The brain’s major excitatory neurotransmitter is glutamate. Glutamate, which is released from presynaptic vesicles, is the neurotransmitter which activates postsynaptic ligand-gated ion channel receptors (NMDA, AMPA and kainate receptors) to secure fast synaptic transmission.1 It also activates metabotropic glutamate (mGlu) receptors which, as well as modulating the activity of other synapses, also modulate its release and postsynaptic response.2,3
Many other neuropathologies are linked to glutamate such as anxiety, pain, ischemia, epilepsy, schizophrenia, and Parkinson’s disease. Because of these modulating properties, mGlu receptors have been firmly established as promising therapeutic targets.3,4 It has been predicted that drugs acting at mGlu receptors will regulate the glutamatergic system without compromising the normal synaptic transmission.
mGlu receptors are forms of G-protein coupled receptors (GPCRs). Three groups have been formed (I-III) from the identification and classification of eight subtypes, based upon their sequence homology, transduction mechanism and pharmacological profile (see Figure 1). The first group, Group I, includes mGlu1 and mGlu5 receptors which couple to Gq and activate phospholipase C (PLC).
The second of these, Group II (mGlu2, mGlu3) and the third, group III (mGlu4, mGlu6, mGlu7, mGlu8) contain receptors which couple Gi /Go and inhibit adenylyl cyclase (AC). Group I receptors are mostly located postsynaptically, and therefore their activation increases excitability. In contrast, Group II/III receptors are largely presynaptic and the activation thereof lessens glutamate release. For each group, selective ligands have been located, and some of the subtypes, as detailed in the following descriptions.2,5–8
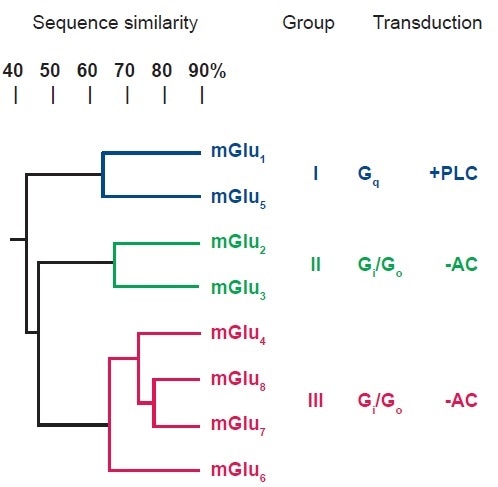
Figure 1. Classification of the Subtypes of mGlu Receptors. Image credit: Tocris Bioscience
mGlu receptors are Class C members of the GPCR superfamily.9 Similarly to all GPCRs, they have a heptahelical domain (HD) in the membrane region. Additionally, much like all other members of class C, a large extracellular amino-terminal domain (ATD) where the glutamate binding site is found characterizes all these receptors (see Figure 2).
A bilobate structure similar to LIVBP (Leucine Isoleucine Valine Binding Protein) is adopted by this protein, which is a bacterial periplasmic protein involved in the transport of hydrophobic amino acids.10–13 These amino acids bind to an open conformation of the protein, which traps them in between the two lobes by closing. For glutamate and competitive agonists in the LIVBP domain (LIVBPD) of mGlu receptors (see Figure 2), a similar binding mode has been proposed.
Furthermore, it has been demonstrated that receptor activation requires the closed conformation of this domain.14 Analysis of the glutamate binding site in the eight mGlu receptor subtype crystal structures (mGlu1, mGlu3, mGlu7)12,13 or homology models15–19 has revealed a common binding motif for the a-amino and a-carboxylic functions of glutamate,20 although residues that bind the distal g-carboxylate vary.17 Therefore, predictably, all competitive agonists are a-amino acids, possessing various selective functional groups on their side chain6 including virtual screening hits and derivatives. The second generation of allosteric modulators which bind in the HD followed first generation of orthosteric ligands.21 CPCCOEt was the first molecule to be described as a non-competitive mGlu receptor antagonist, in the late 90s.22 Since that period, we have seen the discovery of numerous allosteric modulators by high-throughput screening (HTS) in pharmaceutical companies.7,8,23
This article is designed to review our current pharmacological knowledge of mGlu receptors. Thanks to the publication of several detailed reviews2,4–6,8,24 this article will only present the most potent and selective known ligands and its emphasis placed on compounds that were more recently disclosed.
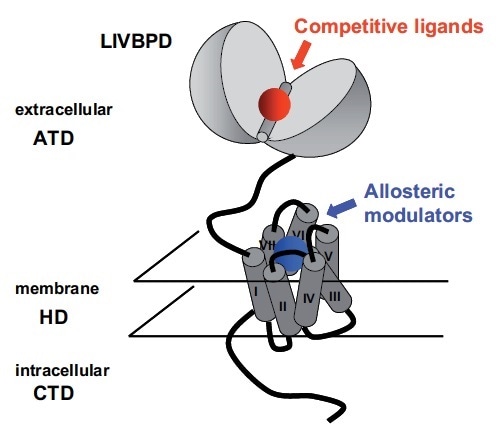
Figure 2. Schematic Representation of an mGlu Receptor: The Two Orthosteric and Allosteric Binding Sites are Indicated. Image credit: Tocris Bioscience
Competitive Ligands
An a-amino acid moiety can be located in all mGlu receptor competitive ligands (agonists and antagonists) and the majority of the side chains hold an acidic function. The spatial disposition of these functional groups, in the ligand active conformations, is that of glutamate in an extended conformation (as anticipated by pharmacophore25 and homology models17 and found in X-ray structures12,13). These compounds have been considered for years as valuable research tools, but have not been considered as drug candidates due to their poor partition coefficient (LogP), which is related to their lack of selectivity and highly polar chemical structures.
