Coronaviruses are a group of enveloped viruses having a single-stranded positive-sense RNA genome. They belong to 4 genetic groups – alpha, beta, delta, and gamma coronaviruses. They infect a wide range of hosts, including bats, humans, rodents, camels, and pigs.
Currently, there are seven known coronavirus species that can infect humans, of which four coronaviruses - OC43 (HCoV-OC43), 229E, NL63, and HKU1 – cause flu-like symptoms. Three highly pathogenic and transmissible viruses, severe acute respiratory syndrome coronavirus 1 (SARS-CoV-1) and Middle East respiratory syndrome coronavirus (MERS-CoV) and the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), can cause fatal illness involving respiratory disorders and pneumonia.
The coronavirus disease (COVID-19) that emerged in late December 2019 in Wuhan, China, was caused by the novel coronavirus SARS-CoV-2, which is very similar to SARS-CoV, binding to the same host cell receptor, angiotensin-converting enzyme 2 (ACE-2). However, the novel virus is genetically distinct from SARS-CoV-1. Both SARS-CoV-1 and SARS-CoV-2 are thought to be transmitted across species barriers from bats to humans by way of an intermediary host such as civet cats or pangolins.
COVID-19 disease causes influenza-like symptoms ranging from mild to severe lung dysfunction and multi-organ failure, which can be fatal in patients with comorbidities. The SARS-CoV-2 pandemic has spread across the globe, causing unparalleled public health and economic crises, which presents an urgent need for antiviral countermeasures to fight the infection.
Repurposing existing therapeutic molecules to test for possible effects against SARS-CoV-2
Several antivirals including nucleoside analogs such as remdesivir, favipiravir, and ribavirin, protease inhibitors such as disulfiram, lopinavir, and ritonavir, antiparasitic drugs such as chloroquine and hydroxychloroquine, monoclonal antibodies, peptides, and small-molecule drugs are being repurposed in clinical trials across the world and tested for their possible therapeutic effects against SARS-CoV-2. Remdesivir, in particular, inhibits viral RNA polymerases and is said to be a potent antiviral against SARS-CoV-2.
A single-chain variable antibody fragment with possible antiviral effects against several viruses
3D8, a 27-kDa recombinant antibody fragment, was found in autoimmune-prone Murphy Roths Large mice. The single-chain variable fragment (scFv) has the nucleic acid hydrolyzing activity and is shown to degrade DNA or mRNA of viruses in infected cells.
“scFv has various advantages over traditional monoclonal antibodies such as ease of genetic manipulation, rapid molecular design and characterization, greatly reduced size, production of antibodies against viral proteins, and various expression systems.”
Studies have shown that this protein shows broad-spectrum antiviral effects against influenza virus, herpes simplex virus, and pseudorabies virus in vitro due to its nucleic acid-hydrolyzing property. 3D8 also exhibited in vivo antiviral therapeutic effects against some viruses in C57BL/6 mice and suppressed the transmission of avian influenza and bronchitis viruses in transgenic chickens. However, 3D8 scFv’s antiviral activity against SARS-CoV-2 and other coronaviruses is not well understood.
“The reduction of infectious virus particles accounted for the nuclease activity of 3D8, indicating the degradation of viral nucleic acids prohibited the production of viral genomes and proteins.”
Investigating the antiviral activity of 3D8 scFv against coronaviruses in vitro
Recently, a team of researchers from the Novelgen Co., Ltd., Hallym University, Korea University College of Medicine, Sungkyunkwan University, and Chung-Ang University, Republic of Korea, investigated the antiviral activity of 3D8 scFv against coronaviruses in vitro in their study published on the bioRxiv* preprint server. In this study, the researchers report that 3D8 scFv blocked the replication of human coronavirus OC43, SARS-CoV-2, and porcine epidemic diarrhea virus.
