Precise quantification and annotation of individual lipids is critical for understanding the complicated lipid function in biological systems. But achieving both concurrently is challenging. Researchers are turning to increasingly sophisticated instrumentation and techniques in the hopes of resolving this.
This article showcases TimsMetabo, a company providing accurate lipid quantification in low-input samples, key to studying the biological activities of low-abundance lipids.
High-confidence annotation of low-abundant lipid fragments enables researchers to derive relevant interpretations of lipid functions and metabolic processes from lipidomics data.

Image Credit: Bruker Daltonics
The challenge
Co-elution of interfering chemicals, poor MS performance, and chimeric MS/MS spectra unsuitable for automated annotation all impede accurate quantification and annotation of individual lipids in complex and diluted biological samples.
Solution
Bruker's innovative TimsMetabo LC-TIMS-QTOF solves these challenges by combining LC and TIMS selectivity with remarkable performance in MS and MS/MS modes, allowing for the detection of low-abundance lipid species and the generation of precise fragment spectra.
Boosting quantitation and annotation confidence with enhanced sensitivity
Improved lipid detection allows accurate quantitation even in diluted samples and for low-abundance precursors and fragments.
It can increase annotation confidence by revealing low-abundance fragment ion peaks in MS/MS spectra, leading to more confident, and in many cases higher-level (species-to-molecular species), lipid annotations.
Exceptional performance in MS → accurate quantification at low sample amounts
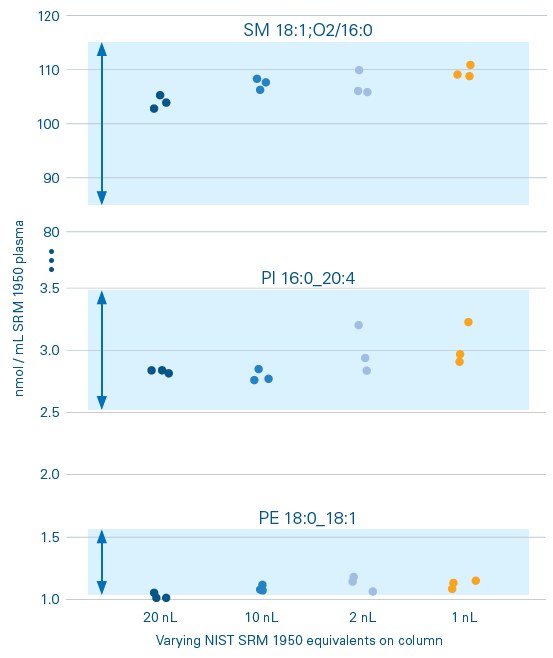
Image Credit: Bruker Daltonics

Figure 1. Displays lipid concentrations for three lipid species (Phosphatidylethanolamine (PE), Phosphatidylinositol (PI), and Sphingomyelin (SM)) analyzed in negative mode. Each dot represents the concentration measured in individual samples across the dilution series. The boxes indicate expected concentration ranges from the interlaboratory study by Bowden et al. (2017)3. All measured concentrations (nmol/mL) fall within these reference ranges, confirming accurate quantification down to a 1:1000 dilution of 1 μL plasma extract. Image Credit: Bruker Daltonics
Lipid extraction and analysis
Prof. Michal Holčapek and his team (Pardubice, Czech Republic) isolated lipids from NIST SRM 1950 reference plasma using MTBE.1
To enable precise lipid quantification, the extracts were spiked with a bespoke mixture of deuterated internal lipid standards, following the recommendations of the lipidomics standards effort.
Extracts were diluted to yield 20 nL to 1 nL equivalents of SRM 1950 plasma. The injection volume was 1 µL. The data was collected using a 26-minute reversed-phase LC procedure in conjunction with the innovative TimsMetabo TIMS-MS equipment in 4D-PASEF® negative mode.2
Representative lipid species from three different classes were identified and quantified. PE 18:0_18:1 was standardized to PE 15:0/18:1-d7. PI 16:0_20:4 was normalized to PI 15:0/18:1-d7. SM 18:1;O2/16:0 was normalized to SM 18:1;O2/18:1-d9.
Results and discussion
Figure 1 shows the concentrations of three lipids obtained in a dilution series of SRM 1950 plasma extracts using negative ionization mode.