Eli Lilly researchers took the lead in demonstrating how such a glutamate analog, LY 354740, could pass the blood-brain barrier and to demonstrate that its peptidyl prodrug, LY 544344, was orally active as an anticonvulsant and anxiolytic.26,27 LY404039, a sulfonyl analog, reached phase III clinical trials, being orally administered as a methionine amide prodrug LY 2140023 for the treatment of schizophrenia.28,29.
Another benefit is that drugs like this are hardly metabolized since they are already quite hydrophilic30, and few side effects are predicted. Other glutamate analogs were demonstrated to be systemically active such as (2R,4R)-APDC, 3′Me-CCG, 3′HM-CCG, (S)-DCPG, ACPT-I and LSP1- 2111. It was also feared that desensitization would occur with continuous activation in the case of group II/III receptors; however, they are resistant to agonist-induced desensitization.31 Together, these results rejuvenate an interest in mGlu receptor competitive ligands.
Agonists
Trans-ACPD (1S,3R isomer) was the first agonist that could discriminate between ionotropic and metabotropic glutamate receptors32. Despite its lack of subtype selectivity, this ligand contributed considerably to the study of mGlu receptors.2,5,24 It is a limited number of molecules that can possess agonist activity across all mGlu receptors. The most potent are the endogenous agonist L-glutamate, L-CCG-I and ABHxD-I.2,5,24 We can note that L-CCG-I and ABHxD-I are conformationally constrained, and mimic the bioactive extended glutamate conformation common to all mGlu receptors.25 Selectivity can be gained when adding new chemical groups onto these structures.
Group I
The most potent Group I agonist is Quisqualate (Quis), but it also activates AMPA receptors, which restricts its use. (S)-3,5-DHPG is the most widely used group I selective agonist, yet it exhibits only moderate potency.2,5,24 It has been claimed that CHPG33 and Z/E-CBQA34 can specifically activate mGlu5 receptors, though the affinity of CHPG33 is quite low. To date, no specific mGlu1 competitive agonists have been disclosed.
Group II
The first mGlu agonist reported to exhibit a nanomolar affinity was LY 354740.26 It is group II selective, as well as its oxy (LY 379268), thia (LY 389795)35 and sulfonyl derivatives (LY 404039).36 The potent activity was retained by introducing a fluorine atom at position 3 (MGS0008) or 6 (MGS0022) and, as in the case of MGS0028, further enhanced when a carbonyl group was added.37
These bicyclic glutamate analogs derive from the general agonist L-CCG-I, wherein increased potency and group II selectivity were gained through the second hydrocarbon ring. In contrast, it was shown that a methyl, hydroxymethyl or cyclopropyl substituent in the 3′ position (3′Me-CCG, 3′HM-CCG and 3′CP-CCG respectively) offered agonists which had a similar potency.38–40 A decreased affinity at mGlu2/3 is caused by replacement of the hydroxyl functionality at C3′ of 3′HM-CCG, by a sulfhydryl 3. This analog (3′SM-CCG), interestingly, remains an mGlu2 agonist but is a full antagonist at mGlu3.41
For the C4βmethyl-substituted analog of LY 354740 (LY 541850), a similar selectivity was also recorded.42 At this same position, substitution by a thiotriazole group (LY 2812233) confers different pharmacological activity at the two subtypes.43 These three compounds selectively activate mGlu2, while the only reported mGlu3 competitive agonist expected to discriminate between the two group II subtypes was NAAG. However, this theory was recently disproven.44 Other group II selective agonists with submicromolar affinity have been described, including (2R,4R)-APDC and DCG-IV.
Group III
The majority of potent group III selective agonists have a diacidic side chain that is able to interact with the highly basic distal binding pocket.17,19 The most potent are (S)-AP4 (L-AP4), (S)-thioAP4,45 (S)-SOP (L-SOP) and (1S,2R)- APCPr46,47 which all exhibit submicromolar affinities at cloned receptors (except for mGlu7, to which all binding affinities are weak). (S)-PPG,48,49 (S)-3,4-DCPG,50 ACPT-I and (+)-ACPT-III,51 (S)-PBPG52 and PCG-153 have additionally been reported as micromolar agonists.
Notably, a CCG derivative bearing a hydroxymethyl group in the 3′ position (3′HMCCG) bears a similar affinity for mGlu8 and mGlu2/3 receptors.39 Recently, PCEP, a new series of agonists deriving from a virtual screening hit was disclosed.54–56 LSP1-308157 is among these,
which demonstrates potency close to L-AP4 and LSP1-211158 and LSP4- 202256 that show a preference or selectivity for the mGlu4 receptor respectively.56
Additionally, following systemic injection in animal models, these agonists alleviate Parkinson’s disease and anxiolytic symptoms.58–60 However, very few group III mGlu receptor agonists are subtype-selective. FP429 is a full mGlu4 and partial mGlu8 agonist,61,62 N-benzyl- APDC (BnAPDC)62 and (S)-homoAMPA64 act at mGlu6 and (S)-3,4-DCPG at mGlu8 with an EC50 over 2 orders of magnitude lower than at other group III receptors.50 An interesting thing to note is that cinnabarinic acid, which is an endogenous metabolite of the kynurenine pathway, is a weak mGlu4 agonist, as the first orthosteric agonist with non-amino acid structure.65
Antagonists
The complete closing of the two lobes of the LIVBPD is prevented by most competitive antagonists. Substitution of the a-proton of glutamate analogs by a methyl group turns the corresponding agonists (4CPG, AP4 and L-CCG-I) into antagonists, which is the case for MCCG, MCPG and MAP4, or a bulkier group as seen in LY 341495. Nonetheless, when the residues responsible for the hindrance are mutated, agonist properties can be recovered.14 Closing can also be disturbed by ionic repulsion, as seen with ACPT-II.14
Group I
4-carboxyphenylglycine derivatives such as the widely used (S)- MCPG comprise the first generation of group I mGlu receptor antagonists. When the a-methyl group was changed to a-thioxanthylmethyl as seen in LY 367366, its affinity was improved; but this derivative can also antagonize group II/III receptor activation.5
Subsequently, the highest potency was to be found with a-substituted 3-carboxycyclobutylglycines such as LY 393675 (cis isomer) and its trans isomer,5 or a cis/trans mixture like LY 393053.66 The latter mixture thereof was proven to be systemically active and to inhibit both mGlu1 and mGlu5 as well as other group II/III mGlu receptors.66 Though mildly less potent, both LY 367385 (4C2MPG) and LY 339840 (4C3H2MPG) demonstrate subtype I selectivity;67 yet LY 367385 also demonstrably inhibited the cystine/glutamate exchanger.68 To date, no mGlu5 selective and competitive antagonists have been described.
