Introduction
Continuous live-cell monitoring techniques have witnessed a significant development in recent years, transforming the way cells are used in the laboratory.
These technologies are able to extract valuable cellular information that was not possible before. Their features are also closely aligned with the key changes in the realm of cellular research.
In an effort for greater significance, biomedical researchers are moving from basic recombinant cell systems to stem-cell derived cells and primary cells that are usually human and specific to patients.
Sophisticated tissue organoid models and in vitro co-cultures are becoming increasingly common. Though target-specific assays will continue to be significant, cell-based, phenotypic assays are becoming more popular because they enable holistic measurement of integrated cell functions.
Although these advancements present opportunities for novel studies, they also bring new requirements for cellular workflows and technical complications. This article shows how these changing requirements are addressed by advancements in continuous live-cell monitoring and analysis techniques.
Immortalized cell lines
HEK, CHO, HeLa cells, and other classic cell lines are simple and routine. Through advanced molecular approaches, researchers are able to manipulate or express genes in these cell lines. Kit-based, turnkey methods can also be used to induce stable gene expression for various cell passages.
Recombinant cells are predictable, robust, and grow quickly, reliably, and homogeneously for a long duration of time without any major complexities induced by loss of expression. These cells are low-cost and high-yield, allowing investigators to discard unused ‘spare’ cells frequently.
Primary cells
Primary cells, for example neurons, can accurately show the in-vivo function of biological systems when compared to immortalized cell lines. However, they cannot be easily acquired in large numbers, and many also not divide, meaning they cannot be scaled-up further.
There are certain primary cell types that do not survive for an extended period of time in culture. The genetic manipulation of primary cells is also more complicated when compared to immortalized cells. Transfection procedures and special constructs may also be required.
Stem cells
Compared to immortalized and primary cells, stem cells provide better flexibility. With next-generation technology, fully differentiated mature cells can be derived from replicating stem cell pre-cursors.
Such cells provide several therapeutic and diagnostic applications, for instance, genetic pre-disposition and disease etiology can be studied by comparing the properties of cells obtained from different patients.
Differentiated cells and mesechymal stem cells can also be thought of as potential therapies. These methods combined with the recent developments in precision gene editing bring us closer to the concept of truly personalized medicine.
However, the procedures involved for differentiating, reprogramming, and scaling stem cells are quite lengthy, complicated, and poorly defined. In order to resolve these challenges, several commercial suppliers provide differentiated, cryopreserved cells. These cells are expensive and make it unfeasible for laboratories to use on a routine basis.
As stem cells mature and differentiate, their properties change over time and in many situations only a low yield of healthy, fully differentiated cells is obtained. Due to the dynamic nature of primary and stem-cell derived cultures, more emphasis is placed on the requirement for real-time decision making and active management by researchers, particularly in comparison to immortalized cells.
The role of the microenvironment
Another emerging factor is the extent to which the microenvironment influences cell behavior. In the quest to simulate true pathology and physiology, scientists are developing more comprehensive multi-culture and co-culture cell systems.
Growing cells in scaffolds and bio-matrices to form 3D microtissues and organoids is becoming popular and so are organ-on-a-chip systems (Figure 1). However, these experiments can be complicated, need a large amount of optimization, and place greater demands on the detection systems employed for analysis and quality control of target cells.
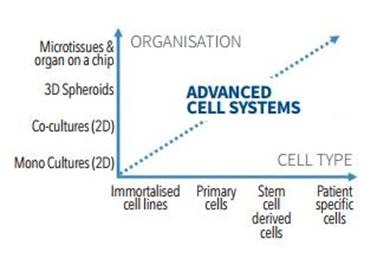
Figure 1. Researchers are combining more relevant cell types with more relevant organizational structures to create advanced and more translational model systems.
Cellular models are changing, the new commercially available cell systems may prove to be more translational and applicable but they are more expensive and complicated.
When using advanced cellular models, increased time and effort has to be devoted to the experimental studies. The cells are dynamic, requiring careful observation and they are valuable so shouldn’t be wasted.
 Sartorius
Sartorius
Sartorius is a leading international pharmaceutical and laboratory equipment supplier. With our innovative products and services, we are helping our customers across the entire globe to implement their complex and quality-critical biomanufacturing and laboratory processes reliably and economically.
The Group companies are united under the roof of Sartorius AG, which is listed on the Frankfurt Stock Exchange and holds the majority stake in Sartorius Stedim Biotech S.A. Quoted on the Paris Stock Exchange, this subgroup is comprised mainly of the Bioprocess Solutions Division.
Innovative Technologies Enable Medical Progress
A growing number of medications are biopharmaceuticals. These are produced using living cells in complex, lengthy and expensive procedures. The Bioprocess Solutions Division provides the essential products and technologies to accomplish this.
In fact, Sartorius has been pioneering and setting the standards for single-use products that are currently used throughout all biopharmaceutical manufacturing processes.
Making Lab Life Easier
Lab work is complex and demanding: Despite repetitive analytical routines, lab staff must perform each step in a highly concentrated and careful way for accurate results.
The Lab Products and Services Division helps lab personnel excel because its products, such as laboratory balances, pipettes and lab consumables, minimize human error, simplify workflows and reduce physical workloads
Sponsored Content Policy: News-Medical.net publishes articles and related content that may be derived from sources where we have existing commercial relationships, provided such content adds value to the core editorial ethos of News-Medical.Net which is to educate and inform site visitors interested in medical research, science, medical devices and treatments.