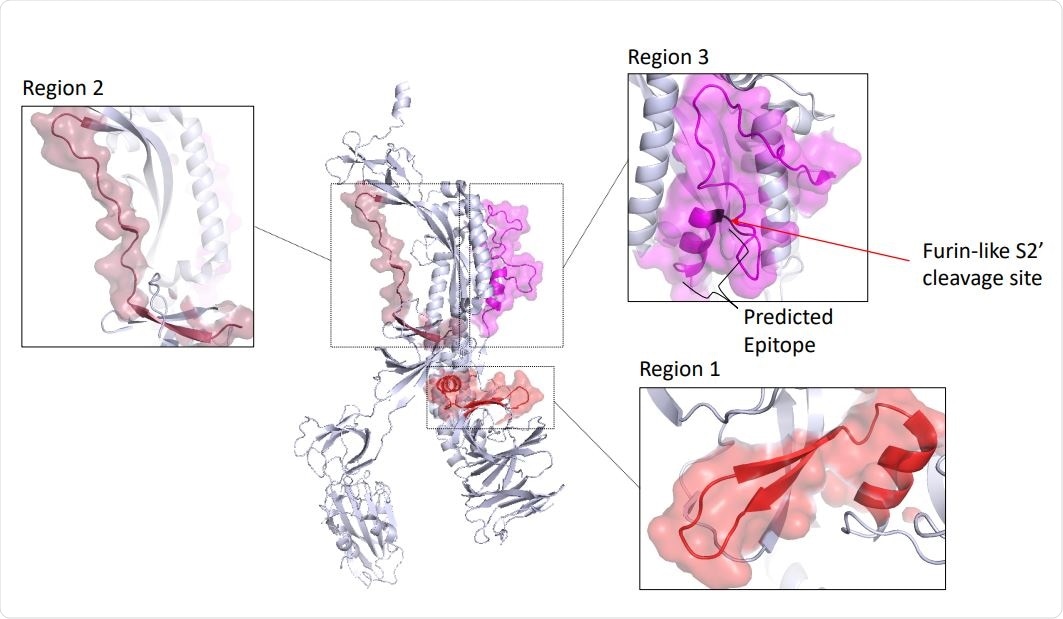
Antigenic regions and predicted antibody binding sites of the spike (S) protein. A monomer of the S 684 protein of SARS-CoV-2 in the prefusion conformation (PDB id: 6VXX, chain A), with the 685 regions 1, 2 and 3 shown enlarged. FP, fusion peptide; HR1 and HR2, heptad repeat 1 and 2.
Differential targeting of structural and non-structural SARS-CoV-2 proteins
This comprehensive screen for antiviral antibody repertoires across three distinct human cohorts revealed a large number of peptides that exhibit novel linear epitopes in several proteins of endemic human coronaviruses. Among them, there were 25 new immunodominant linear B cell epitopes.
Furthermore, by using stringent and robust cut-off values, the researchers have also estimated that the seroprevalence of endemic human coronaviruses ranges from approximately 4% to 27%, depending on the species and studied cohort.
A surprising and somewhat unexpected finding of this study was that circulating IgG antibodies in pediatric subjects (when compared to the adult ones) differentially targeted structural and non-structural proteins of human coronaviruses.
For example, antibodies in children showed affinity towards structural proteins such as nucleocapsid protein and spike glycoprotein. At the same time, in adults, specific regions of the non-structural polyprotein known as 'pp1ab' were their primary target.
Furthermore, some antibodies were found to be broadly cross-reactive with peptides of epidemic human and non-human coronavirus isolates, which may play a protective role against endemic and epidemic infections – particularly among children that seem to target such functionally relevant B cell epitopes and seldom present with severe disease outcomes.
To serve and cross-protect
"Information about the targets of immune responses to coronaviruses across different species provides a valuable resource for the prediction of candidate targets of newly emerging coronaviruses" study authors emphasize the importance of their research.
These findings elucidate the humoral immune responses to natural infection with endemic human coronaviruses and may have important implications for understanding highly variable clinical outcomes seen in human infections. But there is also an issue of cross-reactivity.
"It is tempting to speculate that natural infection with endemic human coronavirus may provide some degree of cross-protection and may, therefore, affect health outcomes in individuals infected by epidemic coronaviruses, such as SARS-CoV-2 or MERS-CoV", state study authors.
Consequently, the results of this study may indeed inform further development of prophylactic or therapeutic monoclonal antibodies and vaccine design, with potential broad-spectrum activity against many different coronaviruses.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
Khan, T. et al. (2020). Endemic human coronaviruses induce distinct antibody repertoires in adults and children. bioRxiv. https://doi.org/10.1101/2020.06.21.163394.
- Peer reviewed and published scientific report.
Khan, Taushif, Mahbuba Rahman, Fatima Al Ali, Susie S.Y. Huang, Manar Ata, Qian Zhang, Paul Bastard, et al. 2021. “Distinct Antibody Repertoires against Endemic Human Coronaviruses in Children and Adults.” JCI Insight, January. https://doi.org/10.1172/jci.insight.144499. https://insight.jci.org/articles/view/144499.