Group II
The most potent group II antagonists are obtained when aryl substituents are introduced in specific positions of that glutamate analog, given that potent group II agonists derive from L-CCG-I. Therefore, LY 341495,5 a fluorinated derivative69 and XECCG69 holding a 9′-xanthylmethyl or 9′-xanthylethyl moiety in the a- or 3′-position, show nanomolar affinities.
Two substituted phenyl groups can replace the a-xanthyl moiety while retaining potency (e.g. mCD-CCG).71 As previously stated, stereospecific substitution at the 3-position of the agonist LY 354740 is vital for agonist/antagonist property.17,37,42 HYDIA71,73 and several O-benzyl derivatives such as MGS0039 display high competitive group II antagonist activity.74–76 With both LY 341495 and MGS0039, systemic and antidepressant-like effects were observed.74 Other arylalkyl substituted glutamate and glutamate analogs like ADED (LY 310225), (S)-BnQuis and NM-APDC exhibit group II selectivity with IC50 values in the micromolar range.5
Group III
To date, no highly potent and group III-selective competitive antagonists have been reported. (S)-AP4, the best agonist, becomes a moderate antagonist when it’s a-proton is substituted by a methyl group in MAP4. MCPG, a weak group I/II antagonist becomes a moderate group III antagonist when a phosphonate replaces the 4-carboxylate, such as in the case of MPPG.
Similar group III antagonist activity occurs with the addition of a substituent in the 3-position, but increases selectivity for group III over group II.76 The analog of MPPG bearing an a-cyclopropyl group, CPPG, displays slightly increased potency5,77 in the same range as DCG-IV, which is also a group II agonist.78 The nonselective antagonist LY 34149 produces the best acivity5.79
Allosteric Modulators
Allosteric modulators are the names given to non-competitive ligands which bind in the transmembrane heptahelical domain. Identification has taken the place of both negative (NAMs) and positive modulators (PAMs).7,8,23,80 NAMs do not affect agonist binding but inhibit receptor activation, while PAMs do not activate receptors alone but do enhance agonist activation. Only those mGlu receptor modulators for which biological activities are available will be presented here, of all those that have been described (mostly in patents). Generally, these compounds are highly potent and subtype-selective: not the case for most competitive ligands.
Group I
Both non-competitive inhibitors as well as enhancers have been disclosed for group I receptors.
mGlu1 Antagonists
CPCCOEt the first negative mGlu receptor modulator, has inspired numerous detailed studies.22,81,82 Particular emphasis has been given to specific residues of the HD that bind CPCCOEt, which were identified by a group from Novartis.22 Subsequently, HTS and optimization in various companies discovered other compounds with higher affinities.83
In no particular order, these include the following: NPS 239084,85 (NPS Pharmaceuticals Inc.), Bay 36- 762086 (Bayer AG), LY 45606687,88 and LY 45623689 (Eli Lilly), R21412785/JNJ 1625968590,91 (Johnson & Johnson), 3,5-dimethyl-pyrrole-2,4-dicarboxylic acid diesters (of which DMPPP is the most potent derivative92,93) (GlaxoSmithKline), several analogs of EM-TBPC94,95 (Hoffmann-La Roche), thiazolobenzimidazoles YM 298198,96 YM 20207497 and thienopyrimidine YM 230888 (Yamanouchi Pharma), triazafluorenones such as A 84172098 and more selective tetracyclic derivatives99 (Abbott Laboratories, Schering-Plough), CFMTI100,101 (Banyu Pharmaceutical Co.), pyrazines-2-carboxamides PChPC102 and azaquinazolines such as CMPPA103 (Pfizer) and adamantyl methanone AdPyM104 (Merz Pharmaceuticals).
A homology model was generated of the mGlu1 allosteric binding site and a binding mode proposed for EM-TBPC which was corroborated by mutagenesis and functional assays.94 In addition to this, it was proven that several inhibitors (R214127, CPCCOEt , NPS 2390, Bay 36- 7620) bind to this same site.85 Potentialities in the form of anxiolytic and analgesic effects have been recorded with allosteric mGlu1 receptor antagonists; though potential side effects including the possibilities of locomotor and cognition impairment were also discovered, an impediment to their development.83,105
mGlu1 Positive Modulators
Ro 01-6128, Ro 67-4853105,107 and Ro 67- 7476 were the first allosteric potentiators of rat mGlu1 receptors.108,109 Chimeric and mutated receptors were generated to validate the transmembrane localization of the binding site of these ligands, which are subtype I selective.108 An interesting point to note is the lack of effect had by Ro 67-7476 and Ro 01-6128 on human mGlu1 receptor activation, where Ro 67-4853 produces a pronounced enhancement.108
Though CDPPB was known as an mGlu5 selective potentiator, meaning that it has the phenyl substituent of the pyrazole core in the 4 rather than the 3 position, VU 71 –– it was discovered to be a selective mGlu1 potentiator, interacting with a distinct site from that of NAMs.110
Initially, mGlu5 Antagonists SIB 1757 and SIB 1893111 were located and optimized into MPEP112, since widely used to explore the physiological roles of mGlu5 receptors as a potential therapeutic target.113 Novartis-led investigations developed a methoxy derivative M-MPEP, which can be easily tritium-labeled114. Lead optimization generated AFQ056 (Mavoglurant), used in clinical trials for the symptomatic treatment of Fragile X Syndrome and Parkinson’s disease levodopa-induced dyskinesia (PD-LID).