Lipid concentrations may be reliably detected down to 1 nL SRM 1950 plasma extract equivalents, which is equivalent to a 1 µL extract diluted up to 1:1000. The findings were consistent with the interlaboratory investigation by Bowden et al. (2017).3 The determined concentrations are constant across the acquired dilution series*.
*It is important to note that Bowden's reference concentrations may be influenced by numerous lipid species. In this work, the primary lipid species were compared, which may explain slight variations between observed and reference values.
Conclusion
The TimsMetabo platform provides accurate lipid quantification in both high-volume and limited-sample applications.
Exceptional performance in MS/MS → more confidence
![Extracted Ion Mobilogram for [M-H]<sup>–</sup> of PE 16:0_18:2](https://www.news-medical.net/images/appnotes/ImageForAppNote_6175_17713251657176953.png)
Figure 2. Extracted Ion Mobilogram for [M-H]– of PE 16:0_18:2. Image Credit: Bruker Daltonics
![Extracted Ion Mobilogram for [M+H]<sup>+</sup> of PC 18:1_22:6](https://www.news-medical.net/images/appnotes/ImageForAppNote_6175_17713251877037021.png)
Figure 3. Extracted Ion Mobilogram for [M+H]+ of PC 18:1_22:6. Image Credit: Bruker Daltonics
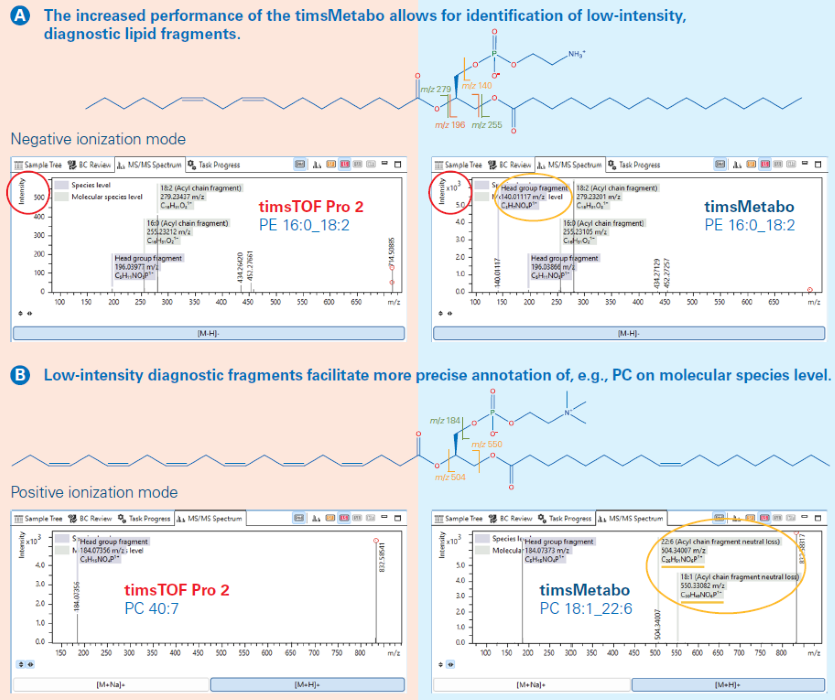
Figure 4. Comparison of MS/MS spectra on timsTOF Pro2 and timsMetabo based on A PE 16:0_18:2 in negative ion mode and B PC 40:7 (PC 18:1_22:6) in positive ion mode, respectively. Molecular structures are shown, highlighting the fragments observed in the PASEF MS/MS spectra. Image Credit: Bruker Daltonics
Lipid extraction and analysis
Lipids were isolated from NIST SRM 1950 reference plasma using IPA, using an NPC-adapted technique.4 The extracts were diluted, and 66 nL plasma equivalents were put into the column.
A 14-minute reversed-phase LC method was used in conjunction with both the timsTOF Pro2 and the innovative TimsMetabo TIMS-MS systems, using 4D-PASEF acquisition in both ionization modes.
Results and discussion
Figures 2 and 3 show the Extraction Ion Mobilograms (EIMs) for [M-H]- of PE 16:0_18:2 and [M+H]+ of PC 18:1_22:6, respectively. TimsMetabo resulted in a ∼7-15x increase in precursor ion intensity compared to timsTOF Pro 2, indicating better ion transfer.