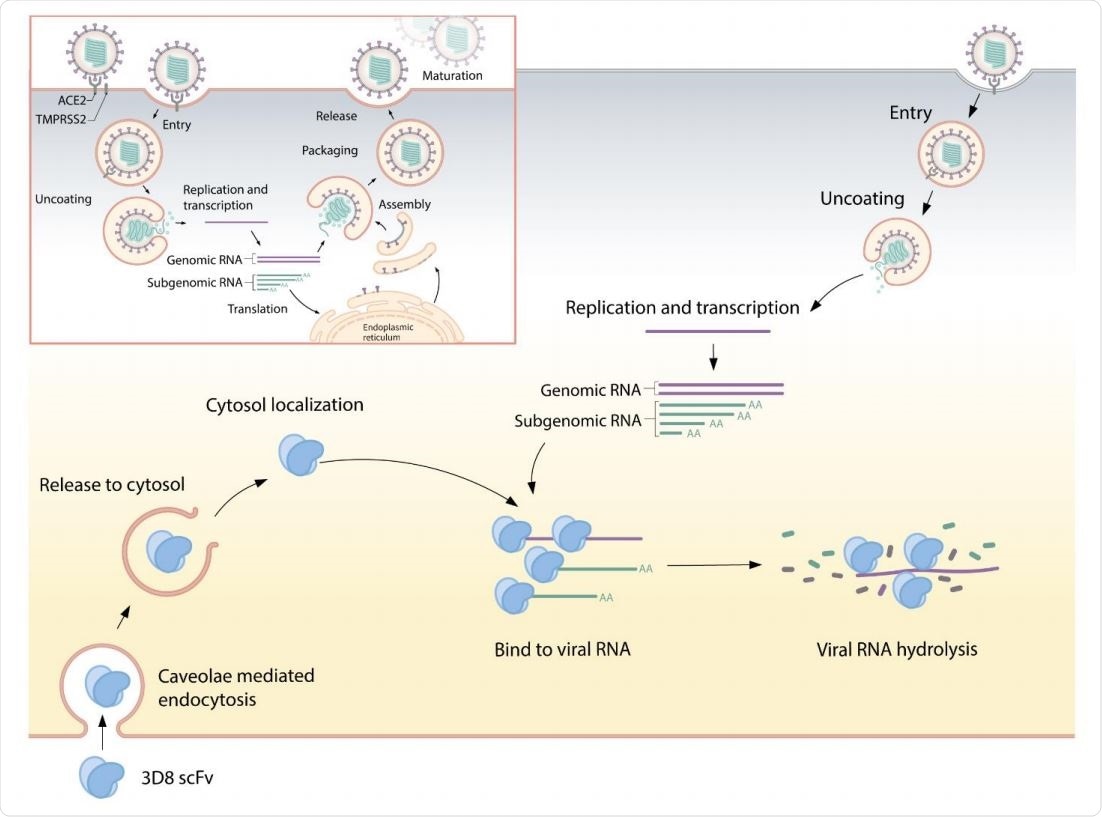
Suggested mode of action for 3D8 - 3D8, a single chain variable fragment (scFv), is internalized into the cell through caveolae-mediated endocytosis. After release from the endosomal compartment, 3D8 localizes to the cytosol. In the cytosol, it binds to the viral nucleic acid and degrades it to prevent its amplification, thus inhibiting viral growth. 3D8 confers nuclease activity without sequence specificity and hydrolyzes viral RNA genomes or transcripts.
3D8 may be a potential candidate for anti-SARS-CoV-2 therapeutics

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
The results of the study demonstrated 3D8 scFv’s prophylactic and therapeutic effects against SARS-CoV-2 in Vero E6 cells. The researchers performed immunoblot and plaque assays that showed the absence of coronavirus nucleoproteins and infectious viral particles, respectively, in 3D8 scFv-treated cells. Moreover, they observed 3D8’s strong antiviral effects against HCoV-OC43 and porcine epidemic diarrhea virus.
“Taken together, the 3D8, a nucleic-acid hydrolyzing mini-antibody, might be a potential candidate for antivirals due to the broad spectrum, the easy penetration to the cell, and the accessibility to the lung
in vivo”
Based on these findings, the authors concluded that their study offers crucial insights into the broad-spectrum antiviral agent, 3D8 scFv and that 3D8 can be considered a potential antiviral therapeutic agent against SARS-CoV-2 and multiple other coronaviruses.

 This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
This news article was a review of a preliminary scientific report that had not undergone peer-review at the time of publication. Since its initial publication, the scientific report has now been peer reviewed and accepted for publication in a Scientific Journal. Links to the preliminary and peer-reviewed reports are available in the Sources section at the bottom of this article. View Sources
Journal references:
- Preliminary scientific report.
3D8, a nucleic acid-hydrolyzing scFv, confers antiviral activity against SARS-CoV-2 and multiple coronaviruses in vitro Gunsup Lee, Shailesh Budhathoki, Hyeok Soon Choi, Kwang-ji Oh, Geum-Young Lee, Yeon Kyoung Ham, Young Jun Kim, Ye Rin Lim, Phuong Thi Hoang, Yongjun Lee, Seok-Won Lim, Jun-Mo Kim, Seungchan Cho, Jin-Won Song, Sukchan Lee, Won-Keun Kim, bioRxiv, 2020.11.25.398909; doi: https://doi.org/10.1101/2020.11.25.398909, https://www.biorxiv.org/content/10.1101/2020.11.25.398909v1
- Peer reviewed and published scientific report.
Lee, Gunsup, Shailesh Budhathoki, Geum-Young Lee, Kwang-ji Oh, Yeon Ham, Young-Jun Kim, Ye Lim, et al. 2021. “Broad-Spectrum Antiviral Activity of 3D8, a Nucleic Acid-Hydrolyzing Single-Chain Variable Fragment (ScFv), Targeting SARS-CoV-2 and Multiple Coronaviruses in Vitro.” Viruses 13 (4): 650. https://doi.org/10.3390/v13040650. https://www.mdpi.com/1999-4915/13/4/650.