The search for new ligands was prompted by the therapeutic potential of mGlu5 antagonists.115–117 Early series contained an alkyne core, but more recent efforts have concentrated on alternative chemotypes. A thiazol derivative of MPEP with improved aqueous solubility, MTEP, was described with similar high mGlu5 affinity118 as well as its tritium-labeled methoxymethyl derivative MM-MTEP,119,120 M-PEPy119 bipyridyl derivative MTEBP121 and fluorine derivatives for PET imaging (F-MTEB and SP203).122,123
Following on from these primary MPEP/ MTEP derivatives,124 multiple disubstituted alkyne compounds have been reported which include the following: ADX10059 (useful for migraine and gastroesophageal reflux but a cause of liver function abnormalities in patients); ADX48621117 (Dipraglurant), in phase II clinical trials for PD-LID; ethynylbenzamides (effective in anxiety models);125 ethynylpyrrolopyrazines;126 MRZ 8676127 and CTEP, which exhibits high oral bioavailability and a long half-life of 18h.128
Despite this, during the ethynyl series development, it became rapidly apparent that structural changes unexpectedly modulated the pharmacology (a “molecular switch”), which turned full NAMs into partial antagonists, PAMs or silent/neutral allosteric modulators (SAMs).129,130 This process is demonstrated with 5-MPEP, where moving the methyl substituent of the MPEP pyridyl ring to the neighboring carbon turns this analog into a neutral modulator,129 or demonstrated further still with the 5-(phenylethynyl) pyrimidine series wherein the 3-methylphenyl derivative is a potent antagonist and the 4-methyl isomer a potentiator.130
Multiple proposals have been made for biosteric replacements of the alkyne core. These include carboxamides,132 arylquinolines,133,134 heterocycles (e.g. tetrazole),135 oxazolo-azepine136 or oxadiazole (VU 0285683).137 HTS campaigns, in parallel to this, provided new scaffolds which were modulated into a plethora of chemical structures116, for instance: aryl benzoxazoles138 (illustrated by BOMA), dipyridyl amides (ACDPP),139 phenyloxadiazoles and phenyltetrazoles,140 carbamoyloximes,141 thiazolotriazoles (such as GSK 2210875),142 pyrrolidinylpyridines,143 piperidylamides,144 benzimidazoles,145 and anilinoquinazolines.146
In one such campaign, results demonstrated that the known anxiolytic drug fenobam was in reality a potent non-competitive mGlu5 antagonist.147 New derivatives were also developed based on this discovery.148 Several reviews have detailed potential therapeutic application of mGlu5 antagonists.3,115,124,149 In addition, molecular determinants of the high-affinity binding site of MPEP have been defined150 and a strong similarity with critical residues of the mGlu1 binding site has been observed.151
mGlu5 Positive Modulators
The development of mGlu5 potentiators as promising new antipsychotics was reported based on the first PAMs.152 In consequence, these reports encouraged numerous research programs conducted over the last few years.152 DFB,153 CPPHA,154,155 CDPPB156,157 and ADX47273 were the first mGlu5 PAMs to be identified.158,159 Substituent modifications of mGlu5 PAMs led to a molecular switch, as observed with mGlu5 NAMs: replacing the fluorine atoms of DFB by methoxy groups transmutes this ligand into an antagonist, while a neutral modulator is dichlorobenzaldazine (DCB) which attenuates the potentiation conferred by DFB.153
Like modulations were reported with close analogs of MPEP, as detailed above.130,160 Development of acetylenic mGlu5 PAM led to MRZ 3573,151 VU 0360172137 and to the 2-aminomethyl-pyrimidine phenylethynyl derivative that, unlike several other mGlu5 PAMs that are ‘ago-potentiators’, is a pure PAM’.131 Various efforts to ameliorate these PAMs’ metabolic stability resulted in the N-aryl piperazine (VU 0364289)161,162 and piperidine amide series162 and the phenoxymethyl pyridooxazines that are devoid of phenylacetylene and carbonyl functionalities.163
Molecular switching164 was also recorded when building SAR around the ADX47273 structure165 but was not recorded around CDPPB 166. This is despite the fact that moving a phenyl substituent changes the selectivity of VU 1545 (while mGlu5 PAM) to VU 71 (an mGlu1 PAM). HTS also identified a benzamide scaffold, chemical modulation led to the discovery of VU 0357121 but also to neutral (silent) modulators (such as VU 0365396).167 CPPHA and analogs seem to bind to a different site than MPEP while ethynyl PAM and NAM binding sites overlap.