TIMS separation paired with PASEF acquisition produces cleaner MS/MS spectra, which are critical for accurate lipid identification. TimsMetabo improves sensitivity in both MS and MS/MS by synchronizing TIMS-MX and Athena Ion Processor (AIP) actions, increasing annotation confidence.
Figure 4A compares the MS/MS spectra of PE 16:0_18:2 obtained with both systems in negative mode. TimsMetabo has a ∼10× higher sensitivity and a unique 140 m/z fragment, confirming the head group (Figure 4B) and validating the PE 16:0_18:2 annotation.
Figure 4C shows the MS/MS spectra for PC 18:1_22:6, demonstrating TimsMetabo's increased sensitivity. Unlike conventional platforms, TimsMetabo detects low-abundance diagnostic fragment ions at m/z 504 and 550 (acyl chain neutral losses), allowing for accurate molecular species annotation.
This increased specificity is a big step forward in routine lipidome screening of complex materials, paving the way for more insights into metabolic pathways.
Conclusion
TIMS separation, along with increased ion transfer in TimsMetabo, increases the identification of low-abundance fragments, such as neutral losses and headgroup ions. This results in a large increase in reliable lipid annotations at the molecular species level.
Prof. Michal Holčapek: University of Pardubice, Czech Republic
“The sensitivity of the new TimsMetabo instrument is truly impressive. It enables researchers to accurately quantify lipids in complex extracts using MS data and provides more specific molecular species-level annotations thanks to its enhanced MS/MS performance.”
Unlock the power of precision lipidomics with timsMetabo
Bruker's TimsMetabo LC-TIMS-QTOF improves lipidomics sensitivity and confidence. Designed to address the issues of low-abundance lipid detection and complex sample matrices, TimsMetabo provides
- 4D-TIMS separation provides cleaner spectra and up to 10x higher MS and MS/MS sensitivity
- Accurate measurement down to 1.0 nL plasma equivalents
- Significantly increased lipid annotations at the molecular species level
Whether you are working with valuable clinical samples or pushing the limits of lipid biomarker research, TimsMetabo gives you unrivaled annotation confidence and quantification accuracy - even at high dilutions
References
- Matyash, V., et al. (2008). Lipid extraction by methyl-tert-butyl ether for high-throughput lipidomics. Journal of Lipid Research, 49(5), pp.1137–1146. DOI: 10.1194/jlr.d700041-jlr200. https://www.sciencedirect.com/science/article/pii/S002222752042601X,
- Vaňková, Z., et al. (2022). Retention dependences support highly confident identification of lipid species in human plasma by reversed-phase UHPLC/MS. Analytical and bioanalytical chemistry, (online) 414(1), pp.319–331. DOI: 10.1007/s00216-021-03492-4. https://link.springer.com/article/10.1007/s00216-021-03492-4.
- Bowden, J.A., et al. (2017). Harmonizing lipidomics: NIST interlaboratory comparison exercise for lipidomics using SRM 1950–Metabolites in Frozen Human Plasma. Journal of Lipid Research, 58(12), pp.2275–2288. DOI: 10.1194/jlr.m079012. https://linkinghub.elsevier.com/retrieve/pii/S0022227520335549.
- npc-open-lcms/Protocols/NPC.SOP.MS003_v2.1_PP_Lipid blood.pdf at main · phenomecentre/npc-open-lcms · GitHub
About Bruker Daltonics
Empowering Science – Improving Life
Bruker Daltonics delivers cutting-edge mass spectrometry solutions and workflows that help scientists and industry leaders tackle real-world challenges and make new discoveries. From life sciences and pharmaceutical research to food and contaminant analysis, environmental monitoring, forensics, and industrial quality control, our technologies and instruments provide the precision and reliability you need to make confident decisions.
Our innovative platforms - such as timsTOF, scimaX, neofleX, and DART-TQ - combined with advanced software like SCiLS™ Lab, MetaboScape®, and Biopharma Compass®, transform complex data into actionable insights. Breakthrough innovations like Trapped Ion Mobility (TIMS), Omnitrap®, and dual ionization GC-HRMS are redefining what’s possible in mass spectrometry.
Trusted by leading research institutes, universities, government agencies, and industrial partners worldwide, Bruker Daltonics is committed to driving scientific progress and delivering solutions that matter.
(For Research Use Only. Not for use in clinical diagnostic procedures).
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.net, which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.