mGlu2 Positive Modulators
The launch of numerous research programs was instigated by the potential treatment of psychiatric diseases with mGlu2 potentiators, which additionally resulted in the discovery of multiple modulators.168,169 A pyridylmethylsulfonamide, LY487379, was the first of its kind reported to potentiate the activity of glutamate at mGlu2 receptors with an EC50 value of 0.3 μM and to be highly selective for this subtype.170
Also, LY 487379 binds to a pocket in the transmembrane domain which is different from the orthosteric site in the ATD, it was demonstrated.170 The discovery of 1-methylbutoxy analog occurred after further SAR studies (2,2,2-TEMPS) with improved potency (EC50 = 14 nM) and selectivity.171,172 Soon after, a new chemical series of phenyl-tetrazolyl acetophenones (e.g. PTBE) was disclosed as selective mGlu2 potentiators,173 followed by extensive SAR studies.174–177 Later, new chemotypes were disclosed after further SAR studies and HTS hits.178 The compounds presented here are largely those which were selected among the series for in vivo assays and those which provide the best compromise between metabolic stability and potency: biphenylindanone (BINA),179 recently optimized into benzizothiazolone,180 benzimidazole-1181 and benzimidazole-2 (GSK 1331268),182 oxazolidinone-1183 and oxazolidinone-2184 optimized into oxazolobenzimidazoles (TBPCOB),185 imidazopyridine,186 1,5-disubstituted pyridine,187 imidazole carboxamide (THIIC),188 isoquinolones,189 and imidazomethylpiperidine.190
mGlu2/3 Antagonists
Only mGlu2/3 NAMs have been disclosed to date, in the main by the researchers at Hoffmann-La Roche.169 Heterocyclic enol ethers, like Ro 64-5229, were reported as the first selective noncompetitive mGlu2 receptor antagonists.191 Later, a series of dihydrobenzo[b][1,4] diazepin-2-one derivatives was disclosed, which displayed nanomolar inhibition of receptor activation by LY 354740.192 Like Ro 4491533 that was tested in vivo, this series was ameliorated further in several derivatives.193–196
Two novel antagonists, Ro 4988546 (from a new pyrazolo[1,5-a]pyrimidine scaffold) and Ro 5488608, were more recently disclosed and used in an investigation of the structural determinant at the mGlu2 NAM binding site.197
mGlu3
Specific mGlu3 PAMs an NAMs were provided by a recent screening campaign. However, at the time of writing, the chemical structures had not yet been disclosed.196 In an interesting development, it was found that varying a substituent on the PHCCC structure gave rise to a mGlu2/3 SAM or conferred dual mGlu2 NAM - mGlu3 PAM properties.199
Group III
The latest to be identified were Group III modulators, largely including mGlu4 potentiators. Initially described as an mGlu1 receptor antagonist,81 PHCCC was the first mGlu4 receptor PAM to be found as its (–) enantiomer.200,201 SIB 1893 and MPEP, two other mGlu5 antagonists, were reported to enhance agonist potency and efficacy at human mGlu4 at higher concentrations.202
Subsequently, several mGlu4 PAMs were discovered by HTS and hit optimization: VU 0155041;203 a series of phenylpicolinamides VU 0361737,204 VU 0364439,205 VU 366037;206 styryl aminopyrimidine;207 and thiazolopyrazole.208,209 Though several of these ligands showed good brain penetration and benefits in motor dysfunction models, they may possess intrinsic agonist activity (as in the case of VU 0155041) and have therefore been named as ago-potentiators.203 AMN 082 was described as an mGlu7 allosteric agonist210 and yet a fast metabolism was revealed by a recent study.211 Isoxazolpyridones such as MMPIP were determined as mGlu7 antagonists212,213 however, this may be dependent on context.214 An MGlu8 PAM, AZ 12216052, was proved to be systemically active in an animal model of anxiety.215
Conclusion
Early on, mGlu receptor molecular pharmacology efforts offered group selective competitive ligands. Though now it appears possible to uncover subtype-selective orthosteric ligands, allosteric modulators are responsible for most of the recent advances. Generally, these compounds are highly selective and potent.
Furthermore, many of these compounds exhibit in vivo activity and may pave the way to new therapeutic agents. Though we still await some further subtype-selective compounds, particularly for group III mGlu receptors, the panel of available mGlu receptor ligands is now broad. This panel has potential to shed new light on the physiological and pathological roles of the various mGlu receptor subtypes in both the normal and diseased brain. These investigations are currently ongoing and the ensuing developments offer up an exciting future.
Special Mentions
This white paper was based on the original work of:
Francine C. Acher:
Francine C. Acher is currently a CNRS Research Director within the Biomedical Institute of University Paris‑V (France). Her research focuses on structure/function studies and drug discovery using chemical tools (synthetic chemistry, molecular modeling), molecular biology and pharmacology within interdisciplinary collaborations.
Laboratoire de Chimie et Biochimie Pharmacologiques et Toxicologiques, UMR8601-CNRS, Université René Descartes-Paris V, Paris, France
Email: [email protected]
References
- Dingledine et al. (1999) Pharmacol.Rev. 51 7.
- Pin and Acher (2002) Curr.Drug Targets: CNS Neurol.Disord. 1 297.
- Niswender and Conn (2010) Annu.Rev.Pharmacol.Toxicol. 50 295.
- Recasens et al. (2007) Curr.Drug Targets 8 651.
- Schoepp et al. (1999) Neuropharmacology 38 1431.
- Pin et al. (1999) Eur.J.Pharmacol. 375 277.
- Gasparini and Spooren (2007) Curr.Neuropharmacol. 5 187.
- Urwyler (2011) Pharmacol.Rev. 63 59.
- Pin et al. (2003) Pharmacol.Ther. 98 325.
- O’Hara et al. (2003) Neuron 11 41.
- Bessis et al. (2000) Protein Sci. 9 2200.
- Kunishima et al. (2000) Nature 407 971.
- Muto et al. (2007) Proc.Natl.Acad.Sci.U.S.A. 104 3759.
- Bessis et al. (2002) Proc.Natl.Acad.Sci.U.S.A. 99 11097.
- Hampson et al. (1999) J.Biol.Chem. 274 33488.
- Malherbe et al. (2001) Mol.Pharmacol. 60 944.
- Bertrand et al. (2002) J.Med.Chem. 45 3171.
- Rosemond et al. (2002) J.Biol.Chem. 277 7333.
- Rosemond et al. (2004) Mol.Pharmacol. 66 834.
- Acher and Bertrand (2005). Biopolymers 80 357.
- May and Christopoulos (2003) Curr.Opin.Pharmacol. 3 551.
- Litschig et al. (1999) Mol.Pharmacol. 55 453.
- Kew (2004) Pharm.Ther. 104 233.
- Conn and Pin (1997) Ann.Rev.Pharmacol.Toxicol. 37 205.
- Bessis et al. (1999) Neuropharmaco logy 38 1543.
- Monn et al. (1997) J.Med.Chem. 40 528.
- Dunayevich et al. (2008) Neuropsychopharmacology 33 1603.
- Patil et al. (2007) Nat.Med. 13 1102.
- Mezler et al. (2010) Curr.Opin.Investig.Drugs 11 833.
- Johnson et al. (2002) Drug Metab.Disp. 30 27.
- Lennon et al. (2010) Eur.J.Pharmacol. 649 29.
- Palmer et al. (1989) Eur.J.Pharmacol. 166 585.
- Doherty et al. (1997) Neuropharmacology 36 265.
- Littman et al. (1999) J.Med.Chem. 42 1639.
- Monn et al. (1999) J.Med.Chem. 42 1027.
- Monn et al. (2007) J.Med.Chem. 50 233.
- Nakazato et al. (2000) J.Med.Chem. 43 4893.
- Collado et al. (2002) J.Med.Chem. 45 3619.
- Collado et al. (2004) J.Med.Chem. 47 456.
- Stanley et al. (2010) Bioorg.Med.Chem. 18 6089.
- Gonzalez et al. (2005) Bioorg.Med.Chem. 13 6556.
- Dominguez et al. (2005) J.Med.Chem. 48 3605.
- Monn et al. (2011) Curr.Neuropharmacol. 9 Suppl.1, 44.
- Chopra et al. (2009) J.Pharmacol.Exp.Ther. 330 212.
- Selvam et al. (2007) J.Med.Chem. 50 4656.
- Kroona et al. (1991) J. Med. Chem. 34 1692.
- Sibille et al. (2007) J.Med.Chem. 50 3585.
- Gasparini et al. (1999) J.Pharm.Exp.Ther. 290 1678.
- Gasparini et al. (2000) Bioorg.Med.Chem.Lett. 10 1241.
- Thomas et al. (2001) Neuropharmacology 40 311.
- Acher et al. (1997) J.Med.Chem. 40 3119.
- Filosa et al. (2006) Bioorg.Med.Chem. 14 3811.
- Amori et al. (2006) Bioorg.Med.Chem.Lett. 16 196.
- Selvam et al. (2010) J.Med.Chem. 53 2797.
- Triballeau et al. (2005) J.Med.Chem. 48 2534.
- Selvam et al. (2011) submitted.
- Cuomo et al. (2009) J.Neurochem. 109 1096.
- Beurrier et al. (2009) FASEB J. 23 3619.
- Wieronska et al. (2010) Neuropharmacology 59 627.
- Goudet et al. (2011) submitted.
- Schann et al. (2006) Bioorg.Med.Chem.Lett. 16 4856. References
- Frauli et al. (2007) Mol.Pharmacol. 71 704.
- Tückmantel et al. (1997) Bioorg.Med.Chem.Lett. 7 601.
- Ahmadian et al. (1997) J.Med.Chem. 40 3700.
- Fazio et al. (2012) Mol.Pharmacol. 81 643.
- Chen et al. (2000) Neurosci. 95 787.
- Kingston et al. (2002) Neurosci.Lett. 330 127.
- Melendez et al. (2005) J.Pharmacol.Exp.Ther. 314 139.
- Sakagami et al. (2008) Bioorg.Med.Chem. 16 4359.
- Pellicciari et al. (2001) Bioorg.Med.Chem.Lett. 11 3179.
- Sørensen et al. (2003) Bioorg.Med.Chem. 11 197.
- Woltering et al. (2008) ChemMedChem 3 323.
- Lundstrom et al. (2009) ChemMedChem 4 1086.
- Chaki et al. (2004) Neuropharmacology 46, 457.
- Nakazato et al. (2004) J.Med.Chem. 47 4570.
- Yasuhara et al. (2006) Bioorg.Med.Chem. 14 3405. 77. Conway et al. (2001) Bioorg.Med.Chem.Lett. 11 777.
- Brabet et al. (1998) Neuropharmacology 37 1043.
- Wright et al. (2000) Naunyn Schmiedebergs Arch.Pharmacol. 362 546.
- Sabbatini and Micheli (2004) Expert Opin.Ther.Patents 14 1593.
- Annoura et al. (1996) Bioog.Med.Chem.Lett. 6 763.
- Ott et al. (2000) J.Med.Chem. 43 4428.
- Owen (2011) ACS.Chem.Neurosci. 2 394.
- Van Wagenen et al. (1998) Society for Neuroscience Abstract 24 576.
- Lavreysen et al. (2003) Mol.Pharmacol. 63 1082.
- Caroll et al. (2001) Mol.Pharmacol. 59 965.
- Li et al. (2002) Neuropharmacology 43 A79.
- Ambler and Baker (2001) Patent WO 01/32632.
- Shannon et al. (2005) Neuropharmacology 49 188.
- Lavreysen et al. (2004) Neuropharmacology 47 961.
- Mabire et al. (2005) J.Med.Chem. 48 2134.
- Micheli et al. (2003) Bioorg.Med.Chem. 11 171.
- Di Fabio et al. (2007) Bioorg.Med.Chem.Lett. 17 2254.
- Malherbe et al. (2003) J.Biol.Chem. 278 8340.
- Binggeli et al. (2002) Patent WO02051418.
- Kohara et al. (2005) J.Pharmacol.Exp.Ther. 315 163.
- Kohara et al. (2008) Brain Res. 1191 168.
- Zheng et al. (2005) J.Med.Chem. 48 7374.
- Wu et al. (2007) J.Med.Chem. 50 5550.
- Ito et al. (2009) Bioorg.Med.Chem.Lett 19 5310.
- Satow et al. (2009) J.Pharmacol.Exp.Ther. 330 179.
- Owen et al. (2007) Bioorg.Med.Chem.Lett. 17 486.
- Mantell et al. (2009) Bioorg.Med.Chem.Lett. 19 2190.
- Noeske et al. (2009) Bioorg.Med.Chem. 17 5708.
- Lesage and Steckler (2010) Eur.J.Pharmacol. 639 2.
- Bleicher et al. (2000) Patent WO0063166.
- Vieira et al. (2009) Bioorg.Med.Chem.Lett. 19 1666.
- Knoflach et al. (2001) Proc.Nat.Acad.Sci. 98 13402.
- Wichmann et al. (2002) Farmaco. 57 989.
- Hemstapat et al. (2006) Mol.Pharmacol. 70 616.
- Varney et al. (1999) J.Pharmacol.Exp.Ther. 290 170.
- Gasparini et al. (1999) Neuropharmacology 38 1493.
- Spooren et al. (2001) T.I.P.S. 22 331.
- Gasparini et al. (2002) Bioorg.Med.Chem.Lett. 12 407.
- Jaeschke et al. (2008) Expert.Opin.Ther.Pat. 18 123.
- Emmitte (2011) ACS.Chem.Neurosci. 2 411.
- Rocher et al. (2011) Curr.Top Med.Chem. 11 680.
- Cosford et al. (2003) J.Med.Chem. 46 204.
- Cosford et al. (2003) Bioorg.Med.Chem.Lett. 13 351.
- Anderson et al. (2002) J.Pharmacol.Exp.Ther. 303 1044.
- Roppe et al. (2004) Bioorg.Med.Chem.Lett. 14 3993.
- Simeon et al. (2007) J.Med.Chem. 50 3256. tocris.com | 19 Metabotropic Glutamate Receptors: Molecular Pharmacology
- Simeon et al. (2011) J.Med.Chem. 54 901.
- Slassi et al. (2005) Curr.Top.Med.Chem. 5 897.
- Gilbert et al. (2011) Bioorg.Med.Chem.Lett. 21 195.
- Micheli et al. (2008) Bioorg.Med.Chem.Lett. 18 1804.
- Dekundy et al. (2010) J.Neural Transm. 118 1703.
- Lindemann et al. (2011) J.Pharmacol.Exp.Ther. 339 474.
- Rodriguez et al. (2005) Mol.Pharmacol. 68 1793.
- Wood et al. (2011) Biochemistry 50 2403.
- Sharma et al. (2009) J.Med.Chem. 52 4103.
- Kulkarni et al. (2009) J.Med.Chem. 52 3563.
- Milbank et al. (2007) Bioorg.Med.Chem.Lett. 17 4415.
- Zhang et al. (2010) Bioorg.Med.Chem. 18 3026.
- Roppe et al. (2004) J.Med.Chem. 47 4645.
- Burdi et al. (2010) J.Med.Chem. 53 7107.
- Rodriguez et al. (2010) Mol.Pharmacol. 78 1105.
- Wang et al. (2004) Bioorg.Med.Chem. 12 17.
- Bonnefous et al. (2005) Bioorg.Med.Chem.Lett. 15 1197.
- Wagner et al. (2010) Bioorg.Med.Chem.Lett. 20 3737.
- Galambos et al. (2010) Bioorg.Med.Chem Lett. 20 4371.
- Pilla et al. (2010) Bioorg.Med.Chem.Lett. 20 7521.
- Weiss et al. (2011) Bioorg.Med.Chem.Lett. 21 4891.
- Spanka et al. (2010) Bioorg.Med.Chem.Lett. 20 184.
- Carcache et al. (2011) ACS.Med.Chem.Lett. 2 58.
- Felts et al. (2009) Bioorg.Med.Chem.Lett. 19 6623.
- Porter et al. (2005) J.Pharmacol.Exp.Ther. 315 711.
- Jaeschke et al. (2007) Bioorg.Med.Chem.Lett. 17 1307.
- Gasparini et al. (2008) Curr.Opin.Drug Discov.Devel. 11 655.
- Malherbe et al. (2003) Mol.Pharmacol. 64 823.
- Vanejevs et al. (2008) J.Med.Chem. 51 634.
- Stauffer et al. (2011) ACS.Chem.Neurosci. 2 450.
- O’Brien et al. (2003) Mol.Pharmacol. 64 731.
- O’Brien et al. (2004) J.Pharmacol.Exp.Ther. 309 568.
- Zhao et al. (2007) Bioorg.Med.Chem.Lett. 17 1386.
- Lindsley et al. (2004) J.Med.Chem. 47 5825.
- Kinney et al. (2005) J.Pharmacol.Exp.Ther. 313 199.
- Le Poul et al. (2005) Neuropharmacology 49 252.
- Liu et al. (2008) J.Pharmacol.Exp.Ther. 327 827.
- Sams et al. (2011) Bioorg.Med.Chem.Lett. 21 3407.
- Zhou et al. (2010) ACS.Chem.Neurosci. 1 433.
- Xiong et al. (2010) Bioorg.Med.Chem.Lett. 20 7381.
- Varnes et al. (2011) Bioorg.Med.Chem.Lett. 21 1402.
- Lamb et al. (2011) Bioorg.Med.Chem.Lett. 21 2711.
- Engers et al. (2009) ChemMedChem 4 505.
- de Paulis et al. (2006) J.Med.Chem. 49 3332.
- Hammond et al. (2010) ACS.Chem.Neurosci. 1 702.
- Trabanco et al. (2011) Curr.Med.Chem. 18 47.
- Sheffler et al. (2011) ACS.Chem.Neurosci. 2 282.
- Schaffhauser et al. (2003) Mol.Pharmacol. 64 798.
- Barda et al. (2004) Bioorg.Med.Chem.Lett. 14 3099.
- Hu et al. (2004) Bioorg.Med.Chem.Lett. 14 5071.
- Pinkerton et al. (2004) J.Med.Chem. 47 4595.
- Pinkerton et al. (2004) Bioorg.Med.Chem.Lett. 14 5867.
- Pinkerton et al. (2005) Bioorg.Med.Chem.Lett. 15 1565.
- Cube et al. (2005) Bioorg.Med.Chem.Lett. 15 2389.
- Govek et al. (2005) Bioorg.Med.Chem.Lett. 15 4068.
- Fraley (2009) Expert Opin.Ther.Pat. 19 1259.
- Bonnefous et al. (2005) Bioorg.Med.Chem.Lett. 15 4354.
- Dhanya et al. (2011) J.Med.Chem. 54 342.
- Zhang et al. (2008) Bioorg.Med.Chem.Lett. 18 5493.
- D’Alessandro et al. (2010) Bioorg.Med.Chem.Lett. 20 759.
- Duplantier et al. (2009) Bioorg.Med.Chem.Lett. 19 2524.
- Brnardic et al. (2010) Bioorg.Med.Chem.Lett. 20 3129.
- Garbaccio et al. (2010) ACS.Med.Chem.Lett. 1 406.
- Tresadern et al. (2010) Bioorg.Med.Chem.Lett. 20 175.
- Cid et al. (2010) ACS.Chem.Neurosci. 1 788.
- Fell et al. (2011) J.Pharmacol.Exp.Ther. 336 165.
- Trabanco et al. (2011) Bioorg.Med.Chem.Lett. 21 971.
- Zhang et al. (2011) J.Med.Chem. 54 1724.
- Kolczewski et al. (1999) Bioorg.Med.Chem.Lett. 9 2173.
- Woltering et al. (2007) Bioorg.Med.Chem.Lett. 17 6811.
- Woltering et al. (2008) Bioorg.Med.Chem.Lett. 18 1091.
- Woltering et al. (2008) Bioorg.Med.Chem.Lett. 18 2725.
- Woltering et al. (2010) Bioorg.Med.Chem.Lett. 20 6969.
- Hemstapat et al. (2007) J.Pharmacol.Exp.Ther. 322 254.
- Lundstrom et al. (2011) Br.J.Pharmacol.
- Pratt et al. (2011) Comb.Chem.High Throughput Screen 14 631.
- Schann et al. (2010) J.Med.Chem. 53 8775.
- Maj et al. (2003) Neuropharmacology 45 895.
- Marino et al. (2003) Proc.Natl.Acad.Sci.U.S.A. 100 13668.
- Mathiesen et al. (2003) Br.J.Pharmacol. 138 1026.
- Niswender et al. (2008) Mol.Pharmacol. 74 1345.
- Engers et al. (2009) J.Med.Chem. 52 4115.
- Engers et al. (2010) Bioorg.Med.Chem.Lett. 20 5175.
- Engers et al. (2011) J.Med.Chem. 54 1106.
- East et al. (2010) Bioorg.Med.Chem.Lett. 20 4901.
- East and Gerlach (2010) Expert Opin.Ther.Pat. 20 441.
- Hong et al. (2011) J.Med.Chem. 54 5070.
- Mitsukawa et al. (2005) Proc.Natl.Acad.Sci.U.S.A. 102 18712.
- Sukoff Rizzo et al. (2011) J.Pharmacol.Exp.Ther. 338 345.
- Suzuki et al. (2007) J.Pharmacol.Exp.Ther. 323 147.
- Nakamura et al. (2010) Bioorg.Med.Chem.Lett. 20 726. 214.
- Niswender et al. (2010) Mol.Pharmacol. 77 459
- Duvoisin et al. (2010) Behav.Brain Res. 212 168.
- Adam et al. (1999) Neuropharmacology 38 A1 abstract 3. 217. Naples and Hampson (2001) Neuropharmacology 40 170.
About Tocris Bioscience
Tocris Bioscience is your trusted supplier of high-performance life science reagents, including receptor agonists & antagonists, enzyme inhibitors, ion channel modulators, fluorescent probes & dyes, and compound libraries. Our catalog consists of over 4,500 research tools, covering over 400 protein targets enabling you to investigate and modulate the activity of numerous signaling pathways and physiological processes.
We have been working with scientists for over 30 years to provide the life science community with research standards, as well as novel and innovative research tools. We understand the need for researchers to trust their research reagents, which is why we are committed to supplying our customers with the highest quality products available, so you can publish with confidence.
Tocris is part of the protein sciences division of Bio-Techne, which also includes the best in class brands R&D Systems, Novus Biologicals, ProteinSimple, and Advanced Cell Diagnostics. Bio-Techne has united these brands to provide researchers with a full portfolio of research reagents, assays, and protein platforms. For more information on Bio-Techne and its brands, please visit bio-techne.com.
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices, and treatments.